 Every few months we highlight a subset of the new plasmids and viral preps in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Every few months we highlight a subset of the new plasmids and viral preps in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Here's what you'll find in this post:
- Yeast-based protein evolution
- Fluorescent tools for mammalian cells
- Nanobody purification
- CRISPRi and CRISPRa system for hPSCs
- The CRISPR corner
- New from the viral service
Faster catalysis after yeast-based protein evolution
Mateo Sanchez and Alice Ting have used one biotechnology tool to optimize another. The authors have created a yeast-based directed evolution approach with some notable improvements. Directed evolution techniques mimic natural selection of sequences that evolve a measurable property of a protein. As a proof of concept, Sanchez and Ting sought to improve the catalytic efficiency of tobacco etch virus protease (TEV), a highly sequence-specific protease used to cleave affinity tags from purified recombinant proteins at small and large scale.
This new technique involves selection in the cytosol (instead of on the yeast extracellular surface) which enables tracking of protease activity with a fluorescent protein--no affinity purification required! The team used other molecular techniques to introduce time modulation of the TEV cleavage and improve utility of the system for use with low affinity variants of TEV. The new system has been demonstrated to work for evolution of other target proteins and might be useful for your studies.
Find the plasmids to set up your screening systems at Addgene!
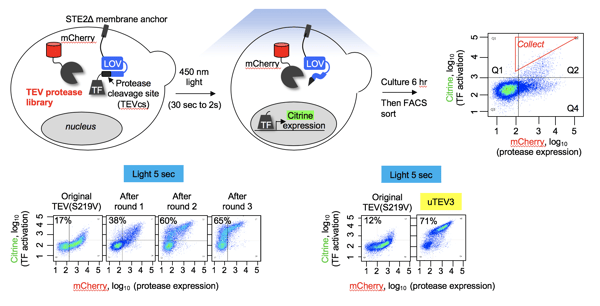 |
| Schematic of the evolution platform to generate TEV protease variants. Catalysis by TEV protease allows a transcription factor to enter the nucleus and modulate transcription of the mCitrine gene. mCherry is constitutively expressed. Cells are then cultured and FACS sorted. Image from Mateo Sanchez and Alice Ting. |
Sanchez MI et al., Nat methods. 2019. https://doi.org/10.1038/s41592-019-0665-7
Optimizing fluorescent tools for labeling structures and compartments in mammalian cells
Genetically encoded fluorescent proteins are often used to label specific structures, compartments, or specific localization of biomolecules within cells. However, sometimes these fluorescent proteins might not localize to the right places, which can provide misleading information on where exactly a structure or protein is located within a cell. The Goedhart lab, Gadella lab, and collaborators have an ongoing effort to improve genetically encoded fluorescent markers for labeling specific structures within cells.
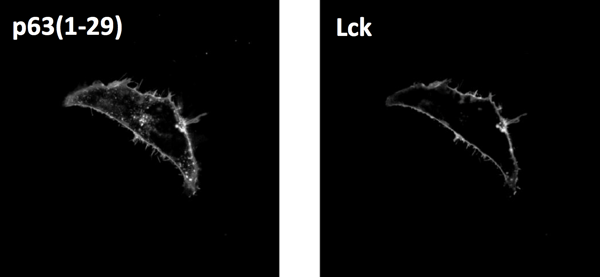 |
| HeLa cells were transfected with plasmids encoding FPs tagged with the plasma membrane targetin sequence derived either from p63(1-29) and Lck. Image from Chertkova et al., 2020. |
In their BioRxiv post, the authors generated and described several improved genetically encoded fluorescent markers tagged with mTurquoise2, mNeonGreen, and mScarlet-I for labeling a list of specific structures within cells. For example, to improve nuclear localization of fluorescent proteins, they found that adding a triple nuclear localization signal or including fusions with Histone 2A or 2B can result in bright targeted expression while minimizing non-specific labeling. For labeling the plasma membrane, they tested different target sequences for improved selectivity and found that the best motif was from the Lck gene. For marking microtubules of mammalian cells, the author found that too high expression of these markers can lead to non-specific labeling. They generated plasmids tagged with EB3 to mark the growing tips of microtubuli and plasmids tagged with MapTau to label entire microtubules.
Find these fluorescent mammalian cell structure markers plasmids at Addgene!
Chertkova, AO, et al., bioRxiv. 2020. https://doi.org/10.1101/160374
Nanobodies- there's a simple method for that!
The benefits of nanobodies over conventional antibodies can’t be overlooked. Nanobodies are smaller in size, more durable, and usually much easier to clone, express, and purify compared to antibodies. Nanobodies are easily expressed in the cytosol of E. coli, however the nanobody proteins tend to get stuck in inclusion bodies. Takayuki Iwaki, Kimiko Hara, and Kazuo Umemura have described a simple method which alleviates the inclusion body problem and makes nanobody production in E.coli as simple as following a DNA miniprep protocol!
First, the authors created the pMAK backbone by modifying pMAL-T-Avi-His/BirA, a vector commonly used for in vivo biotinylation via BirA and AviTAG. pMAK contains the signal peptide of MBP, followed by the desired nanobody (NB) sequence, an AviTAG, and 6xHis tag. The MBP signal peptide allows the NB to be exported to the periplasm of bacteria where it can be secreted into the supernatant. Four plasmids containing different nanobodies (NB) were generated in pMAK:
- pMAK461(NBαRabitb-IgG-Fc, Addgene #140699)
- pMAK462 (NBαMouse/Rat-IgG1-Fc, Addgene #140700)
- pMAK463 (NBαMouse-IgK, Addgene #140701)
- pMAK464 (NBαEGFP, Addgene #140702)
The pMAK461-64 plasmids were expressed in SHuffle T7 Express competent E.coli (NEB) using IPTG induction. To biotinylate the nanobodies, the authors used biotin dissolved in DMSO which allows both biotin and biotin-4-fluorescein to be dissolved at a higher concentration, and results in uniform NB biotinylation. Once induced, the NBs could be easily purified by one-step immobilized metal ion adsorption chromatography (IMAC) using Ni-NTA magnetic beads.
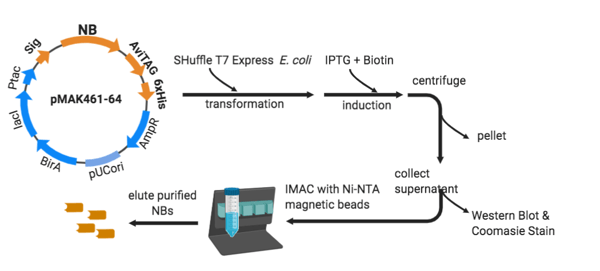 |
| Workflow of nanobody expression and purification. |
Iwaki et al., Protein Expr Purif, 2020. https://doi.org/10.1016/j.pep.2020.105607
A new multiplexed CRISPRi and CRISPRa system for human pluripotent stem cells
CRISPR gene interference (CRISPRi) and gene activation (CRISPRa) are great ways to modulate the expression of nearly any gene via specific guide RNA (gRNA) sequences. But these approaches rely on continual expression of both dCas9 and the gRNA. This expression can vary significantly depending on transgene design and delivery.
To overcome these variations in human pluripotent stem cells(hPSCs), Lindy Barrett’s lab at the Broad Institute developed fluorescently labeled piggyBac vectors for uniform and sustained expression of single or multiplexed gRNAs and of dCas9 fusions. The dCas-KRAB activator and the dCas-VPR repressor are introduced into the hPSC genome using AAV1 and contain an EGFP reporter. Expression of the dCas9-KRAB activator or dCas9-VPR repressor in hPSCs is also driven by a doxycycline inducible promoter. The gRNA piggyBac plasmids include a mRFP reporter. With this system, they could quantify and track cells expressing both dCas9 and gRNAs.
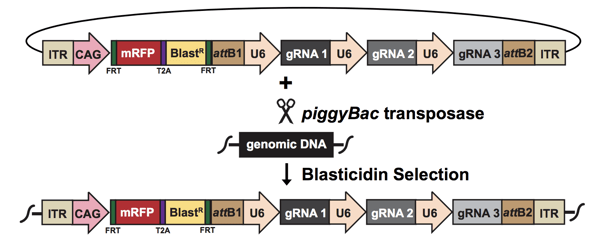 |
| The multi-gRNA piggyBac vector integrates into genomic DNA with the help of the piggyBa transposase. Image from Hazelbaker et al., 2020. |
The lab tested this system by repressing and activating expression of TCF4 while monitoring transcript and protein levels. Plasmids for multiplexed piggyBac gRNA delivery and the all-in-one dCas9-KRAB and dCas9-VPR targeting plasmids can be found at Addgene.
Check out these plasmids for piggyBac gRNA delivery and the all-in-one dCas9 targeting plasmids!
Hazelbaker et al., Sci Reports. 2020. https://doi.org/10.1038/s41598-020-57500-1
The CRISPR corner
Here are a few highlights from recent CRISPR plasmids. To find all of the CRISPR plasmids available from Addgene, head over to our CRISPR Plasmids and Resources page.
- PAC-MAN (prophylactic antiviral CRISPR in human cells) is a CRISPR-Cas13 based antiviral method to degrade RNA from SARS-CoV-2 sequences and live influenza A virus. It was demonstrated to work in human lung epithelial cells.
- The MoClo CRISPR/Cas Toolkit for plants includes 95 plasmids consisting of CRISPR/Cas nucleases, base editors, gRNA backbones, and promoters for expression in monocots and dicots.
- CRISPR RNP electroporation and AAV donor infection (CRISPR-READI) enables genome engineering using HDR donors lengths up to 4.9 kb.
- A new dual-deaminase base editor enables concurrent adenine and cytosine editing.
- A new tool for synthetic biology uses CRISPRi to control gene expression. This includes a CRISPRi-based synthetic oscillator, bistable network, stripe pattern-forming incoherent feed-forward loop.
New from the viral service
By Ina Ersing
We regularly add new viral aliquots from our plasmid collection to provide ready-to-use viral preps. Here are some of the new viral preps from recent months:
- New GPCR activation-based norepinephrine (GRABNE) sensors allow for sensitive and high spatiotemporal measurements of norepinephrine. These norepinephrine sensors include GRAB_NE1h (high affinity) and GRAB_NE1m (medium affinity), as well as GRAB_NEmut as a negative control.
- New yellow fluorescence protein-based calcium sensors (jYCaMP1), variants of the calcium indicator jGCaMP7, that are optimized for excitation wavelengths above 1,000 nm and enable two-color imaging with red fluorescence protein-based calcium sensors. These new calcium sensors include syn-jYCaMP1s (slow variant), the Cre-dependent syn-FLEX-jYCaMP1s, and the axon-targeted syn-FLEX-axon-jYCaMP1s.
- Cre-dependent EYFP AAV in the new serotype PHP.V1 that exhibits efficient transduction of vesicular brain cells.
- New serotypes for the Cre-dependent Flpo recombinase pEF1a-DIO-FLPo-WPRE-hGHpA, which is now also available in AAV5 and AAVrg.
- New controls, biosensor, chemogenetic and optogenetic AAV for expression in forebrain GABA-ergic interneurons. These include mDlx-GFP, hDlx-FLEX-GFP, hDlx-Flex-dTomato, mDlx-GCaMP6f, hDlx-GiDREADD-dTomato and mDlx-ChR2-mCherry.
Topics: Other Plasmid Tools, Plasmids






Leave a Comment