Originally published Jan. 14, 2014 and updated Apr. 14, 2020.
Any newcomer who joins a molecular biology lab will undoubtedly be asked to design, modify, or construct a plasmid. A plasmid is a small circular piece of DNA found in bacterial cells, and someone new to plasmids may need some extra guidance to understand the specific components that make up a plasmid and why each is important.
Our “Plasmids 101” series designed to educate all levels of scientists and plasmid lovers - serves as an introduction to plasmids. Plasmids 101 will provide you with an overview of general molecular biology knowledge and techniques, and empower you with a firm understanding of the fundamentals. Our mission is to curate a one-stop reference guide for plasmids, so that you can spend less time researching the basics and spend more time developing cleverly designed experiments and innovative solutions necessary for advancing the field.
What is a plasmid?
At their most basic level, plasmids are small circular pieces of DNA that replicate independently from the host's chromosomal DNA. They are mainly found in bacteria, but also exist naturally in archaea and eukaryotes such as yeast and plants. In nature, plasmids provide one or more functional benefits to the host such as resistance to antibiotics, degradative functions, and/or virulence. All natural plasmids contain an origin of replication (which controls the host range and copy number of the plasmid) and typically include a gene that is advantageous for survival, such as an antibiotic resistance gene.
In contrast, plasmids utilized in the lab are usually artificial and designed to introduce foreign DNA into another cell. Minimally, lab-created plasmids have an origin of replication, selection marker, and cloning site. The ease of modifying plasmids and the ability of plasmids to self-replicate within a cell make them attractive tools for the life scientist or bioengineer.
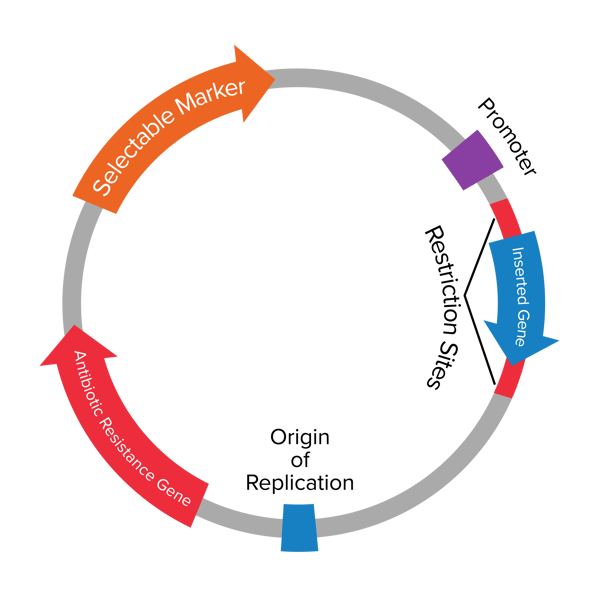 |
| Figure 1: Map of a plasmid with its elements described below. |
| Vector Element | Description |
| Origin of Replication (ORI) | DNA sequence which allows initiation of replication within a plasmid by recruiting replication machinery proteins |
| Antibiotic Resistance Gene | Allows for selection of plasmid-containing bacteria. |
| Multiple Cloning Site (MCS) | Short segment of DNA which contains several restriction sites allowing for the easy insertion of DNA. In expression plasmids, the MCS is often downstream from a promoter. |
| Insert | Gene, promoter or other DNA fragment cloned into the MCS for further study. |
| Promoter Region | Drives transcription of the target gene. Vital component for expression vectors: determines which cell types the gene is expressed in and amount of recombinant protein obtained. |
| Selectable Marker | The antibiotic resistance gene allows for selection in bacteria. However, many plasmids also have selectable markers for use in other cell types. |
| Primer Binding Site | A short single-stranded DNA sequence used as an initiation point for PCR amplification or sequencing. Primers can be exploited for sequence verification of plasmids. |
How is a plasmid constructed in the lab?
Due to their artificial nature, lab plasmids are commonly referred to as “vectors” or “constructs.” To insert a gene of interest into a vector, scientists may utilize one of a variety of cloning methods (restriction enzyme, ligation independent, Gateway, Gibson, etc). The cloning method is ultimately chosen based on the plasmid you want to clone into. Regardless, once the cloning steps are complete, the vector containing the newly inserted gene is transformed into bacterial cells and selectively grown on antibiotic plates.
Importantly, because the bacteria from which plasmids are isolated grow quickly and make more of the plasmids as they grow, scientists can easily make large amounts of plasmid to manipulate and use in later work.

How do scientists use plasmids?
Generally, scientists use plasmids to manipulate gene expression in target cells. Characteristics such as flexibility, versatility, safety, and cost-effectiveness enable molecular biologists to broadly utilize plasmids across a wide range of applications. Some common plasmid types include cloning plasmids, expression plasmids, gene knock-down plasmids, reporter plasmids, viral plasmids, and genome engineering plasmids.
Some of the many things that plasmids can be used to do include:
- Produce large amounts of a protein so that scientists can purify and study it in a controlled setting. Read more:
- Produce proteins that glow so that scientists can track their location or quantity inside a cell
- Monitor the level of a chemical in a particular environment
- Produce enzymes that will make specific, controlled changes to an organism’s genome (genome engineering)
- Produce synthetic viruses that can be used in research or for therapeutics
Addgene has compiled various educational resources to facilitate plasmid use in the lab. Addgene’s Molecular Biology Reference includes information about molecular cloning, how to choose a plasmid vector, molecular biology tools and references, and how to maintain your plasmid stocks. The guide also contains multiple protocols and troubleshooting tips to make plasmid usage as simple and straightforward as possible.
If you have a question about a specific plasmid element that you would like answered or any topic suggestions for our Plasmids 101 series, please let us know in the comments.
Note:Marcy Patrick contributed to the writing of this article.

Topics: Plasmids 101, Plasmids





Leave a Comment