 Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Listen to the Hot Plasmids Podcast!
MarathonRT: A highly processive reverse transcriptase
Article contributed by Will Arnold
![]() Listen to the MarathonRT podcast segment
Listen to the MarathonRT podcast segment
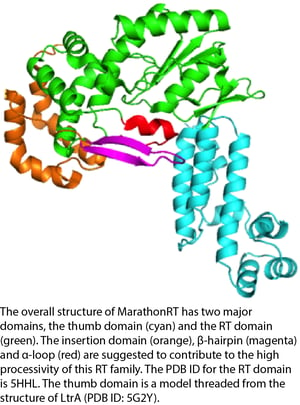
Researchers from Dr. Anna Pyle’s lab at Yale University recently deposited a plasmid for expressing a novel reverse transcriptase (RT), MarathonRT. Reverse transcriptases are critical tools for many fields of biology and enable the conversion of RNA templates to DNA but suffer from several important shortcomings including low processivity (ie. the probability of the polymerase continuing to copy the template rather than dissociating). MarathonRT, isolated from Eubacterium rectale, significantly outperforms other RTs in this regard and is even able to reverse transcribe the extremely long, highly structured HCV genome (~9.6 kb of highly structured template!). MarathonRT also faithfully transcribes RNA to DNA with low error rates compared to other RTs.
MarathonRT represents a major step forward in facilitating techniques that are dependent on long, error-free, reverse transcription of RNA such as long-read cDNA sequencing and RNA sequencing of complex heterogeneous RNA samples. Furthermore, such high processivity enzymes and the structural understanding thereof have the potential to provide improvements to existing technologies based on polymerase processivity.
Please find the MarathonRT plasmid here!
Zhao C, et al. RNA. 2018 PubMed PMID:29109157.Nanobody toolkit to monitor retrograde transport at the cell surface
Article contributed by Marcy Patrick
![]() Listen to the nanobody toolkit podcast segment
Listen to the nanobody toolkit podcast segment
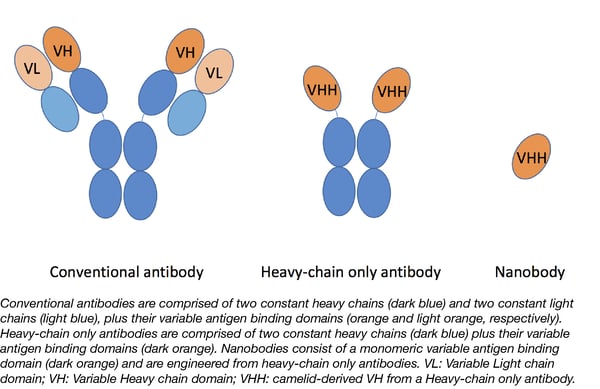
Nanobodies are single domain antibody fragments typically engineered from heavy-chain only antibodies found in camelids. They possess many advantages over conventional antibodies: they are small (12-15 kDa), stable, highly soluble, and non-immunogenic yet are still just as specific to their target as regular antibodies. These properties make nanobodies easier to transform and express in bacteria, which, in turn, makes them more adaptable to a broad range of applications.
Recently, the Spiess lab developed a set of functionalized nanobodies to study retrograde transport from the cell surface. Their toolkit includes a set of plasmids each containing an anti-GFP nanobody fused with a different functional element (tyrosine sulfation consensus sequence, fluorophore, or APEX2), allowing for quantitative analysis using different detection methods. Since the anti-GFP nanobody is able to bind fluorescent proteins derived from A. victoria the functionalized nanobodies described here can be used to target any protein fused with GFP, YFP, mCerulean, or other related fluorophores.
Find all of the nanobody toolkit plasmids here.
- Buser DP, et. al. PNAS 2018. PubMed PMID: 29915061.
A new method to identify proteins occupying a specific genomic locus: Caspex
Article contributed by Susanna Bachle
![]() Listen to the Caspex podcast segment
Listen to the Caspex podcast segment
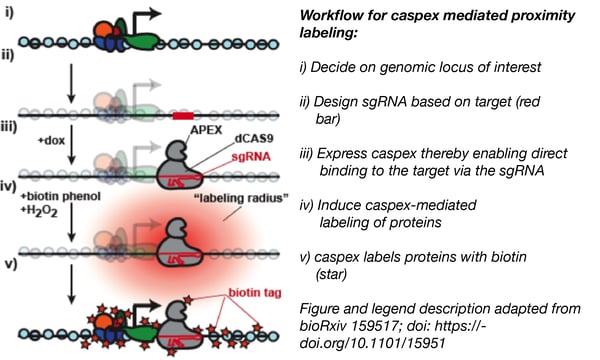
Steven Carr’s lab at the Broad Institute focuses on proteomics: developing and applying new technologies to quantify proteins in tissues, cells, and biofluids. Recently, Sam Myers, a Scientist in the group, developed a CRISPR-based approach to detect proteins that associate with specific genomic loci. He made use of catalytically inactive dCas9 by fusing it to an engineered peroxidase, (APEX) that labels proximal proteins with biotin as an affinity tag. This ‘Caspex’ construct is guided by a pre-defined sgRNA sequence to a specific genomic locus and labels the proteins in its vicinity with biotin. The biotin-labeled proteins can then be isolated and analysed. In this work, the authors specifically investigated protein binding at the TERT gene, but this method enables the unbiased discovery of proteins involved in transcriptional regulation and chromatin structure at your locus of choice.
Find the Tet-inducible caspex plasmid here.
- Myers SA et al. Nature Methods. 2018. PMID: 29735997
- bioRxiv preprint posted July 4, 2017
A new method to identify proteins occupying a specific genomic locus: C-BERST
Article contributed by Klaus Wanisch
![]() Listen to the C-BERST podcast segment
Listen to the C-BERST podcast segment
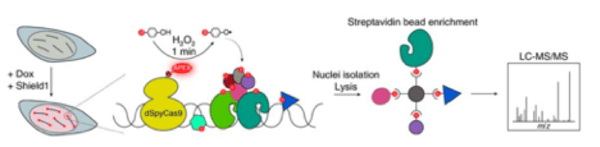
Similarly to the Carr lab, Erik Sontheimer and his group from the University of Massachusetts Medical School have developed a group of plasmids (here) that can be used to label all proteins within a radius of approximately 20 nm of a specific DNA sequence in only 1 min. Using these lentiviral plasmids, catalytically inactive Cas9 (dSpyCas9) fused to APEX2 is expressed from a Tet/Shield1 inducible CMV promoter and targeted to a sequence of interest with an sgRNA expressed from a separate plasmid. When cells are treated with biotin–phenol and H2O2, APEX2 labels electron-rich amino acid side chains (e.g. Tyrosine) with biotin. These biotin-labeled proteins can them be analyzed by mass spectrometry allowing a researcher to specifically identify proteins that are enriched in the 20 nm radius surrounding the dSpyCas9-APEX2 fusion.
This Sontheimer lab used this technology, which they call C-BERST (dCas9–APEX2 biotinylation at genomic elements by restricted spatial tagging), to confirm the composition of proteins found at the telomere, and hope it can be used to expand our understanding of proteome localization in future studies of other genomic loci.
Find the C-BERST plasmids here.
-
Gao at al. Nature Methods (2018) Pubmed PMID: 29735996.
Topics: Other Plasmid Tools, Plasmids





Leave a Comment