 Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
New toolkit for N or C terminal tagging of proteins for degradation using AID
Article contributed by Alyssa Cecchetelli
Auxin Inducible Degradation (AID) uses plant hormones called auxins to conditionally and reversibly degrade proteins of interest fused to a degron, a domain that induces degradation. In plants, auxin binds to TIR1 of the SCF-TIR1 ubiquitin ligase complex, promoting its interaction with the degron sequence and ultimately its degradation. While other eukaryotes don’t naturally have this response, they do contain the specific ubiquitin ligase pathway needed for AID. By tagging proteins with a 7kD degron called mAID and expressing the essential plant protein TIR1 in non-plant cells, which interacts with the cells’ endogenous ubiquitin ligase, the Kanemaki lab was able to translate AID to yeast, as well as mammalian and human cell lines (Nishimura et al., 2009, Natsume et al., 2016).
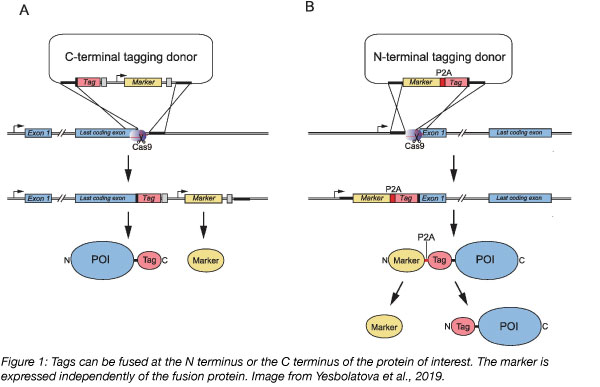
The Kanemaki lab previously developed a CRISPR/Cas9 based method to tag endogenous proteins in HCT116 cells with mAID. Using this method, the lab was able to generate conditional alleles of a few essential proteins (Natsume et al., 2016). Now, the Kanemaki lab has created a new toolkit of plasmids for the N- or C- terminal tagging of proteins with mAID and other tags (Yesbolatova et al., 2019) using CRISPR. This toolkit makes it easy to tag your protein of interest to create conditional human cell lines. Lastly, the Kanemaki lab demonstrated that auxinole, a TIR1 inhibitor, sufficiently suppresses basal degradation of tagged proteins, a common problem of the AID technology (Yesbolatova et al., 2019).
Find the mAID tagging toolkit at Addgene
Yesbolatova et al., Methods, 2019. PubMed PMID: 31026591.
Expanding the targeting range of CRISPRCas9 by engineering SpCas9
Article contributed by Shreya Vedantam
CRISPR/Cas9 is widely used genome editing tool. However, Cas9’s endonuclease activity can be limited by the need for a specific PAM (protospacer adjacent motif) in order to bind the target sites for editing. PAM sequences can be from 2-6 base pairs in length and follow the DNA sequence targeted by the Cas9 nuclease.
The Streptococcus pyogenes Cas9 (SpCas9) requires an NGG PAM to recognize the target. Osamu Nureki’s lab engineered SpCas9 (SpCas9-NG) with relaxed preferences for the third base of the PAM so that it now recognizes NG rather than NGG. In addition to the engineered SpCas9, they also fused the SpCas9-NG with the activation-induced cytidine deaminase Target-AID-NG, which mediated a C to T conversion in the target sites adjacent to an NG PAM in human cells. The increased versatility of the engineered SpCas9-NG and its variants can be used to engineer specific sites that were previously difficult.
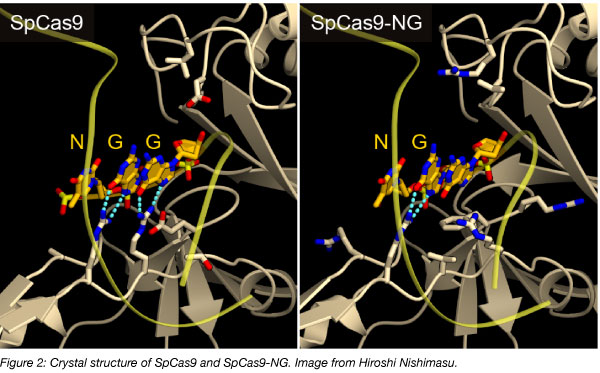
Find SpCas9 and Target-AID-NG plasmids at Addgene!
Nishimasu H et al., Science, 2018. PubMed PMID: 30166441.
MerMAIDs - marine channelrhodopsins for the optogenetic toolbox
Article contributed by Susanna Bachle
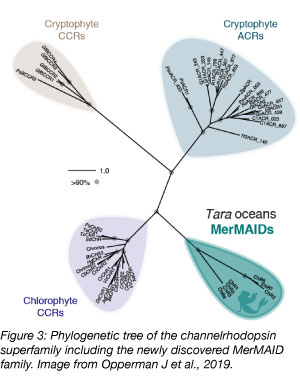 Anion-conducting channelrhodopsins (ChR) are ion channels that can be used to silence the activity of neurons. Earlier this year, the Hegemann lab and collaborators report their discovery of a new family of seven ChRs derived from marine microorganisms based on metagenomic data. MerMAIDs (Metagenomically discovered Marine, Anion-conducting and Intensely Desensitizing ChRs) differ from currently used ChRs in that they cause a rapid and almost complete desensitization when exposed to continuous bright light. Desensitization is mediated by a conserved cysteine. With the introduction of MerMAIDs, neurons can be turned off quickly and with high precision - a valuable optogenetics feature when studying neuronal activity.
Anion-conducting channelrhodopsins (ChR) are ion channels that can be used to silence the activity of neurons. Earlier this year, the Hegemann lab and collaborators report their discovery of a new family of seven ChRs derived from marine microorganisms based on metagenomic data. MerMAIDs (Metagenomically discovered Marine, Anion-conducting and Intensely Desensitizing ChRs) differ from currently used ChRs in that they cause a rapid and almost complete desensitization when exposed to continuous bright light. Desensitization is mediated by a conserved cysteine. With the introduction of MerMAIDs, neurons can be turned off quickly and with high precision - a valuable optogenetics feature when studying neuronal activity.
The researchers identified MerMAIDS from their analysis of different publicly available metagenomic data including the Tara Oceans Project datasets. This study demonstrates the power of combining the analysis of publicly available metagenomic datasets with biophysical studies and the potential of mining datasets for research and innovation.
Find MerMAIDs plasmids at Addgene
Opperman J et al., Nature Communications, 2019. PubMed PMID: 31346176.
Using LADL to loop with light
Article contributed by Eric Perkins
Mammalian genomic DNA is not just a 2-D linear molecule snaking through our cells’ nuclei--our chromatin is comprised of thousands of loops. These loops not only provide a 3-D structure for packing genomic DNA into nuclei, but are also thought to play a key role in the transcriptional regulation of many genes. A number of groups have designed means of studying the effects of endogenous looping on distal promoters and enhancers, or have attempted to engineer loops to create genomic interactions that wouldn’t normally occur. The synthetic looping factors developed thus far for these latter experiments, such as transcription factors tethered to dCas9, require long time scales with constitutively expressed proteins that are not inducible.
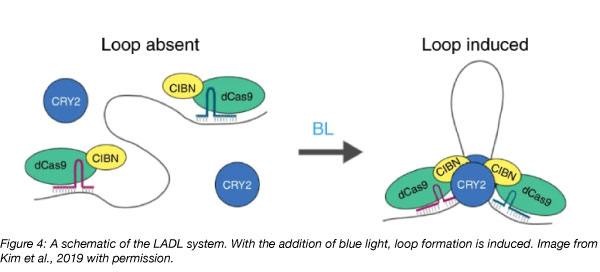
To further develop dCas9 for synthetic looping, the Phillips-Cremins lab developed the light-activated-dynamic-looping (LADL) system. The group fused dCas9 to a truncated CIB1 protein from Arabidopsis (CIBN), which they call the LADL Anchor. Using specifically designed gRNAs, they can then recruit these LADL Anchors to genomic targets, such as a distal promoter and an enhancer. What makes LADL unique from previous iterations is the addition of CRY2, another Arabidoposis protein that heterodimerizes with CIBN upon blue light exposure (see Fig 1*). When the group targeted the enhancer region of Klf4 and the promoter region of Zfp462, which are over 500kB apart, they were able to observe loop formation as well as increased Zfp462 expression within 4 hours of blue light induction (6-fold faster than previously developed systems). The LADL system promises to be a better model for the dynamic nature of genomic looping due to it’s briefer time scale and its inducible and reversible nature.
Find the LADL plasmids at Addgene
Kim et al., Nat Methods. 2019, PubMed PMID: 31235883.
Topics: Other Plasmid Tools, Plasmids






Leave a Comment