 In complex metazoans, rapid cell division and large scale cell mobility are essential processes during embryonic development. These are required for a growing organism to make the complicated transition from a clump of cells to a fully differentiated body. In contrast, these dynamic processes are largely absent in adult organisms, where tissues structures are more stable and local movements predominate (e.g. a basal progenitor cell migrating to the epithelium). At this stage, only cells from the immune system show wide scale mobility with movement from the bone marrow and other lymphoid organs to specific tissues where they can scan for any signs of danger. In this post we’ll focus on how fluorescent proteins can and have been used to monitor cellular movements in the immune system. The techniques used here could be adapted to studying other systems in which there is large scale cellular movement throughout an organism.
In complex metazoans, rapid cell division and large scale cell mobility are essential processes during embryonic development. These are required for a growing organism to make the complicated transition from a clump of cells to a fully differentiated body. In contrast, these dynamic processes are largely absent in adult organisms, where tissues structures are more stable and local movements predominate (e.g. a basal progenitor cell migrating to the epithelium). At this stage, only cells from the immune system show wide scale mobility with movement from the bone marrow and other lymphoid organs to specific tissues where they can scan for any signs of danger. In this post we’ll focus on how fluorescent proteins can and have been used to monitor cellular movements in the immune system. The techniques used here could be adapted to studying other systems in which there is large scale cellular movement throughout an organism.
Studying cell mobility within these complex environments has never been easy and, for many years, researchers lacked good tools to directly follow immune cells in lymphoid organs. The development of cell sorters together with the engineering of fluorescently labelled antibodies made it possible to track cells from one organ to another. Thus several labs were able to decipher the journey of lymphocytes from the bone marrow to the thymus where they gain their specificity or from the thymus to the lymph nodes. These methods gave researchers much more information regarding immune cell mobility than the static microscopy images of chemically coloured lymphoid organs did in the past.
How do researchers track fluorescently labeled cells?
1. Epifluorescence microscopy
This technique uses a fluorescence microscope where the light source is mounted above (epi) the specimen and the excitation light passes through the microscope objective lens on its way toward the specimen. It allows the visualization of fluorescent proteins expressed in the specimen.
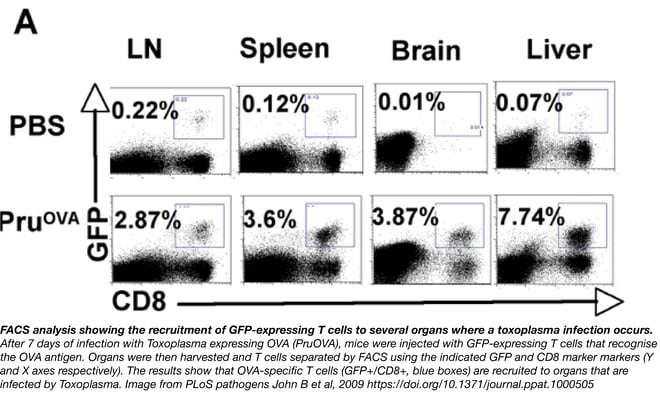
2. Fluorescence Activated Cell Sorter (FACs)
A machine that sorts cells according to whether or not they have been tagged with a fluorescent protein or dye. It separates the cells mechanically in a vibrating nozzle, imparting a positive or negative charge to cells that fluoresce, and then passing the cells through an electric field to deflect them into appropriate containers. The machine is useful for distinguishing cell populations that have been tagged with different fluorescent proteins or dyes.
 3. Two-photon microscopy
3. Two-photon microscopy
Two-photon excitation microscopy is a fluorescence imaging technique that allows imaging of living tissue up to about one millimeter in depth. Two low-energy photons (typically from the same laser) cooperate to cause a higher-energy electronic transition in a fluorescent molecule (see figure to the right). The excitation generated by these two-photons occurs only at a chosen focal volume and thus the microscope captures only fluorescence coming from this volume. The consequence is that you can detect signal in thick specimens. See the video below for an example of two-photon intravital microscopy from PLoS Pathogens - Kamenyeva et al, 2015.
Cellular trafficking studies enabled by fluorescent proteins
The optimization of GFP and its derivatives by the late Roger Tsien changed everything. For the first time, fluorescent proteins gave researchers the ability to track immune cells within organs and visualize how cells interact after a particular stimulus. Using an epifluorescence microscope and GFP fusion proteins controlled by promoters specific to the immune lineage, scientists could easily track GFP expressing cells. For example, by harvesting lymph nodes at different times after antigen challenge, scientists were able to track where GFP labelled B cells were in mounted slices of these organs. These GFP labelled cells could also be tracked by cell sorter giving immunologists the ability to count cells and statistically analyze cell movements. By using cells labelled by fluorescent chemicals or by other fluorescent proteins, labs were also able to see movement and interaction between several different immune cells. These methods allowed immunologists to develop a spatiotemporal view of an immune response, but the study of these dynamics was still laborious as epifluorescence microscopy wasn’t able to give a clear view of cellular movement in real time.
Two-photon microscopy was the next revolution in cell imaging, giving researchers the ability to monitor cellular movements in real time. This technique was first Introduced to immunology by three papers in the journal Science in 2002. Now, this technique and the in vivo live-cell imaging (intravital imaging) studies it’s enabled are revealing the cellular behaviors that mediate adaptive and innate immunity in diverse tissue environments. These studies provide quantitative measurements of cellular motility, interactions, and response dynamics.
In the past 15 years efforts have been made to create transgenic mice expressing fluorescent protein reporter constructs in the immune lineage. By specifically labeling T cells, B cells, and antigen presenting cells, scientists have been able to decypher cellular dynamics in lymphoid organs. In lymph nodes, for example, two-photon microscopy allowed scientists to better understand T cell initiation in the adaptive immune response. It was known that T cells were able to extravasate from the bloodstream to invade lymph nodes and scan antigen presenting cells, but two-photon microscopy enabled scientists to see how chaotic and random the scanning was and how fast T cells could jump from one antigen presenting cell to another. Indeed, we now know that T cells are able to crawl more rapidly than any other cell type in the body!
Intravital imaging powered by two-photon microscopy has uncovered host-pathogen interactions leading to the understanding of effector function in infected tissues. Before the use of two-photon imaging, our understanding of pathogen-immune cell interaction relied on in vitro studies where it was difficult to apprehend the key role of specialized cell types and organs that exist in vivo. The use of fluorescent pathogens and fluorescent cells enables scientists to monitor cell interactions in diverse tissues, including lymph nodes, brain, liver, gut and skin.
Two-photon intravital microscopy. The video above depicts the infiltration of neutrophils expressing LysM-GFP (green) into a draining lymph node after infection with S. aureus (red). You can then see neutrophils swarming around S. aureus and eating the bacteria. Video reproduced from Kamenyeva et al. 2015.
Some of the many questions scientists have been able to help answer using two-photon microscopy include:
- How do T cells and/or Neutrophils travel to the sites of infection or damage (Peters et al. 2008, Chtanova et al. 2008, Kamenyeva et al. 2015)?
- How do blood monocytes patrol blood vessels during immune responses (Auffray et al. 2007, Finsterbusch et al. 2016)?
- How do dendritic cells migrate to lymph nodes to activate T cells (Celli et al 2008, Kitano et al. 2016, Cavanagh et al. 2008)?
Technical challenges of two photon microscopy
Intravital imaging is an easy way to understand what is going on in the body at steady state or under physio-pathological challenge. However technical specificities have to be taken into consideration especially in the use of fluorescent proteins to track cellular movements and interactions. Even if virtually all fluorescent proteins can be used in your experiments, experience has shown that some are more reliable and more efficient at tracking cells. Good two-photon probes when using a standard Ti:Sapphire Laser include:
- Blue-green fluorescent proteins like EGFP
- Yellow-orange fluorescent proteins like TagRFP, tdTomato, DsRed, the mKate series, or tdKatushka2 (Drobizhev et al. 2011)
The commonly used orange and red fluorescent proteins are excited by 750 nm to 760 nm laser light, enabling dual color imaging studies with blue or cyan proteins without changing excitation wavelength (Salomonnson et al. 2012). They can also be excited efficiently at wavelengths between 1,000 nm and 1,200 nm, where there is relatively little tissue absorption, weak tissue scattering and small amounts of tissue auto-fluorescence (Drobizhev et al. 2011). But this requires other lasers than the standard Ti:Sapphire laser. It is also important to finely tune the excitation wavelength as it has been shown that even a small incremental change of the excitation wavelength can significantly affect emission intensities from fluorescent proteins (Salomonnson et al. 2012).
Finally, it is worthwhile to test the photostability of your fluorescent protein when using 2-photon microscopy as this hasn't been well reported for many fluorescent proteins.
Additional fluorescent protein tools
Photoactivable fluorescent proteins (proteins that fluoresce after a light-induced chemical reaction) or photoconvertible Keade proteins (protein that undergoes light-induced irreversible photoconversion from green fluorescence to red fluorescence) are also useful tools to use with intravital imaging. These tools allow one to study the kinetics of cell migration and movement by marking cells at a specific time. Photoactivable fluorescent proteins, for example, have been successfully used to study B and T cell dynamics within the germinal center of murine lymph nodes. However, one must pay attention to time scale when using photoactivatable FPs - their degradation within labelled cells limits the timespan available for tracking certain cell types.
Another useful tool for marking and tracking cells is the the Cre/lox recombination system. A researcher can flank fluorescent proteins in a plasmid with loxP sites such that their expression is turned on or off in cells expressing Cre. The result of a system like this is a mosaic of cells labelled with different fluorescent proteins. Using one iteration of this system called Brainbow, the Ubow mouse strain has been created to fate map Langerhans cells and follicular dendritic cells within the skin and lymph node respectively. The permanent labelling of cells using cre-lox is advantageous when compared to photoactivable or photoconvertible proteins, as it enables tracking throughout the lifetime of the cell.
The past 15 years have seen a lot of development surrounding the use of fluorescent proteins and intravital imaging in understanding the key roles of immune cells in triggering an immune response. New fluorescent proteins can be used more efficiently in two-photon microscopy and new constructs have facilitated the creation of mouse lines that can be used in fate mapping experiments. Immune response dynamics are much better understood thanks to these technologies but many questions remain. The use of other technologies such as optogenetics and CRISPR/Cas9 will help immunologists create even better tools and models to further our understanding of the immune system and its dynamics.
References
1. Peters, Nathan C., et al. "In vivo imaging reveals an essential role for neutrophils in leishmaniasis transmitted by sand flies." Science 321.5891 (2008): 970-974. PubMed PMID: 18703742. PubMed Central PMCID: PMC2606057.
2. Chtanova, Tatyana, et al. "Dynamics of neutrophil migration in lymph nodes during infection." Immunity 29.3 (2008): 487-496. PubMed PMID: 18718768. PubMed Central PMCID: PMC2569002.
3. Egen, Jackson G., et al. "Macrophage and T cell dynamics during the development and disintegration of mycobacterial granulomas." Immunity 28.2 (2008): 271-284. PubMed PMID: 18261937. PubMed Central PMCID: PMC2390753.
4. Egen, Jackson G., et al. "Intravital imaging reveals limited antigen presentation and T cell effector function in mycobacterial granulomas." Immunity 34.5 (2011): 807-819. PubMed PMID: 21596592. PubMed Central PMCID: PMC3164316.
5. Chieppa, Marcello, et al. "Dynamic imaging of dendritic cell extension into the small bowel lumen in response to epithelial cell TLR engagement." Journal of Experimental Medicine 203.13 (2006): 2841-2852. PubMed PMID: 17145958. PubMed Central PMCID: PMC2118178.
6. Schaeffer, Marie, et al. "Dynamic imaging of T cell-parasite interactions in the brains of mice chronically infected with Toxoplasma gondii." The Journal of Immunology 182.10 (2009): 6379-6393. PubMed PMID: 19414791.
7. John, Beena, et al. "Dynamic imaging of CD8+ T cells and dendritic cells during infection with Toxoplasma gondii." PLoS Pathog 5.7 (2009): e1000505. PubMed PMID: 19578440. PubMed Central PMCID: PMC2700268.
8. Hickman, Heather D., et al. "Direct priming of antiviral CD8+ T cells in the peripheral interfollicular region of lymph nodes." Nature immunology 9.2 (2008): 155-165. PubMed PMID: 18193049.
9. Filipe-Santos, Orchidée, et al. "A dynamic map of antigen recognition by CD4 T cells at the site of Leishmania major infection." Cell host & microbe 6.1 (2009): 23-33. PubMed PMID: 19616763.
10. Drobizhev, Mikhail, et al. "Two-photon absorption properties of fluorescent proteins." Nature methods 8.5 (2011): 393-399. PubMed PMID: 21527931. PubMed Central PMCID: PMC4772972.
11. Salomonnson, Emma, et al. "Cell-based and in vivo spectral analysis of fluorescent proteins for multiphoton microscopy." Journal of biomedical optics 17.9 (2012): 0960011-0960019. PubMed PMID: 22975677. PubMed Central PMCID: PMC3438408.
12. Kamenyeva, Olena, et al. "Neutrophil recruitment to lymph nodes limits local humoral response to Staphylococcus aureus." PLoS Pathog 11.4 (2015): e1004827. PubMed PMID: 25884622. PubMed Central PMCID: PMC4401519.
13. Auffray, Cedric, et al. "Monitoring of blood vessels and tissues by a population of monocytes with patrolling behavior." Science 317.5838 (2007): 666-670. PubMed PMID: 17673663.
14. Finsterbusch, Michaela, et al. "Patrolling monocytes promote intravascular neutrophil activation and glomerular injury in the acutely inflamed glomerulus." Proceedings of the National Academy of Sciences (2016): 201606253. PubMed PMID: 27528685. PubMed PMCID: PMC5024581.
15. Celli, Susanna, Béatrice Breart, and Philippe Bousso. "Intravital two-photon imaging of natural killer cells and dendritic cells in lymph nodes." Innate Immunity (2008): 119-126. PubMed PMID: 18370151.
16. Kitano, Masahiro, et al. "Imaging of the cross-presenting dendritic cell subsets in the skin-draining lymph node." Proceedings of the National Academy of Sciences 113.4 (2016): 1044-1049. PubMed PMID: 26755602. PubMed Central PMCID: PMC4743831.
17. Cavanagh, Lois L., and Wolfgang Weninger. "Dendritic cell behaviour in vivo: lessons learned from intravital two-photon microscopy." Immunology and cell biology 86.5 (2008): 428-438. PubMed PMID: 18431356.
Additional Resources on the Addgene Blog
- Get Tips on Choosing Your Fluorescent Protein
- Learn Some of the Pitfalls of Working With Fluorescent Proteins
- Use FRET to Monitor Protein-Protein Interactions
Resources on Addgene.org
- Check out Our Fluorescent Protein Guide Pages
- Find Plasmids for Subcellular Localization
- Find Fluorescent Protein Empty Vectors
Topics: Fluorescent Proteins, Fluorescent Proteins 101, Localization with Fluorescent Proteins






Leave a Comment