This post was contributed by Kyle Hooper at Promega.
Researchers have been sharing plasmids ever since there were plasmids to share. Back when I was in the lab, if you read a paper and saw an interesting construct you wished to use, you could either make it yourself or you could “clone by phone”. One of my professors was excellent at phone cloning with labs around the world and had specific strategies and tactics for getting the plasmids he wanted. Addgene makes it so much easier to share your constructs from lab to lab. Promega supports the Addgene mission statement: Accelerate research and discovery by improving access to useful research materials and information. Many of our technology platforms like HaloTag® Fusion Protein, codon-optimized Firefly luciferase genes (e.g., luc2), and NanoLuc® Luciferase are available from the repository. We encourage people to go to Addgene to get new innovative tools. After all, isn’t science better when we share?
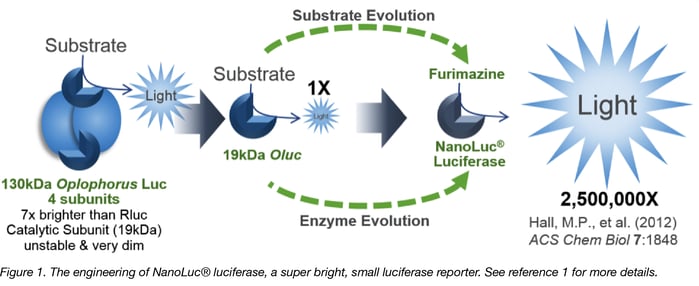
NanoLuc® Luciferase-based tools are fantastic examples of technologies that can accelerate your research. The creation of NanoLuc® Luciferase from a rather dull 19kDa Oplophorus luciferase by Promega’s Advanced Technologies group as well as evolution of the coelenterzine substrate to the more stable yet brighter furimazine by Promega’s Chemistry group is a good story (1). From a deep sea shrimp to a compact powerhouse of bioluminescence, NLuc is 100-fold brighter than our more common luciferases like firefly (FLuc) and Renilla (RLuc) luciferase. This is important not so much for how bright you can make a reaction but for how sensitive you can make a reaction. NLuc requires 100-fold less protein to produce the same amount of light from an Fluc or RLuc reaction. In fact, NLuc is bright enough to detect endogenous tagged genes generated through CRISPR/Cas9 knock-in. Finally, NLuc is very inviting for endogenous protein tagging as it is only 19 kDa. An example is the CRISPaint-NLuc construct (Plasmid #67178) for use in the system outlined in Schmid-Burgk, J.L., et al (2).
In this post, I’ll cover two great applications of the NLuc technology. The first couples NLuc to fluorescent proteins to make better reporters and the second couples NLuc to fluorescent proteins to make better biosensors.
NanoLuc®-fluorescent protein fusions for imaging
Fluorescent proteins are fantastic imaging tools but their use is limited in vivo because they must be excited by external light sources. These external sources can generate autofluorescence and have limited penetration due to absorption by tissues. To avoid absorbance of heme-containing proteins in whole animals, light at or above 600 nm wavelengths should be used.
Researchers often use bioluminescence to overcome these issues as traditional luciferases emit at near optimal wavelengths (Fluc emits at 610 nm) and do not require external light sources. Unfortunately, detecting traditional luciferases’ dim light emissions requires long acquisition times that may be impractical or infeasible for certain experimental setups.
The brightness of NLuc has caught the attention of researchers wanting to perform whole-animal imaging, however NLuc’s emission peak is at 460 nm thus making it non-optimal for in vivo imaging.
The solution is to couple Nluc with a red-shifted fluor. Here, NLuc internally excites the fluor through a process known as bioluminescent resonance energy transfer (BRET). The fluor then emits light at a tissue-penetrating wavelength above 600 nm.
LumiFluors
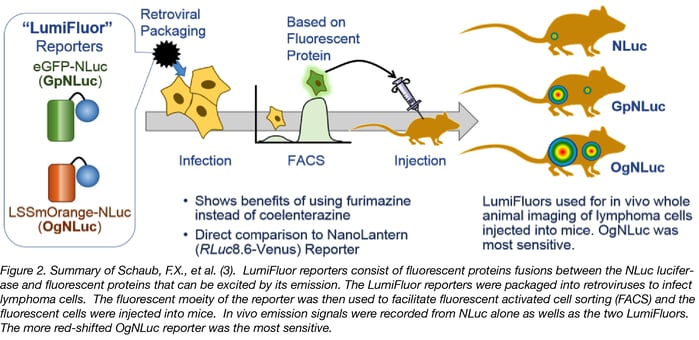
Schaub, F.X., et al. (3) designed two imaging probes. One consists of NLuc fused to enhanced green fluorescent protein (GpNluc) and the other consists of NLuc fused to the more penetrative orange fluorescent protein fusion (OgNLuc). Both fluorescent proteins can be excited by Nluc emission and the lab has dubbed these fusions LumiFluor Reporters. The LumiFluor reporter constructs were designed to be packaged into a retrovirus for infection of host cells. The fluorescent moiety of the BRET reporter was used to sort successfully infected/expressing cells by flow cytometry. The isolated cells were injected into mice and imaged after i.p. or i.v. administration of the furimazine substrate for NanoLuc luciferase. Both GpNLuc and OgNLuc BRET reporters produced bright signals without exogenous excitation and with OgNLuc being the brightest since it had an emission near 600 nm – a clear demonstration of why a BRET reporter can improve deep tissue imaging. The constructs for GpNLuc and OgNLuc are available through Addgene (Cat.# 70185 and 70186, respectively).
Antares
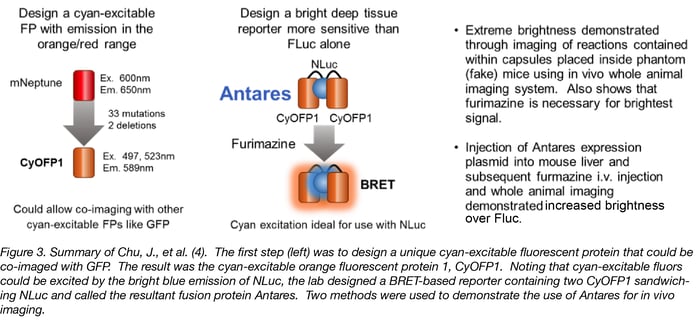
Chu, J., et al. (4), likewise, wanted to produce a BRET probe for imaging. These authors set out to design a better fluorescent protein that could be co-imaged with GFP using the same wavelength for excitation. The result was the cyan-excitable orange fluorescent protein, CyOFP1. CyOFP1 was molecularly evolved from mNeptune2 through 33 mutations and 2 deletions. CyOFP1 has a higher quantum yield, brightness and improved maturation time compared to similar orange fluorescent proteins. In their work, the authors noted how CyOFP1 could be excited by NanoLuc® luciferase (NLuc) and set about to create an in vivo imaging reporter by linking two CyOFP1 molecules to NLuc (CyOFP1-NLuc-CyOFP1; Antares). Direct injection of plasmids for Antares or firefly luciferase into mouse livers followed by i.v. injection of furimazine or luciferin 24 hours later demonstrated a more than 2-fold brighter signal from Antares as compared to Fluc in this model. The construct for Antares is available through Addgene (Cat. # 74279).
Enhanced Nano-Lanterns
Suzuki, K., et al. (5) imaging multiple proteins at the same time in live cells using fluorescent proteins (FPs) can be problematic because excitation light can cause phototoxicity, autofluorescence, and perturb biological phenomena. The technique of coupling FPs with Luciferases can overcome these problems but has been hampered by dim signals from the coupled luciferase. This strategy was first applied using RLuc8, an engineered Renilla luciferase, to produce a series of FP-RLuc8 fusions for multicolor imaging. In all of these fusions coelenterazine intiates the fluorescent signal and these FP-RLuc8 fusions were dubbed Nano-Lanterns.
RLuc8 is 5.5-fold brighter than RLuc but NLuc is 30-fold brighter than RLuc8 (6). Susuki, K, et al. therefore replaced the RLuc8 moieties in their Nano-Lanterns with NLuc and called their improved creations Enhanced Nano-Lanterns. These great imaging tools generate cyan, yellow, green, orange and red fluorescence. The paper reports a wide variety of applications of the Enhanced Nano-Lanterns including monitoring multiple cellular events such as the dynamics of subcellular structures and gene expression. You must see the movies demonstrating Ca2+ dynamics in cardiomyocytes.
Enhanced Nano-Lantern constructs are available through Addgene: The Turquoise (CeNL; Plasmid #85199), Yellow (YeNL; Plasmid #85201), Orange (OeNL; plasmid #85202) and Red (ReNL; plasmid #85203) NLuc fusions. The Nagai lab has additionally deposited more than 25 fusions of the Enhanced Nano-Lanterns to other cellular proteins.
BRET-based biosensors utilizing NanoLuc® luciferase
Many intracellular sensors of events like calcium release have been made using fluorescent resonance energy transfer (FRET) probes. These probes generally consist of a responsive protein (e.g. calcium binding protein) fused to two fluorescent proteins. When the responsive protein binds to calcium it changes conformation and brings the two fluorescent proteins together thereby increasing their FRET signal. FRET is a non-radiative transfer of energy from one fluorescent protein to a compatible fluorescent protein. FRET can be measured by exciting the most blue-shifted fluorescent protein (donor) and measuring the resultant emission from the red-shifted fluorescent protein (acceptor) in the pair. The higher the emission from the acceptor, the more powerful the FRET signal. In well-designed FRET-based biosensors, the intensity of the FRET signal can be correlated to specific concentrations of biological molecules.
FRET sensors face challenges of photobleaching, autofluorescence, and, in the case of exciting cyan-excitable donors, phototoxicity. Another challenge to using FRET sensors comes when employing optogenetic regulators to initiate the event being monitored. Optogenetic regulators respond to specific wavelengths of light to initiate signaling. When FRET donor excitation wavelengths overlap with optogenetic initiation wavelengths, excitation of the optogenetic regulator leads to spurious activation of the FRET sensor.
Researchers have sought to alleviate many of these challenges by exchanging the fluorescent donor for a bioluminescent donor, making BRET probes. Here the bioluminescent donor is unaffected by activation of other fluorescent probes but can still transfer energy to the fluorescent protein acceptor. NLuc is an excellent BRET donor due to its bright luminescence.
Intracellular Calcium Sensor
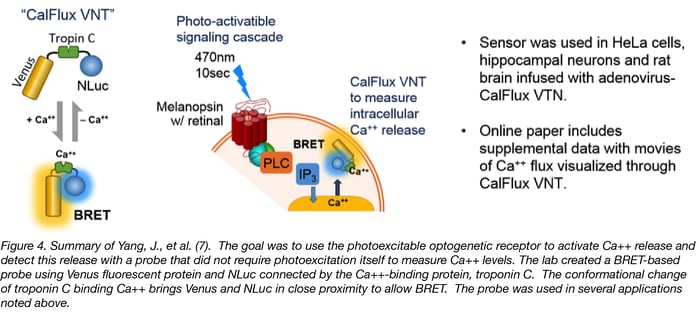
Yang, J. et al. (7) wished to trigger intracellular Ca++ release with an optogenetic receptor, melanopsin, requiring brief exposure to 470nm blue light. To avoid overlap with this optogenetic tool, they needed to create a calcium sensor with the following characteristics:
- No photoexcitation
- High Ion specificity
- High sensitivity
- Easy expression
- Gives ratiometric data
The result was a fusion protein called CalFlux VNT. CalFlux VNT is composed of Venus fluorescent protein, the Troponin C Ca++ binding domain, and NanoLuc® Luciferase. CalFlux VNT responds to changes in intracellular calcium and is useful for direct imaging of events in cells and brain explants. This construct is available through Addgene (Plasmid #83926).
Live-Cell Voltage Sensor
Ingaki and colleagues (8) were interested in using photoactivatable, optogenetic accuators to depolarize or hyperpolarize membranes. Traditional FRET reporters could not be used as the excitation wavelengths needed would also activate the optogenetic accuators. The researchers took a FRET reporter and adapted it into a BRET reporter by using NanoLuc® Luciferase to excite the Venus fluorescent protein in conjunction with a membrane-inserted voltage sensing domain (LOTUS V; Luminescent Optical Tool for Universal Sensing of Voltage). With this sensor, membrane depolarization increases the BRET signal whereas hyperpolarization decreases the signal. The authors additionally demonstrate that LOTUS V works in many model systems. This construct is available through Addgene (Plasmid # 87127).
Pairing NanoLuc® luciferase with fluorescent proteins has allowed researchers to make innovative tools not possible with other luciferases. In these tools, NanoLuc-fluorescent protein fusion shifts NLuc’s bright blue luminescence to wavelenghts more friendly for cellular and in vivo imaging. NLuc additonally allows researchers to convert FRET biosensors into more versatile BRET biosensors that can even be paired with optogenetic regulators. Many of these tools are available from Addgene and we're excited to see how you utilize them in your group!
Many thanks to our guest blogger Kyle Hooper.
 A former Technical Services Scientist, Kyle Hooper, PhD, has also worked with R&D for product development and now specializes in supporting Promega cellular analysis products at Promega.
A former Technical Services Scientist, Kyle Hooper, PhD, has also worked with R&D for product development and now specializes in supporting Promega cellular analysis products at Promega.
References
1. Hall, M.P., et al. (2012) Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazopyrazinone substrate. ACS Chem Biol 7, 1848-57. PubMed PMID: 22894855. PubMed Central PMCID: PMC3501149.
2. Schmid, J.L., et al. (2016) CRISPaint allows modular base-specific gene tagging using a ligase-4-dependent mechanism. Nature Comm. 7, 12338. PubMed PMID: 27465542. PubMed Central PMCID: PMC4974478.
3. Schaub, F.X., et al. (2015) Fluorophore-NanoLuc BRET reporters enable sensitive in vivo optical imaging and flow cytometry for monitoring tumorigenesis. Cancer Res. 75, 5023-33. PubMed PMID: 26424696. PubMed Central PMCID: PMC4668208.
4. Chu, J., et al. (2016) A bright cyan-excitable orange fluorescent protein facilitates dual-emission microscopy and enhances bioluminescence imaging in vivo.Nature Biotechnol. 34, 760-7. PubMed PMID: 27240196. PubMed Central PMCID: PMC4942401.
5. Susuki, K., et al. (2016) Five colour variants of bright luminescent protein for real-time multicolour bioimaging. Nature Comm. 7, 13718. PubMed PMID: 27966527. PubMed Central PMCID: PMC5171807.
6. Machleidt, T., et al. (2015) NanoBRET-A novel BRET platform for the analysis of protein-protein interactions. ACS Chem. Biol. 10, 1797-804. PubMed PMID: 26006698.
7.Yang, J., et al. (2016) Coupling optogenetic stimulation with NanoLuc-based luminescence (BRET) Ca++ sensing. Nature Comm. 7, 13268. PubMed PMID: 27786307. PubMed Central PMCID: PMC5476805.
8. Inagaki, S., et al. (2017) Genetically encoded bioluminescent voltage indicator for multi-purpose use in wide range of bioimaging. Reports 7, 42398. PubMed PMID: 28205521. PubMed Central PMCID: PMC5322354.
Additional Resources on the Addgene Blog
- Learn more about FRET
- Tips for Using FRET in Your Experiments
- Learn more about Fluorescent Protein Biosensors
Additional Resources on Addgene.org
- Browse Luciferase Plasmids
- Browse the Fluorescent Protein Collection
- Find Fluorescent Protein Biosensors
Topics: Fluorescent Proteins, FRET, Luminescence







Leave a Comment