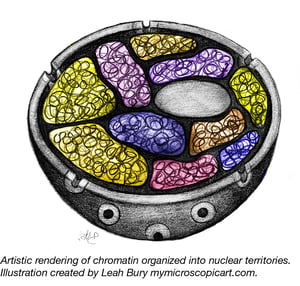
 How many times have you looked at a diagram depicting transcription, or DNA repair, or replication, or any number of CRISPR applications and thought “OK, but how does this work in the context of chromatin?” Though it’s true that adding histones and chromatin architecture to every diagram portraying some aspect of eukaryotic DNA would become busy and potentially detract from the process being depicted, we can’t forget that those other components are still there in real life. Certainly referring to any DNA within the context of a nucleus as “linear” is a misnomer. The DNA packed into our chromosomes is very much 3-dimensional, as you can see by this post’s chromatin illustrations from Leah Bury of Microscopic Art.
How many times have you looked at a diagram depicting transcription, or DNA repair, or replication, or any number of CRISPR applications and thought “OK, but how does this work in the context of chromatin?” Though it’s true that adding histones and chromatin architecture to every diagram portraying some aspect of eukaryotic DNA would become busy and potentially detract from the process being depicted, we can’t forget that those other components are still there in real life. Certainly referring to any DNA within the context of a nucleus as “linear” is a misnomer. The DNA packed into our chromosomes is very much 3-dimensional, as you can see by this post’s chromatin illustrations from Leah Bury of Microscopic Art.
I started thinking about this problem when I recently attended the 2018 Chromosome Architecture and Chromosome Organization Keystone Symposium. As it happens, there are a lot of scientists who not only remember to think about DNA in this context, but also specifically study it. They might be researching enhancer elements that control the transcription of a gene a megabase away, or looking at the histone methylation status at a double-strand break that needs repairing. They may be looking at chromatin at the level of nucleosomes, loops, chromosomal domains, or how stretches of DNA on different chromosomes interact with each other.
Chromatin regulation is complex and influences--directly or indirectly--many other areas of biological research. If you’re looking for resources that will help you study how chromatin effects your favorite biological phenomenon, there’s a reasonable good chance that Addgene can help.
Though it would be difficult to consolidate every conceivable chromatin-related plasmid in a collection that grows on a daily basis, the lists below provide a snapshot of relevant resources and a good jumping off point. The following lists of PIs, articles, genes, and blog posts are not comprehensive, so just because you do not see a name or gene on this list, that does not necessarily mean the repository does not have it. That said, if you know of a plasmid that we definitely don’t have that you really want, please let us know! And if you study chromatin and want to start sharing your plasmids through Addgene, don’t hesitate to start the process!
 Addgene depositors who study chromatin
Addgene depositors who study chromatin
The PIs linked below are partially curated from the presentations and posters I saw at the 2018 Keystone Chromosome Architecture and Chromosome Organization. These aren’t the only investigators in the chromatin field who have deposited with Addgene, but you’ll find many great tools by clicking on the links below and visiting their associated PI pages on the Addgene website.
| In-exhaustive List of Addgene Depositors Who Study Chromatin | |||
| Cheryl Arrowsmith | James Bradner | Kerstin Bystricky | Courtney Hodges |
| Sharon Dent | Archa Fox | Elaine Fuchs | Kai Ge |
| Robert Kingston | Thoru Pederson | Phil Sharp | Julie Ahringer |
| Alexander Stark | Bas van Steensel | Richard Young | Yi Zhang |
| Leonard Zon | You! | ||
Chromatin-related genes with plasmids available from Addgene
The links below pull up lists of plasmids containing the indicated gene, and you can subscribe to these pages to learn when Addgene receives new plasmids containing these genes. As with the PIs above, many of these genes were discussed at the 2018 Keystone meeting for Chromatin. Note that histone genes frequently have many different variant names, not all of which can be listed here. If you don’t see the gene you’re looking for on this list, please consider searching directly on our website.
| In-exhaustive Chromatin-related genes with plasmids on Addgene.org | |||
| AGO1 | BRCA1 | CREBBP | CUL1 |
| H2AFX | H2AFY | H3F3A | H3.1 |
| H3.3 | HIST1H1E | HIST1H2BB | HIST2H2BE |
| RBX1 | STAG2 | Xist | Your favorite gene! |
| SWI/SNF Genes: | |||
| BAF170 (SMARCC2) | ARID1A | ||
Chromatin research articles with plasmids available from Addgene
Many articles reporting chromatin-related research describe the production of plasmids that might be useful for your work. The links below direct you to pages containing lists of plasmids from the indicated publications. You can request all of these plasmids directly from Addgene. Note that there are plasmids from hundreds of articles in our collection; the following articles are just a small cross-section spanning 20 years of research.
- DNA Cross-Bridging Shapes a Single Nucleus from a Set of Mitotic Chromosomes. Samwer M, Schneider MWG, Hoefler R, Schmalhorst PS, Jude JG, Zuber J, Gerlich DW Cell. 2017 Aug 24;170(5):956-972.e23
- Screen identifies bromodomain protein ZMYND8 in chromatin recognition of transcription-associated DNA damage that promotes homologous recombination. Gong F, Chiu LY, Cox B, Aymard F, Clouaire T, Leung JW, Cammarata M, Perez M, Agarwal P, Brodbelt JS, Legube G, Miller KM Genes Dev. 2015 Jan 15;29(2):197-211.
- DNA Cross-Bridging Shapes a Single Nucleus from a Set of Mitotic Chromosomes. Samwer M, Schneider MWG, Hoefler R, Schmalhorst PS, Jude JG, Zuber J, Gerlich DW Cell. 2017 Aug 24;170(5):956-972.e23
- Rapid and phosphoinositol-dependent binding of the SWI/SNF-like BAF complex to chromatin after T lymphocyte receptor signaling. Zhao K, Wang W, Rando OJ, Xue Y, Swiderek K, Kuo A, Crabtree GR Cell. 1998 Nov 25. 95(5):625-36.
- An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Clemson CM, Hutchinson JN, Sara SA, Ensminger AW, Fox AH, Chess A, Lawrence JB Mol Cell. 2009 Mar 27;33(6):717-26.
- Reporter-nanobody fusions (RANbodies) as versatile, small, sensitive immunohistochemical reagents. Yamagata M, Sanes JR Proc Natl Acad Sci U S A. 2018 Feb 13. pii: 1722491115.
Addgene blog posts related to chromatin
This is not the first Addgene blog post about chromatin--multiple Addgene colleagues and guest bloggers have written posts describing new methods and tools for studying chromatin and the epigenome. If you have been considering some new techniques, the posts below may steer you in the right direction!
CAPTURE-ing Chromatin Interactions: Using CRISPR-dCas9 to Study Gene Regulation
CRISPR 101: Epigenetics and Editing the Epigenome
CUT&RUN: An Improved Method for Studying Protein-DNA Interactions
CRISPRainbow and Genome Visualization
CRISPR Between the Genes: How to Experiment with Enhancers and Epigenomics
I would like to thank the many scientists with me at the Keystone Chromatin meeting, especially Karen Reddy (@theReddyLab), Thoru Pederson, Joanna Wysocka, Kerstin Bystricky, Ben Martin, and Berkley Gryder. Addgene depositors or not, your time is valuable
Topics: Other Plasmid Tools, Plasmids





Leave a Comment