By Balázs Csoma, Hungarian Research Network
CRISPR nucleases are remarkably precise molecular tools for cutting DNA. But “remarkably precise” does not mean “perfectly specific.” In reality, CRISPR nucleases occasionally make mistakes and cleave DNA sequences that only resemble the intended target. These unintended cuts cause the dreaded off-target effects that can wreck experiments and, in clinical settings, raise serious safety concerns.
Off-target effects are a matter of sensitivity. Therefore, the key question is not whether off-target effects exist, but what the required level of specificity is for a given application — and how we can achieve that efficiently.
Here, I outline a practical strategy on how to minimize off-target activity, based on a deeper understanding of the underlying molecular mechanisms involved, without adding any unnecessary hassle.
Increased-fidelity SpCas9 variants: Useful, but not a silver bullet
To address off-target effects, many increased-fidelity SpCas9 nuclease variants (IFNs) have been engineered. These variants are designed to maintain on-target activity while reducing cleavage at mismatched sites.
In practice, however:
- Increased fidelity to a given sequence often comes at the sacrifice of some or all on-target activity.
- The balance between on-target and off-target cutting varies tremendously between targets.
- On-target prediction tools do exist for a few IFNs, but there is currently no reliable way to predict the off-target effects of a specific IFN.
You may think that off-target cleavage is determined solely by the number of or the position of mismatches between the spacer and a genomic site. However, the targeted DNA sequences (or spacers) themselves have an inherent differential propensity to cleavage, and this often-overlooked factor has a major impact on whether a mismatched site will be cut or not. This explains why ‘one-size-fits-all’ solutions to eliminate off-target effects rarely work.
Fortunately, IFNs can be arranged along a spectrum of decreasing activity/increasing fidelity (activity rank) and likewise, target sites themselves form a spectrum based on how easily they can be cleaved (cleavability rank). By combining these two trends, the cleavage rule can be established:
If a nuclease variant cleaves a target, all variants with higher activity will cleave it too.
This leads to the powerful concept of using target-matched nucleases.
The target-matched nuclease strategy
The idea is very simple: choose the least active SpCas9 variant that still efficiently cuts your on-target site.
If a nuclease is just active enough to cleave the intended target, it does not have enough activity to cut off-target sites.
_min.png?width=565&height=345&name=fig1(1)_min.png) |
|
Figure 1: Rectangles represent the on-target activity of a given nuclease (indicated at the top of each column) on a given target (indicated at the beginning of each row). Targets that are “easily” cleaved are cut efficiently by most variants (rows with mostly green rectangles). However, only the least active variant which still retains on-target activity (highlighted by a bold frame) provides off-target–free editing. In contrast, “hard-to-cleave” targets are not edited by low-activity variants (red rectangles). However, among the higher-activity nucleases, the weakest variant that is still active can achieve off-target–free editing for this type of targets.
|
The CRISPRecise kit available at Addgene includes 17 IFNs, covering a spectrum of activity levels fine enough to provide a target-matched variant for virtually any target (Kulcsár et al., 2023).
The challenge? There is no prediction tool that can reliably identify this optimal variant in advance, so minimal experimental testing is required.
So how can you do this? Testing all IFNs is completely unnecessary to identify the target-matched variant for a given application. Here, I give you a step-by-step guide that can be scaled to how much specificity you need for your particular experiment.
Step 1: Choose your target well
If multiple target sites are available:
- Use on-target prediction tools to increase the likelihood of high efficiency cleavage. In our experience, DeepWT provides the most reliable prediction (Wang et al., 2019).
- Choose targets with fewer close genomic matches, just in case.
You may not get a perfect prediction, but you get a great preliminary risk reduction at a minimal cost.
Step 2: Rough screening of nuclease variants by next-generation sequencing (NGS)
After cloning your sgRNA:
- Check on-target activity of WT SpCas9 and a small set of strategically selected IFNs on your samples by NGS. We suggest using e-plus, Blackjack-HF1, and Blackjack-HypaR, as these variants are located in different ranges of the activity spectrum (Figure 2).
- Identify the least active variant that still shows the right amount of on-target editing.
This variant will typically show substantially reduced off-target activity, and the whole screening required only minimal extra effort (Figure 3). This first-pass screen also quickly tells you which variants to test if you want further refinement.
 |
|
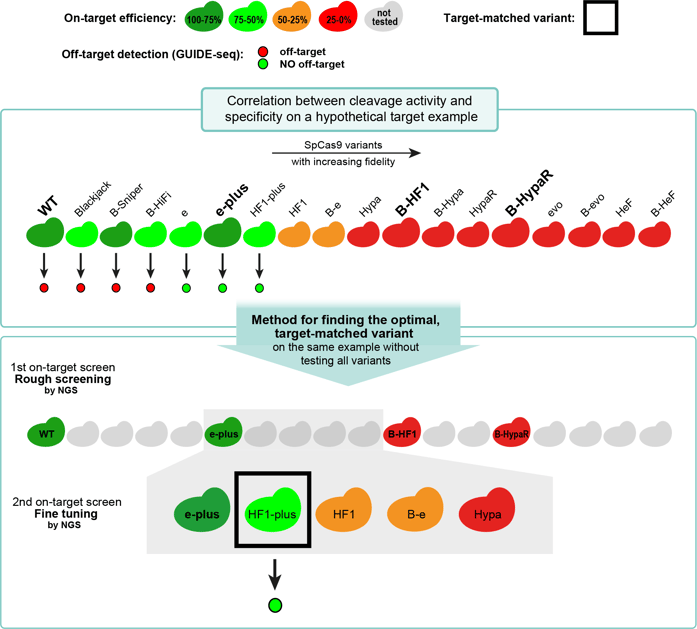
Figure 2: Schematic representation of the two-step screening method used on a hypothetical target example. The top panel shows the on- and off-target activity of a set of IFNs with increasing fidelity on a hypothetical target example. The second panel shows the screening method, which identifies the optimal variant for the target without having to test all the IFNs. In the first step, a rough on-target screen is performed, where we test WT SpCas9 and three selected IFNs that divide the target ranking range into four approximately proportional sections. The second step is a fine-tuning on-target screen, that involves the not-yet-used variants with lower activity than the lowest-ranking active (green) variant but with higher activity than the highest-ranking inactive (red) variant from the first screen. This second screen thus identifies the target-matched variant (active variants with the lowest activity, and thus highest fidelity, in the full set). Adapted from Kulcsár et al., 2023 under a CC BY 4.0 license.
|
 Pro tip! If on-target efficiency of WT SpCas9 is below ~5–10%, consider selecting a different target. Remember, if WT SpCas9 does not cleave the target efficiently, neither will the IFNs.
Pro tip! If on-target efficiency of WT SpCas9 is below ~5–10%, consider selecting a different target. Remember, if WT SpCas9 does not cleave the target efficiently, neither will the IFNs.
Step 3 (optional): Fine tuning for when zero off-target cleavage is your goal
For applications where the highest achievable specificity is paramount:
- Pick and test some more variants from the CRISPRecise set located between the last active and the first inactive one from step 2. (In the example in Figure 3, the least active still working variant was B-HypaR, thus variants with lower activity — evo, B-evo, and B-HeF — should be tested in this screen.)
- Identify the weakest variant that still remains active.
 |
|
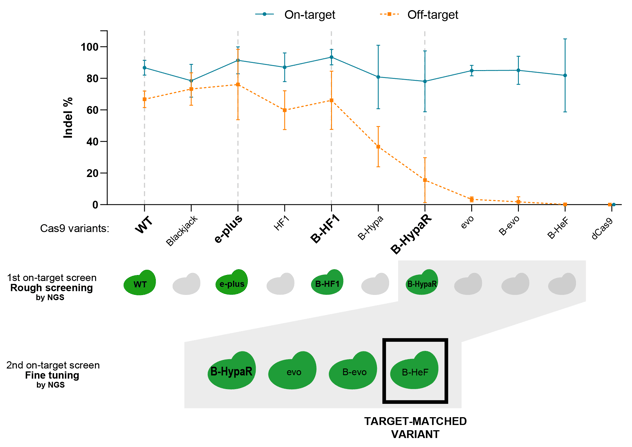
Figure 3: Activity of 9 IFNs with a sgRNA targeting a perfectly matching genomic target (blue line) and a single mismatch-containing genomic target (orange line) are shown. Under the graph the hypothetical results of the first and the second screen are shown.
|
This second round of screening will generally provide off-target-free editing. (Note: this conclusion was reached based on deductive reasoning rooted in the nature of IFNs and targets (i.e., the cleavage rule), and assumptions were validated on some critically selected challenging targets by GUIDE-seq assessment (Kulcsár et al., 2023), as, of course, testing every IFN-target combination would be impossible.)
According to the cleavage rule, virtually all targets — with rare exceptions — can be cleaved without off-target effects.
Step 4: Validation made easy
- Grow clonal populations from the cells edited successfully by the target-matched variant.
- Screen and expand clones carrying the desired on-target modification.
Using this approach, highly specific genome editing can generally be achieved without using genome-wide off-target assays, which are time-consuming and expensive.
A practical sgRNA design note
When expressing gRNAs from a U6 promoter, remember that transcription typically requires a purine base (A or G) at the 5′ end. If your spacer does not naturally begin with a purine, extend the gRNA with a 5′ G (21G design).
Be aware that only certain IFNs, typically those containing Blackjack mutations, retain sufficient activity with 5′ modified sgRNAs (Kulcsár et al., 2020).
 |
|
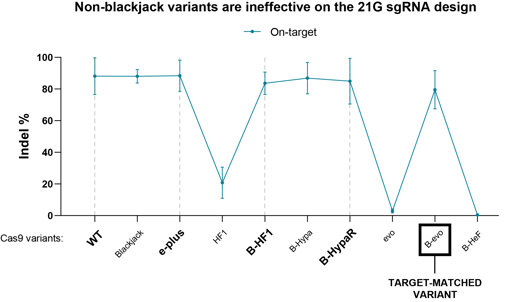
Figure 4: On-target activity of 9 IFNs with a 21G-sgRNA on matching genomic target. Data were provided courtesy of Zoltán Ligeti. The B-evo variant is the target-matched variant here. The HF1 and evo variants that do not contain the Blackjack mutations (otherwise indicated in the variants’ name as B) have diminished activity with a 21G-sgRNA and thus should be excluded from the screen when 21G-sgRNAs are tested.
|
Take-home message
Off-target cleavage is not a black or white kind of situation. You can absolutely achieve application-appropriate specificity while preserving activity without excessive additional workload by combining:
- Careful target selection,
- Informed selection of increased-fidelity SpCas9 variants,
- And minimal but purposeful experimental screening.
Sometimes to reach exactly what you want, cutting just enough is the best approach.
This post was written by Balázs Csoma, a CRISPR biologist working on improved genome editing at HUN-REN RCNS. He is passionate about sharing science and enjoys troubleshooting challenging plasmid cloning problems.
References and Resources
References
Kulcsár, P. I., Tálas, A., Ligeti, Z., Tóth, E., Rakvács, Z., Bartos, Z., Krausz, S. L., Welker, Á., Végi, V. L., Huszár, K., & Welker, E. (2023). A cleavage rule for selection of increased-fidelity SpCas9 variants with high efficiency and no detectable off-targets. Nature Communications, 14(1), 5746. https://doi.org/10.1038/s41467-023-41393-5
Kulcsár, P. I., Tálas, A., Tóth, E., Nyeste, A., Ligeti, Z., Welker, Z., & Welker, E. (2020). Blackjack mutations improve the on-target activities of increased fidelity variants of SpCas9 with 5′G-extended sgRNAs. Nature Communications, 11(1), 1223. https://doi.org/10.1038/s41467-020-15021-5
Tálas, A., Huszár, K., Kulcsár, P. I., Varga, J. K., Varga, É., Tóth, E., Welker, Z., Erdős, G., Pach, P. F., Welker, Á., Györgypál, Z., Tusnády, G. E., & Welker, E. (2021). A method for characterizing Cas9 variants via a one-million target sequence library of self-targeting sgRNAs. Nucleic Acids Research, 49(6), e31. https://doi.org/10.1093/nar/gkaa1220
Wang, D., Zhang, C., Wang, B., Li, B., Wang, Q., Liu, D., Wang, H., Zhou, Y., Shi, L., Lan, F., & Wang, Y. (2019). Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning. Nature Communications, 10, 4284. https://doi.org/10.1038/s41467-019-12281-8
Additional resources on the Addgene blog
Resources on addgene.org
Topics:
CRISPR,
Cas Proteins,
CRISPR Protocols and Tips
_min.png?width=565&height=345&name=fig1(1)_min.png)








Leave a Comment