 In our first few Plasmids 101 posts, we focused mainly on the elements required for plasmid maintenance within an E. coli cell, but vectors can be widely utilized across many different cell types and each one requires different elements for vector propagation. This post, along with a future companion post on mammalian vectors, will catch you up on the core replication and resistance features of yeast vectors and explain how they differ from the bacterial elements previously described.
In our first few Plasmids 101 posts, we focused mainly on the elements required for plasmid maintenance within an E. coli cell, but vectors can be widely utilized across many different cell types and each one requires different elements for vector propagation. This post, along with a future companion post on mammalian vectors, will catch you up on the core replication and resistance features of yeast vectors and explain how they differ from the bacterial elements previously described.
Why do scientists use yeast vectors?
Yeast are eukaryotes and thus contain complex internal cell structures similar to those of plants and animals. Unlike bacteria, yeast can post-translationally modify proteins yet they still share many of the same technical advantages that come with working with prokaryotes. This includes but is not limited to: rapid growth, ease of replica plating and mutant isolation, a well-defined genetic system, and a highly versatile DNA transformation system.Unlike most other microorganisms, yeast have both a stable haploid and diploid state which is useful for genetic analysis, as well as an efficient mechanism of homologous recombination to facilitate simple gene replacement/mutation. Yeast expression plasmids used in the lab typically contain all the necessary components to allow shuttling between E. coli and yeast cells. To be useful in the lab, the vectors must contain a yeast-specific origin of replication (ORI) and a means of selection in yeast cells, in addition to the bacterial ORI and antibiotic selection markers.
The yeast origin of replication
(Please note: This first section primarily pertains to ORIs in budding yeast, Saccharomyces cerevisiae; however, we’ve also noted some features required for the replication of fission yeast, Schizosaccharomyces pombe, vectors at the end.)
In a previous post, we briefly discussed how the regulation of bacterial ORIs determines plasmid copy number within the bacterial cell. Similarly, the specific ORI elements included within a yeast vector determine how the plasmid is replicated and maintained within the yeast cell. These elements control not only the number of plasmids found in each cell, but also whether the plasmid gets integrated into the host DNA or is independently replicated as an episome. The four main types of yeast plasmids are defined below:
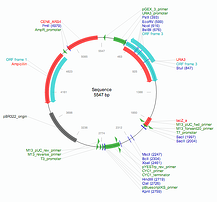
- Yeast Integrating plasmids (YIp): These plasmids lack an ORI and must be integrated directly into the host chromosome via homologous recombination.
- Yeast Replicating plasmids (YRp): These vectors contain an Autonomously Replicating Sequence (ARS) derived from the yeast chromosome. As the name suggests, these vectors can replicate independently of the yeast chromosome; however, they tend to be unstable and may be lost during budding.
- Yeast Centromere plasmids (YCp): These are considered low copy vectors and incorporate part of an ARS along with part of a centromere sequence (CEN). These vectors replicate as though they are small independent chromosomes and are thus typically found as a single copy. Unlike the ARS vectors, CEN vectors are stable without integration.
- Yeast Episomal plasmids (YEp): These are most similar to bacterial plasmids and are considered “high copy”. A fragment from the 2 micron circle (a natural yeast plasmid) allows for 50+ copies to stably propagate per cell. The copy number of these vectors can also be controlled if specific regulatable elements are included (reviewed here).
Plasmids for use in S. pombe, on the other hand, do not require a well defined ORI. Instead, the size and A-T content of the DNA (apparently independent of a known specific sequence) dictate the replication of these vectors. S. pombe plasmids oftentimes utilize an ARS to aid in high transformation efficiency; however, this region does not necessarily promote replication.
Selection markers for yeast
Historically, scientists have utilized auxotrophic selection rather than antibiotic selection when working with yeast, due to high rates of spontaneously occurring resistant mutants and the insensitivity of yeast strains to some antibiotics. Auxotrophy is defined as the inability of an organism to synthesize a particular organic compound required for its growth. Many auxotrophic strains of yeast exist which can be easily maintained when grown on media containing the missing nutrient. Scientists can exploit these host mutations by including a copy of a functional gene which complements the host’s auxotrophy. When grown on media NOT containing the nutrient, the host cells will die unless they have incorporated the plasmid carrying the required gene.
The table below lists some of the most commonly used selection markers in yeast and provides the element needed to overcome the auxotrophy as well as additional uses for said element. This link provides a more extensive list of yeast auxotrophic markers and includes the associated references.
| Yeast selection marker | What does the marker help synthesize? |
Counterselection? (growth-based positive selection for the loss of the marker gene) |
For use in: | Can this be used in other yeast species? | Can this be used for auxotrophic selection in E. coli? | Additional considerations |
| HIS3 | L-hisitidine | no | S. cerevisiae | no | yes | |
| URA3 | pyrimidine (uracil) | yes - Grow with 5-FOA. | S. cerevisiae | yes - This can complement ura4- S. pombe, but the complementation is weak. | yes | |
| LYS2 | L-lysine | yes - Grow on plates containing alpha-aminoadipate in the absence of a nitrogen source. | S. cerevisiae | no | no | |
| LEU2 | L-leucine | no | S. cerevisiae | yes - This can complement leu1- S. pombe, but multiple copies are required. | yes | |
| TRP1 | L-tryptophan | yes - Grow with 5-FAA. | S. cerevisiae | no | no | TRP1 alters some yeast phenotypes. This marker should not be used in gene disruption experiments. |
| MET15 | L-methionine and overproduces hydrosulfide ions | yes - Grow with methylmercury. | S. cerevisiae | no | no | Can be used for color and growth selection if divalent lead ions are used in the growth media. |
| ura4+ | pyrimidine (uracil) | yes - Grow with 5-FOA*. | S. pombe | no | no | *FOA in fission yeast induces mutation in the ura5+ gene in addition to ura4+. |
| leu1+ | L-leucine | S. pombe | no | no | ||
| ade6+ | purine (adenine) | S. pombe | no | no |
Considerations when using auxotrophic selection
Of course, there are some drawbacks to using auxotrophic markers as a means of selection:
- A specific selection marker needs to be used with a yeast strain deficient in that compound. Therefore known auxotrophic strain/ selection element pairs must be utilized or a new combination needs to be created in advance of the experiment.
- The marker provided by the plasmid may be expressed at higher than normal physiological levels due to high copy numbers. This creates a potential metabolic burden on the yeast cells.
- Some phenotypes may be altered due to the presence of the selection marker at non-physiological levels.
Scientists have tried varied approaches to combat these issues. One method to reduce the amount of marker gene expression is to use a partially defective promoter to drive expression of the selection marker. This reduces the amount of gene product present in the cell, thus allowing the yeast to maintain higher copy numbers. Additionally, improvements in antibiotic selection have made utilizing the more traditional drug selection methods feasible in yeast as a complement or alternative to using auxotrophic markers. The MX series of antibiotic resistance cassettes is most commonly employed, with the KanMX being the most prevalent due to its versatility and ability to be used in bacterial, yeast, and mammalian cells.
Topics: Yeast, Plasmids 101, Plasmids







Leave a Comment