Every few months we highlight a subset of the new plasmids and viral preps in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need.

Here’s what you’ll find in this article:
- CRISPR-based editing in organoid
- Biosensors for detecting strigolactones
- The CRISPR Corner
- New ready-to-use viral preps
New toolkit for CRISPR-based genome editing in organoids
Organoids are three-dimensional cultures that mimic aspects of organ morphology and physiology in vitro and have emerged as a powerful model for understanding human physiology, development, and disease. In order to expand their model’s impact, organoid users have recently begun to develop CRISPR-based approaches for studying gene function in these systems. As a part of this effort, members of the Rawlins Lab have compiled a new toolkit for performing genetics studies in organoids. The Organoid EasyTag System is a straightforward workflow for doing gene targeting, reporter generation, controllable gene knockouts, and inducible gene inactivation or activation in an organoid model (Sun, et al. 2021).
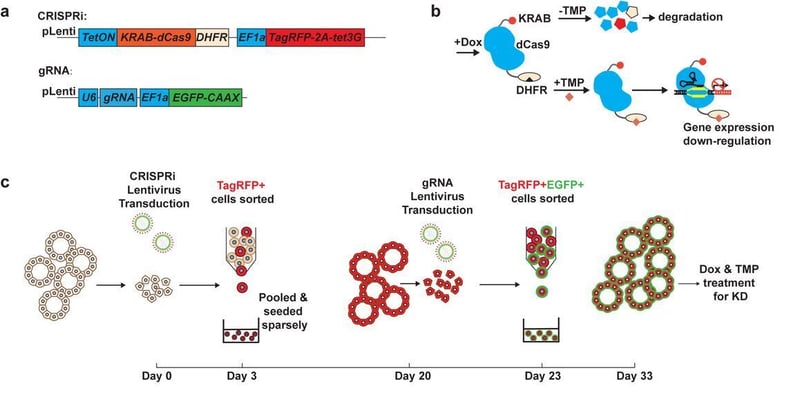 |
| Figure 1: Example schematic from the Organoid EasyTag system, showing the workflow for inducible CRISPR inhibition (CRISPRi). A) Lentiviral vector design. B) Strategy for leak-free transcriptional control. C) Iterative transduction workflow overview. Image from Sun et al. 2021. |
The Organoid EasyTag workflow combines thoughtfully designed plasmid components with Fluorescence Activated Cell Sorting (FACS), allowing for enrichment of successfully targeted cells that can be used to generate organoid cultures with a modification of interest. While the authors optimized the workflow for use in a fetal-lung derived organoid system, the approach is intended to be amenable to organoids derived from other tissue-types. By bringing together CRISPR-based approaches and organoid models, toolkits like this one provide new opportunities for investigating genetic mechanisms related to morphogenesis, cell fate specification, and human disease.
Sun, et al. bioRxiv. 2021. https://doi.org/10.1101/2020.05.04.076067
Novel fluorescent enzyme biosensors provide direct, specific and sensitive detection of the plant hormones strigolactones
Current methods used to detect strigolactones (SLs) are costly, non quantitative or require heavy optimization. Claudia Vickers’s lab has designed novel SL fluorescent biosensors by integrating a SL-insensitive fluorophore (LSSmOrange) as an internal control and a SL-sensitive fluorophore (circularly permuted green fluorescent protein or cpGFP) to the petunia SL enzymatic receptor DAD2. SLs induce a conformational change of the receptor which modulates the fluorescence intensity of the cpGFP without altering that of the internal control (Fig 1). Because this method relies on the fluorescence ratio between the 2 fluorophores, rather than on the fluorescence intensity, it provides a direct and quantitative detection of SLs. The authors also showed that the receptor-based fluorescent biosensor has a high sensitivity (50 nM-500nM range) and a high specificity for SLs in vitro, and they further demonstrated that when tested in tobacco protoplasts, the enzymatic biosensor retains its catalytic activity.
As SLs are also known to trigger seed germination of parasitic plants, this approach was also used to build and validate a ratiometric fluorescent biosensor based on the parasitic Striga hermonthica SL receptor HTL7. This novel type of biosensors could be used in high throughput screening for agrochemical compounds that have the potential to regulate the parasitization of cereal crops. As a result, agricultural applications of these novel biosensors could have a major economic impact.
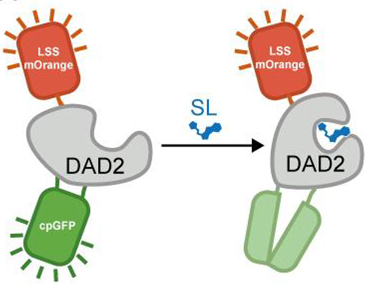 |
| Design and mechanism of action of the receptor-based SL fluorescent biosensor. Image from Chesterfield et al., 2020 |
Chesterfield et al., ACS Synthetic Biology 2020. https://pubs.acs.org/doi/10.1021/acssynbio.0c00192
 The CRISPR corner
The CRISPR corner
New CRISPR plasmids are always being added to the repository. To find all of the CRISPR plasmids available from Addgene, head over to our CRISPR Plasmids and Resources page. Here are some highlights from the past couple months:
- A new lentiviral CRISPR library from Alec Kimmelman targets mouse metabolic genes with ~6 gRNAs per gene.
- CRISPR-Act3.0, developed by Yiping Qi's lab, allows simultaneous gene activation in plants.
- FnCas12a, unlike other Cas12a nucleases identified, functions at temperatures up to 43 °C and can be used for genome editing in moderate thermophiles.
- These evolved SpCas9 variants have improved activity on NAG PAMS and reduced activity on NGG PAMs relative to wild type.
- A new genome-wide CRISPR knockout library from Xiaole Shirley Liu's lab targets all mouse genes with 10 gRNAs per gene. The guides are optimized to maximize on-target cleavage and minimize off-target effects.
 New from the viral service
New from the viral service
By Jason Nasse
We regularly add new viral aliquots from our plasmid collection to provide ready-to-use viral preps. Here are some of the new viral preps from recent months:
- Our selection of GCaMP8 AAV preps has expanded. Check out the new selection of ready-to-use AAV9 and AAVrg vectors.
- New red-shifted INTRSECT 2.0 calcium sensor available in AAV8! Expand your imaging experiments with the Cre-OFF/FLP-ON vector 137128-AAV8: Ef1a-Coff/Fon-sRGECO
Topics: Hot Plasmids, Plasmids








Leave a Comment