This post was contributed by guest bloggers Joachim Goedhart and Marieke Mastop from the Section of Molecular Cytology and Van Leeuwenhoek Centre for Advanced microscopy, University of Amsterdam.
The previous post in this series described a practical approach to selecting a bright fluorescent protein. In the second post of this series, we will discuss how to select a photostable fluorescent protein.
Photobleaching is the irreversible destruction of a fluorophore under the influence of light. Any fluorescent molecule will photobleach at some point. For live-cell imaging, it is desirable to have fluorescent proteins that are photostable. On top of photobleaching, fluorescent proteins may display reversible intensity changes (Shaner et al, 2008; Bindels et al, 2017) and photoswitching (Kremers et al, 2009), which usually are undesired properties. In the ideal situation, a fluorescent proteis should emit a stable fluorescence signal, showing no or little deterioration or change of the signal during the course of the experiment.
The best fluorescent proteins for live cell imaging can be excited many times, thereby producing a large number of emitted photons before they are destroyed.
Factors that affect photostability
The photobleaching rate of a fluorophore primarily depends on the excitation power and the excitation wavelength. A fluorophore is less likely to bleach when it is excited off the peak wavelength, since it will absorb the excitation light less efficiently. Similarly, decreasing the power of the excitation light reduces the number of excitation/emission cycles per unit of time, decreasing the likelihood that a fluorophore will bleach. This is, however, not the full story. The photobleaching rate does not depend on excitation power in a linear fashion. This implies that reducing the power 2-fold does not reduce photobleaching by half. How exactly the photobleaching rate depends on power is a property of the fluorescent protein. This can differ between fluorescent proteins in the same spectral class (Cranfill et al, 2016).
This non-linear dependence on excitation power is important since different fluorescence imaging strategies use widely varying amounts of excitation power (Shaner et al, 2008). In confocal laser scanning microscopy, fluorescent proteins are excited with very intense light (for a brief time), whereas in widefield imaging fluorophores are excited with relatively low light levels (for a prolonged time). Other illumination strategies, such as 2-photon-excitation, selective plane illumination, TIRF or spinning disk confocal use entirely different excitation power regimes. Therefore, it is unpredictable which fluorescent protein is most photostable for each of these conditions. On top of these factors, environmental conditions such as cellular redox state and oxygen concentration may affect photobleaching rates (Shaner et al, 2008). This brings us to the key question: what is the best way to compare the photostability of different fluorescent proteins?
Measuring photostability
 Since it is clear that a photostable fluorescent protein is a key requirement for quantitative live-cell imaging, it is important to be able to quantify the photostability of fluorescent proteins. To determine photostability, an experiment is performed that measures fluorescence intensity over time. In order to predict how fluorophores perform in ‘real experiments’, it is advisable to perform time-lapse imaging of cells producing the fluorescent protein of interest under realistic conditions, i.e. with low excitation power. By repeating these measurements for different fluorescent proteins and by comparing how the fluorescence intensity changes over time, the photostability of the fluorescent proteins can be directly compared.
Since it is clear that a photostable fluorescent protein is a key requirement for quantitative live-cell imaging, it is important to be able to quantify the photostability of fluorescent proteins. To determine photostability, an experiment is performed that measures fluorescence intensity over time. In order to predict how fluorophores perform in ‘real experiments’, it is advisable to perform time-lapse imaging of cells producing the fluorescent protein of interest under realistic conditions, i.e. with low excitation power. By repeating these measurements for different fluorescent proteins and by comparing how the fluorescence intensity changes over time, the photostability of the fluorescent proteins can be directly compared.
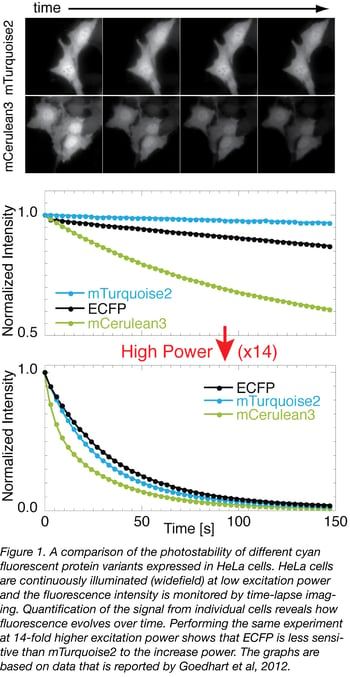
An example of a photostability measurement of various cyan fluorescent proteins is depicted in figure 1. Of note, the photobleaching rate does not depend on fluorescence intensity or protein distribution. Therefore, photostability measurements can be performed with soluble fluorescent proteins or localized fusion proteins and do not require dedicated plasmids or constructs.
It should be noted that the photostability measurements reported in the literature are performed in different ways. We describe a number of issues with some of the experimental designs used to make these measurements and hope this information can help guide your understanding of reported data.
Photobleaching results in a loss of fluorescence, which, in its simplest form, can be described by a mono-exponential decay (analogous to radioactive decay). Consequently, the intensity decay is often described by the t½, which is the time after which half of the initial fluorescence intensity remains. The first issue is that, in several cases, the fluorescence intensity decay of a fluorophore does not follow a simple mono-exponential decay (Shaner et al, 2008; Bindels et al, 2017) and cannot be described by a single parameter. Therefore, it is necessary to know how the fluorescence intensity evolves over time.
The second issue is that, in experiments that measure photostability, high excitation powers are often used to reduce the length of the experiments. As the photobleaching rate is not linearly related to power, the conclusions reached at high power may not translate to real applications in which far less power is used. For example you might find that you can get useful data from your chosen FP for much longer than predicted by these high power experiments. Measuring photostability at realistic powers will provide better insight into the photostability in the intended application (Goedhart et al, 2012).
The third issue relates to the environment of the fluorescent protein. Photostability measurements can be performed on purified fluorescent proteins. To avoid diffusion, the proteins are (i) trapped in microdroplets in a water/oil emulsion, (ii) embedded in a gel, or (iii) attached to a substrate. These in vitro methods allow for a well-controlled environment, but they do not mimic the natural situation. Measuring photostability in living cells provides a much more realistic view of photostability.
Selecting a photostable protein
The issues with quantification of photobleaching rates can be summarized as follows: Instead of determining a t½ of a fluorescent protein in a water/oil emulsion at high power, it is far more relevant to measure how fluorescence intensity evolves over time in living cells at a power used for live-cell imaging.
To conclude, the most photostable fluorescent protein can be identified by a head-to-head comparison of several fluorescent proteins of the same spectral class on the microscopy system and in the cell type (or tissue) that is intended to be used with the fluorescent protein. In figure 1 we provide an example of such a comparison between cyan fluorescent proteins using widefield imaging. Our choice for a photostable protein for live cell imaging would be mTurquoise2, based on the data acquired at low excitation powers. Note, however, that, at a 14-fold higher excitation power, the photostability of the variants is comparable (lower panel of figure 1).
To avoid changes in excitation power, it is important to perform the experiments on a single day on a stable system. A final piece of advice is to measure the excitation power you use in your imaging experiments. It is good practice to measure the excitation power (Grünwald et al, 2008) occasionally and certainly after a set-up has been changed (for instance after replacing a broken lightbulb or exchanging excitation filters). This avoids substantial changes in excitation power and will help to keep the photostability of the probes in check.
Many thanks to our guest bloggers Joachim Goedhart and Marieke Mastop!
 Joachim Goedhart is an assistant professor at the Section of Molecular Cytology and Van Leeuwenhoek Centre for Advanced microscopy (University of Amsterdam). He develops, characterizes and uses genetically encoded fluorescent probes. You can follow him on twitter: @joachimgoedhart.
Joachim Goedhart is an assistant professor at the Section of Molecular Cytology and Van Leeuwenhoek Centre for Advanced microscopy (University of Amsterdam). He develops, characterizes and uses genetically encoded fluorescent probes. You can follow him on twitter: @joachimgoedhart.
 Marieke Mastop is a PhD candidate at the Section of Molecular Cytology (University of Amsterdam) where she develops genetically encoded FRET based biosensors.
Marieke Mastop is a PhD candidate at the Section of Molecular Cytology (University of Amsterdam) where she develops genetically encoded FRET based biosensors.
Additional Resources on the Addgene Blog
- Which Fluorescent Protein Should I Use?
- Choosing Your Fluorescent Proteins for Multicolor Imaging
- Learn about Fluorescent Protein Biosensors
Additional Resources on Addgene.org
- Find Empty Backbones with Fluorescent Tags
- Find Plasmids for Subcellular Localization
- Browse the entire Fluorescent Protein Collection






Leave a Comment