Last updated Sep 30, 2020 by Marcy Patrick.
CRISPR technology has been widely adopted for genome editing purposes because it's cheaper, faster, and easier than prior editing techniques. More and more CRISPR tools are being published each month, making CRISPR a great choice for your next experiment!
In this blog post we’ll provide an overview of some CRISPR mammalian expression systems, the typical applications for each, and potential delivery methods.
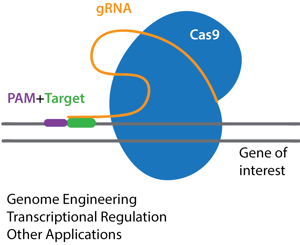
General considerations for planning your CRISPR experiment
As with any experiment, there are many factors that need to be considered during the planning process. For CRISPR experiments, the following framework can help get you started:
- Determine what type of outcome you are trying to achieve: Do you want to permanently knock out or knock-in a gene? Do you want to activate or repress gene expression? Are you trying to create a single point mutation? Do you want to create a fusion with a reporter protein such as GFP? All of these outcomes can be most effectively achieved with different CRISPR components.
- Select the appropriate CRISPR tools for your application: Wildtype Cas9 or Cas9 nickase are appropriate for generating knockouts or knock-ins, or introducing mutations and tags, while a catalytically dead dCas9 can be used in conjunction with activator or repressor domains to control gene expression. Base editors can help you make precise point mutations near the gRNA target site.
- Choose an appropriate expression system and delivery method: Do you need stable integration or is transient expression sufficient? Which cell types will you be editing? Do you want to deliver the components as DNA, or would mRNA or protein delivery be more suitable?
- Determine how you will evaluate the outcome: Will you be detecting insertions/deletions using a mismatch repair assay? Or is PCR followed by gel electrophoresis and/or next-generation sequencing more appropriate?

If you already have an end product in mind, steps 1 and 2 will generally be straightforward. Likewise for step 4, as this ties directly back to your specific goal. When thinking about step 3, however, you may be surprised at the number of options available - how do you choose between them?
One of the first steps is to identify what CRISPR components you will need to deliver. Minimally, one or more sgRNAs and Cas9 are required for any application. If you want to include a homology directed repair (HDR) template to create knock-ins, point mutations, or to add a tag, you will also need to deliver a donor plasmid or single-stranded DNA oligonucleotide, so you will need to make sure your expression system and delivery methods are compatible with all your components.
Next, consider the best form for those CRISPR components based on your model system. CRISPR reagents can be delivered via transfection, nucleofection, viral transduction or injection as either protein, RNA, or DNA. Finally, once you have identified the best expression system, you can then choose the best method for introducing those CRISPR components into your target cells.
Mammalian CRISPR expression systems
Each model system has its own best practices for efficient delivery of CRISPR components. If you are new to CRISPR or your model system, a good first step would be to consult the literature to see if anyone has published a protocol that would work for your system. Addgene has depositor submitted protocols and links to a CRISPR forum where you may be able to find information regarding your system of choice. The table below summarizes the various components included with each expression system as well as suitable applications.
| Expression System | Components of System | Application |
| Mammalian expression vector | Promoter driving Cas9 expression can be constitutive or inducible. U6 promoter is typically used for gRNA. May contain reporter gene (e.g. GFP) to identify and enrich positive cells or a selection marker to generate stable cell lines. | Transient or stable expression of Cas9 and/or gRNA in a mammalian cell line that can be transfected at high efficiency. |
| Lentiviral transduction | Cas9 and gRNA can be present in a single lentiviral transfer vector or separate transfer vectors. May contain reporter gene (e.g. GFP) to identify and enrich positive cells. Likely contains a selection marker to generate stable cell lines. Packaging and Envelope plasmids provide the necessary components to make lentiviral particles. | Stable, tunable expression of Cas9 and/or gRNA in a wide variety of mammalian cell lines. Useful for difficult to transfect cell types and can be used in vivo. A common choice for conducting genome-wide screens using CRISPR. |
| AAV transduction | Only compatible with SaCas9 (packaging limit ~4.5kb). CRISPR elements are inserted into an AAV transfer vector and used to generate AAV particles. | Transient or stable expression of SaCas9 and/or gRNA. Infects dividing and non-dividing cells. AAV is least toxic method for in vivo viral delivery. |
| Cas9 mRNA and gRNA | Plasmids containing gRNA and Cas9 are used in in vitro transcription reactions to generate mature Cas9 mRNA and gRNA. RNA is delivered to target cells using microinjection or electroporation. | Transient expression of CRISPR components, expression decreases as RNA is degraded within the cell. Can be used for generating transgenic embryos. |
| Cas9-gRNA ribonucleoprotein complexes | Purified Cas9 protein and in vitro transcribed gRNA are combined to form a Cas9-gRNA complex and delivered to cells using cationic lipids. | Transient expression of CRISPR components, expression decreases as gRNA and Cas9 protein are degraded within the cell. |
Delivery methods for mammalian cell lines
As mentioned above, the expression system you choose in many ways dictates the best method for introducing those CRISPR components into your target cells. Delivery of these components into mammalian cell lines is quite broad and includes several different methods, so we've summarized some key features of common delivery methods below:
Chemical transfection
| Lipid mediated | Cationic polymers | Calcium phosphate | |
| Cells | Transformed cell lines | Transformed cell lines | Transformed cell lines |
| Cargo | Nucleic acids or protein | Nucleic acids | Nucleic acids |
| Throughput | High | High | High |
| Efficiency | Medium | Medium | Medium |
| Difficulty | Easy | Easy | Easy |
Physical transfection
| Electroporation / Nucleofection | Microinjection | |
| Cells | Many types | Single cells |
| Cargo | Nucleic acids or protein | Nucleic acids or protein |
| Throughput | High | Very low |
| Efficiency | High | Very high |
| Difficulty | Very easy | Very hard |
Viral delivery
| Lentivirus | Retrovirus | AAV | |
| Cells | Dividing or nondividing | Dividing | Dividing or nondividing |
| Cargo | Nucleic acids | Nucleic acids | Nucleic acids |
| Throughput | Low | Low | Low |
| Efficiency | Variable | Variable | Variable |
| Difficulty | Hard | Hard | Hard |
These tables are not inclusive of all methods, nor are these methods limited exclusively to in vitro cell culture. The user should review the current literature about their preferred model.
If your expression system is not well characterized in terms of CRISPR use, you will want to invest some time in optimizing and testing the efficiency of CRISPR delivery. There are a few plasmids at Addgene that have been published as CRISPR testing tools:
Traffic Light Reporter System: Can evaluate lentiviral component delivery as well as genome repair by non-homologous end joining (NHEJ) or HDR.
EGFP validation of sgRNAs: Can evaluate component delivery and sgRNA efficacy by cloning in genome target sequence into EGFP reporter.
Target DNA reporter system: Can evaluate component delivery with validated tools.
Special thanks to Joel McDade for creating the Expression Systems table. Marcy Patrick contributed to the writing of this post.
Additional Resources on the Addgene Blog
- Check out our CRISPR Featured Topic Page
- Learn about CRISPR Delivery Using Ribonucleoproteins (RNPs)
- Read Tips for a 1st Time CRISPR User (by a 1st Time CRISPR User)
Resources on Addgene.org
- Browse CRISPR Plasmids for Mammalian Systems
- Check out our CRISPR Guide Pages
- View CRISPR Protocols
Topics: CRISPR, CRISPR 101, CRISPR Expression Systems and Delivery Methods






Leave a Comment