By Baiyang Liu and James Chappell, Rice University.
For decades, we’ve been designing experiments around two major limitations of plasmids: copy number and incompatibility. While functional, such workarounds are clunky. To address this, we created a synthetic origin of replication (ORI) that allows us to modify the plasmid instead of the experiment.
 |
|
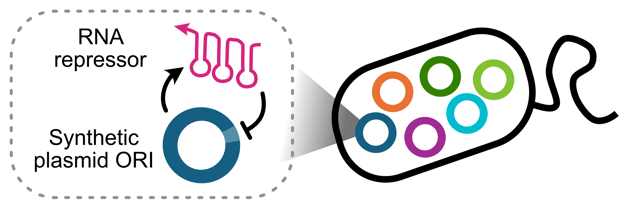
Figure 1: Simplified schematic of SynORI. |
A modular, synthetic ORI
In our paper, we rebuilt the classic pMB1 origin of replication into a modular, programmable origin we call SynORI. Instead of relying on the native ColE1-style feedback loops, SynORI uses synthetic RNA regulators to control replication. Additionally, we refactored the natural system to create a modular architecture that lets researchers plug in synthetic regulatory RNAs to create plasmids with new compatibility groups and tunable copy number.
Expanding plasmid compatibility
A major barrier to multi-plasmid systems is that most plasmids fall into a small number of incompatibility groups, and members of the same group cannot be used in the same cell at the same time. Starting from pMB1, we used a previously-developed orthogonal regulatory RNA library to build six compatible SynORI plasmids. All six could be co-transformed into E. coli in a single reaction and maintained for at least a week. Theoretically, large number of compatible ORIs can be created this way with orthogonal regulatory RNAs designed computationally.
Copy number control
Beyond compatibility, SynORI makes copy number easily tunable rather than a constraint. Because the SynORI plasmids are modular, we can swap elements to dial copy number up or down across a wide range. For example, we achieved a 115-fold change in copy number (from ~1.6 to ~185 copies per cell). We also show that ligand-responsive genetic elements, namely transcription factors and riboswitches, can be used to dynamically control copy number.
Eliminating multimers
Plasmids in the ColE1 family also suffer from a practical headache: they can form multimers, which reduces stability and makes maintenance less reliable. To address this, we added parCBA, a multimer-resolution system adapted from the plasmid RK2. This reduces multimer formation and improves the stability of SynORI plasmids.
Experimental limitations of synthetic ORIs
While we were successful in creating SynORI, an interesting observation was that their stability in the absence of antibiotic selection was reduced compared to the natural pMB1. The underlying cause remains unclear, but the result suggests that refactoring the origin may have compromised a stability feature. As a result, it is important to maintain selective pressure when using SynORI.
Additionally, though the SynORI plasmid no longer uses the naïve negative feedback regulation from ColE1 family, it still contains a truncated RNA primer that may interact with the antisense RNA when co-transformed with ColE1 plasmids. As such, we recommend using SynORIs together with ORIs that are compatible with ColE1 family, such as p15A, CDF, and pSC101.
Interested in using SynORI in your experiment? Find the plasmids here!
This post was written by Baiyang Liu and James Chappell at Rice University, whose team developed SynORI.
Additional resources
Resources on the Addgene blog
- Plasmids 101: Origin of Replication
- Plasmids 101: Plasmid Incompatibility
- Plasmids 101: Dimers and Multimers
Resources on addgene.org
- Watch our video on the Origin of Replication
- Read our Plasmids 101 ebook
- Get practical molecular biology help with our Molecular Biology Reference Pages
Topics: Plasmid Elements, Plasmids





Leave a Comment