This post was contributed by guest blogger Clare O'Connor an Associate Professor at Boston College.
National reports stress the importance of providing authentic research experiences to undergraduate students (1, 2), but educators face significant challenges in designing suitable projects. In the O'Connor lab, we recognized that genome sequencing projects were generating huge amounts of data that could provide the basis for student projects in introductory labs. Genome projects use computational methods to identify genes by their similarities to genes in other species, but these studies generally leave questions about gene function wide open. What if two seemingly similar proteins have acquired divergent functions due to mutations accumulated over time? Undergraduates can help to answer this question!
The Pathways Over Time project
 The Pathways over Time project provides a framework for these undergraduate research questions. The idea is straightforward. Instructors first select a multi-gene pathway to study and a reference organism where deletion strains are available. Then, staff or students clone similar sequences from a less-well studied organism into overexpression plasmids. Students next transform deletion strains from the reference organisms with the overexpression plasmids. Finally, students determine if the presence of the plasmid corrects any known phenotypic abnormalities due to the deletion (i.e. the plasmid complements the deletion). If the phenotype is corrected, students can be relatively confident that the gene from the test species has a similar function to the gene in the reference species.
The Pathways over Time project provides a framework for these undergraduate research questions. The idea is straightforward. Instructors first select a multi-gene pathway to study and a reference organism where deletion strains are available. Then, staff or students clone similar sequences from a less-well studied organism into overexpression plasmids. Students next transform deletion strains from the reference organisms with the overexpression plasmids. Finally, students determine if the presence of the plasmid corrects any known phenotypic abnormalities due to the deletion (i.e. the plasmid complements the deletion). If the phenotype is corrected, students can be relatively confident that the gene from the test species has a similar function to the gene in the reference species.
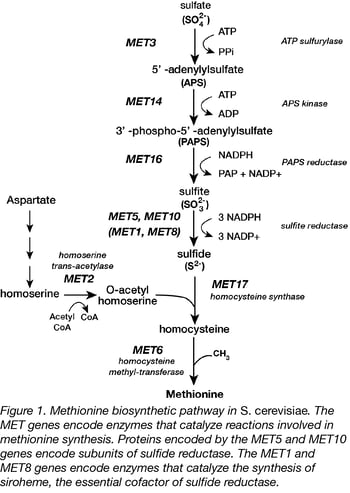
We have been using yeast models (3, 4) to answer questions about the evolution of the genes involved in methionine synthesis (5). The budding yeast, Saccharomyces. cerevisiae, makes a great reference organism for the project because it is non-pathogenic, well-studied and inexpensive. In addition, a complete set of well-annotated deletion strains is available (6). Methionine synthesis in S. cerevisiae (Figure 1) occurs through a well-characterized pathway with multiple enzymatic steps (5). In their experiments, students test whether or not proteins from the fission yeast, Schizosaccharomyces pombe, have the same function as their counterparts in S. cerevisiae. The two yeasts are thought to have diverged from a common ancestor ~350 million years ago. Over time, the two genomes have become very different due to a whole genome duplication and extensive rearrangements in the S. cerevisiae lineage (7).
In our class research project, students transform S. cerevisiae met deletion strains with overexpression plasmids carrying MET/Met genes from either S. pombe or S. cerevisiae. Students then determine if expression of the plasmid-encoded MET gene allows the transformed strain to grow in the absence of methionine (i.e. the plasmid complements the deletion).
Plasmid features enable multiple student experiments
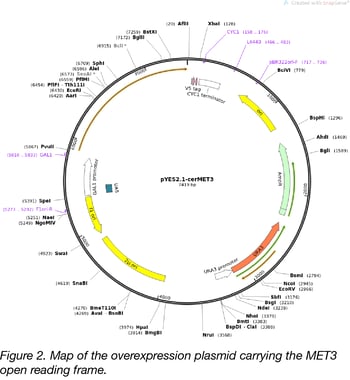 We used TOPO cloning to insert the coding sequences for ten different MET/Met genes (Figure 1) from the two yeast species into the pYES2.1/V5-His yeast overexpression vector. This plasmid contains origins of replication and selectable markers that allow students to propagate the plasmids in either E. coli (ori and ampR) or S. cerevisiae (2µm ori and URA3). The presence of the URA3 gene allows students to select yeast that have been successfully transformed with the plasmids, because the S. cerevisiae deletion strains are ∆ura3 mutants and cannot grow without uracil supplementation (6). Expression of MET/Met genes from the pYES2.1 plasmid is controlled by the GAL1 promoter. The cloning procedure adds both V5 and His6 epitope tags to the C-termini of Met proteins, which can be used to confirm protein expression by western blotting in other experiments (Figure 2).
We used TOPO cloning to insert the coding sequences for ten different MET/Met genes (Figure 1) from the two yeast species into the pYES2.1/V5-His yeast overexpression vector. This plasmid contains origins of replication and selectable markers that allow students to propagate the plasmids in either E. coli (ori and ampR) or S. cerevisiae (2µm ori and URA3). The presence of the URA3 gene allows students to select yeast that have been successfully transformed with the plasmids, because the S. cerevisiae deletion strains are ∆ura3 mutants and cannot grow without uracil supplementation (6). Expression of MET/Met genes from the pYES2.1 plasmid is controlled by the GAL1 promoter. The cloning procedure adds both V5 and His6 epitope tags to the C-termini of Met proteins, which can be used to confirm protein expression by western blotting in other experiments (Figure 2).
Complementation reveals conservation of function
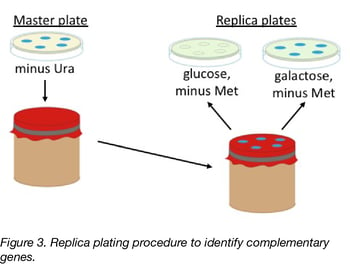 Students have used these plasmids, which have been deposited with Addgene, to successfully analyze Met protein complementation. In the procedure diagrammed in Figure 3, students first transform the S. cerevisiae met deletion strains and then isolate transformed strains on media that lack uracil, but contain methionine. (This allows cells carrying nonfunctional as well as functional MET genes to grow.) Students then use replica plating to determine if expression of MET/Met genes from the plasmids allows the transformed strains to grow in the absence of methionine. Since the GAL1 promoter controls MET/Met gene expression, complementation should only occur when galactose replaces glucose as the carbon source. Student results have shown that most, but not all, S. pombe Met genes complement the corresponding S. cerevisiae met deficiencies (manuscript in preparation), indicating that Met protein functions have been largely conserved since the species diverged.
Students have used these plasmids, which have been deposited with Addgene, to successfully analyze Met protein complementation. In the procedure diagrammed in Figure 3, students first transform the S. cerevisiae met deletion strains and then isolate transformed strains on media that lack uracil, but contain methionine. (This allows cells carrying nonfunctional as well as functional MET genes to grow.) Students then use replica plating to determine if expression of MET/Met genes from the plasmids allows the transformed strains to grow in the absence of methionine. Since the GAL1 promoter controls MET/Met gene expression, complementation should only occur when galactose replaces glucose as the carbon source. Student results have shown that most, but not all, S. pombe Met genes complement the corresponding S. cerevisiae met deficiencies (manuscript in preparation), indicating that Met protein functions have been largely conserved since the species diverged.
Tips on using the Pathways Over Time plasmids
The plasmids are used as part of a semester-long research course that is part of the biology core curriculum at Boston College. Students in the course work in teams of three in sections of 15 students. Within each section, teams are assigned different groups of three genes selected from different parts of the Met pathway (Fig. 1). In addition to the complementation experiments that are used to answer our research question, the plasmids are also used to teach students about restriction mapping and western blotting. Students who participate in the course show significant gains in both content knowledge and research skills (8). A complete set of student and instructor resources is available at our Investigations in Molecular Cell Biology Website.
Many thanks to our guest blogger, Clare O'Connor!
 Clare O'Connor (left with Lab Director Doug Warner on the right) has taught courses in biochemistry, molecular cell biology and genetics at Boston College. She is particularly interested in strategies to incorporate research into undergraduate courses and in communicating science to the public. You can find out more about Clare at the Biology Dept. site.
Clare O'Connor (left with Lab Director Doug Warner on the right) has taught courses in biochemistry, molecular cell biology and genetics at Boston College. She is particularly interested in strategies to incorporate research into undergraduate courses and in communicating science to the public. You can find out more about Clare at the Biology Dept. site.
References
1. American Association for the Advancement of Science. (2011). Vision and Change in Undergraduate Biology Education: A Call to Action, Washington, DC.
2. President's Council of Advisors on Science and Technology [PCAST] (2012). Engage to Excel: Producing One Million Additional College Graduates with Degrees in Science, Technology, Engineering and Mathematics, Washington, DC: US Government Office of Science and Technology.
3. Duina, A. A., Miller, M.E., & Keeney, J.B. Budding yeast for budding geneticists: A primer on the Saccharomyces cerevisiae model system. Genetics 197, 33-48 (2014). PubMed PMID: 24807111. PubMed Central PMCID: PMC4012490.
4. Hoffman, C.S., Wood, V., & Fantes, P.A. An ancient yeast for young geneticists: A primer on the Schizosaccharomyces pombe model system. Genetics 201, 402-423. (2015). PubMed PMID: 26447128. PubMed Central PMCID PMC4596657.
5. Thomas, D., & Surdin-Kerjan, Y. Metabolism of sulfur amino acids in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 61, 503-532. (1997). PubMed PMID: 9409150. PubMed Central PMCID PMC232622.
6. Winzeler, D.A., Shoemaker, D.D., Astromoff, A., Liang, H., Anderson, K., Andre, B., Bnangham, R., Benito, R., Boeke, J.D., Bussey, H. et al. Functional characterization of the Saccharomyces cerevisiae genome by gene deletion and parallel analysis. Science 285, 901-906. (1999). PubMed PMID: 10436161.
7. Dujon, B. Yeast evolutionary genomics. Nat Rev Genet 11, 512-524. (2010). PubMed PMID: 20559329.
8. Reeves, T.D., Warner, D.M., Ludlow, L.H., & O'Connor, C.M. Pathways over time: Functional genomics research in an introductory laboratory course. CBE-Life Sci. Educ. 17, ar1. (2018). PMID: 29326101.
Additional Resources on the Addgene Blog
- Learn about Synthetic Biology with BioBuilder
- Use Addgene Videos to Learn about Basic Lab Techniques
- High School Students Explore Careers at Addgene
Resources on Addgene.org
- Find Yeast Expression Vectors
- Check Out Our Molecular Biology Reference Pages
- Learn Simple Lab Techniques in Our Protocol Videos
Topics: Other Plasmid Tools, Plasmids





Leave a Comment