This post was contributed by Richi Sakaguchi from Kyoto University, and Marcus N. Leiwe and Takeshi Imai from Kyushu University.
Stochastic multicolor labeling is a powerful solution for discriminating between neurons for light microscopy-based neuronal reconstruction. To achieve stochastic multicolor labeling, Brainbow used the Cre-loxP system to express one of the three fluorescent protein (XFP) genes in a transgene. When multiple copies of the transgene cassette are introduced, stochasticity will result in a combinatorial expression of these three genes with different copy numbers, producing dozens of color hues (Livet et al., 2007; Cai et al., 2013). However, the brightness of Brainbow was inherently low. This is because the stochastic and combinatorial expression of fluorescent proteins is only possible at low copy number ranges, resulting in low fluorescent protein level.
The principle of Tetbow
In modern neuroscience, plasmid or viral vector-mediated gene delivery is more common than conventional transgenic strategies. In such vector-based gene delivery systems, the stochastic expression of XFPs is possible by simply introducing a mixture of XFP genes. Stochastic multicolor labeling is possible when each of the XFP genes is introduced at ~2 copies per cell per color, following a Poisson distribution. To overcome the limited expression levels of XFPs in the original method, we used the Tetracycline trans-activator system to boost the expression levels, and we named the system Tetbow (Tetracycline trans-activator Brainbow) (Sakaguchi et al., 2018) (Figure 1). Once introduced, fluorescent proteins were best imaged after tissue clearing with SeeDB2 (Ke et al., 2016, 2018).
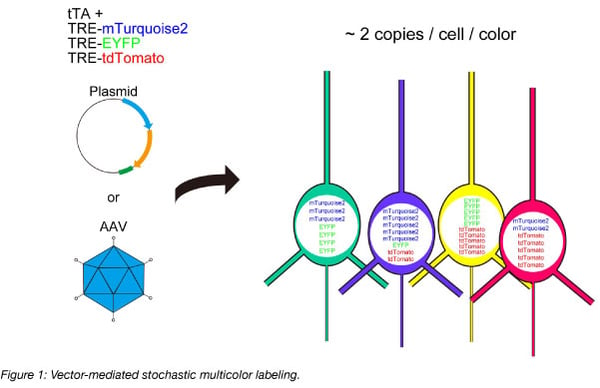
How to use Tetbow for neuronal labeling
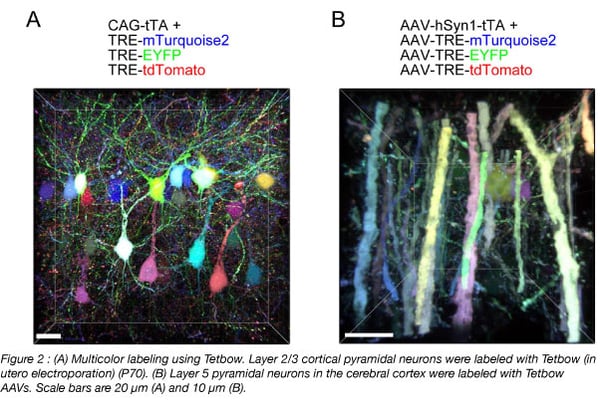
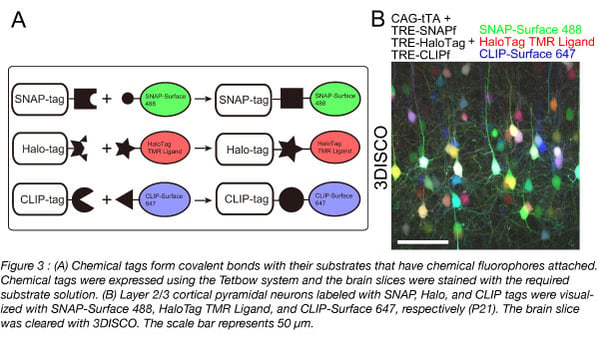
To use Tetbow, you can introduce the four Tetbow plasmids (Addgene #104102 - #104105) using in utero electroporation or with AAVs (prepared with Addgene #104110 - #104112) (Figure 2). We’ve also created constructs with containing chemical tags such as the SNAP-tag, Halo-tag, and CLIP-tag (Kohl et al., 2014) (Addgene #104106 - #104108) (Figure 3) that can be introduced using in utero electroporation. Then at your preferred time points, sacrifice the animals and fix the samples. When you use AAVs, it typically takes a few weeks to achieve optimal expression of XFPs. Any longer and the cells may start to show morphological abnormalities due to the excessive amount of XFPs expressed. Then, clear the samples and visualize.
For more details on this procedure, find the protocol here.
Tips and troubleshooting your Tetbow experiment
- Plasmid concentrations and AAV titer - The XFP genes have to be introduced at ~2 copies per cell per color to achieve the highest color variations. Adjust the plasmid concentrations or AAV titer for the best results.
- The timing of sacrifice when using Tetbow AAVs - You will have to wait a few weeks to achieve an adequate expression level of the Tetbow AAVs. However, you should not wait too long, as too much XFP is toxic to the cells.
- Inadequate expression levels of XFPs - Reduce the amount of tTA. Paradoxically, too much tTA leads to a reduced expression level of XFPs, most likely by suppressing transcription. It is critical to express a minimal amount of tTA to achieve highest expression levels.
- Clearing large tissues - Fluorescent proteins are very stable in SeeDB2. However, SeeDB2 is not powerful enough for large brain samples. For large brain samples, we recommend pretreatment with ScaleCUBIC1 (Susaki et al., 2014), before clearing with SeeDB2.
For more information, visit our SeeDB Resources website. We update the site regularly and hope Tetbow can help you in your research.
To see Tetbow in action, watch the video below depicting dendrite wiring of mitral cells.
Many thanks to our guest bloggers Richi Sakaguchi from Kyoto University, and Marcus N. Leiwe and Takeshi Imai from Kyushu University.
 Richi Sakaguchi is a graduate student at Kyoto University. He is interested in developing genetic tools for neuronal circuit tracing.
Richi Sakaguchi is a graduate student at Kyoto University. He is interested in developing genetic tools for neuronal circuit tracing.
 Marcus Leiwe is an assistant professor at Kyushu University. He has a particular interest in the development of sensory systems focusing on the functional and anatomical maturation of the olfactory system.
Marcus Leiwe is an assistant professor at Kyushu University. He has a particular interest in the development of sensory systems focusing on the functional and anatomical maturation of the olfactory system.
 Takeshi Imai is a professor at Kyushu University. His lab has been studying functional and anatomical aspects of sensory circuit development using light microscopy. Lab website: https://dn-en.med.kyushu-u.ac.jp/
Takeshi Imai is a professor at Kyushu University. His lab has been studying functional and anatomical aspects of sensory circuit development using light microscopy. Lab website: https://dn-en.med.kyushu-u.ac.jp/
References
Cai, Dawen, et al. "Improved tools for the Brainbow toolbox." Nature methods 10.6 (2013): 540. PubMed PMID: 23817127. PubMed Central PMCID: PMC3713494.
Ke, Meng-Tsen, et al. "Super-resolution mapping of neuronal circuitry with an index-optimized clearing agent." Cell reports14.11 (2016): 2718-2732. PubMed PMID: 26972009.
Ke, Meng-Tsen, and Takeshi Imai. "Optical clearing and index matching of tissue samples for High-resolution fluorescence imaging using SeeDB2." Bio-Protocol 8 (2018): e3046.
Livet, Jean, et al. "Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system." Nature 450.7166 (2007): 56. PubMed PMID: 17972876.
Kohl, Johannes, et al. "Ultrafast tissue staining with chemical tags." Proceedings of the National Academy of Sciences111.36 (2014): E3805-E3814. PubMed PMID: 25157152. PubMed Central PMCID: PMC4246963.
Sakaguchi, Richi, Marcus N. Leiwe, and Takeshi Imai. "Bright multicolor labeling of neuronal circuits with fluorescent proteins and chemical tags." eLife 7 (2018): e40350. PubMed PMID: 30454553. PubMed Central PMCID: PMC6245733.
Susaki, Etsuo A., et al. "Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis." Cell 157.3 (2014): 726-739. PubMed PMID: 24746791.
Additional resources on the Addgene blog
- Read about Brainbow
- Learn about using AAVs for neuronal tracing
- Visit the viral vectors blog topic page
Additional resources on Addgene.org
- Learn about Addgene's viral service
- See our fluorescent protein collection
- Find AAVs at Addgene
Topics: Fluorescent Proteins, Cell Tracing






Leave a Comment