We often think of DNA as inert. It generally stays put, which makes it easy to locate in a genome. But there is a type of mobile DNA, called a transposon, that’s a bit hyperactive and likes to jump around from one location in the genome to another. This jumping is what caught the eye of Barabara McClinntock who first discovered transposons due to changes in coloration of corn kernels. Aside from corn, transposons are in the genomes of many other prokaryotes and eukaryotes, and make up ~45% of the human genome (Munoz-Lopez et al., 2010).
So why do transposons matter? The outcome really depends on where a transposon lands. For example, in bacteria, transposons can move antibiotic resistance genes between plasmids or from plasmids to the bacteria’s genome. In humans, if a transposon jumps into a gene, it could be mutagenic and result in human disease. While transposon insertions can be detrimental, it also drives genomic evolution and is used as a tool by scientists to move DNA around in the lab.
Let’s take a look at the key components of transposons, how they are classified, and how they are used in the lab.
What is a transposon?
DNA transposons are mobile, repetitive genetic elements that are found in both prokaryotic and eukaryotic genomes.
Components of a transposon system
- Transposon: The transposon is the DNA sequence that moves around. This DNA sequence encodes the proteins the transposon needs for transposition.
- Transposition protein: This is the protein that’s required for the transposon sequence to move from one DNA site to another. This protein is either a transposase or a reverse transcriptase, depending on the mechanism of transposition.
- Target site: Different transposons insert at different DNA sequences or target sites. Integration of most transposable elements (TEs) results in the duplication of this target site sequence at the site of insertion.
Types of transposons
Transposable elements are split into two classes based on their mechanism of transposition: Class I TEs, also called retrotransposons, and Class II TEs, also called DNA transposons. DNA transposons are commonly used as tools in the lab, so we’ll focus on DNA transposons for much of the rest of this article.
Class I Transposable elements (TEs): Retrotransposons
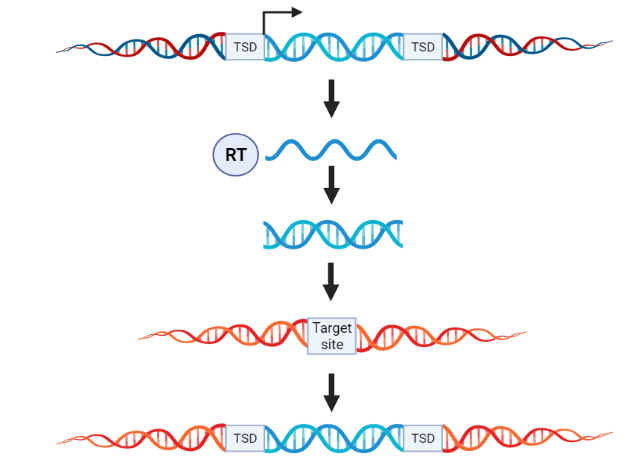
Class I TEs are also known as retrotransposons. They transpose by a ‘copy-paste’ mechanism (Fig. 1). They first copy themselves as RNA transcripts and then rely on a reverse transcriptase enzyme to reverse transcribe back into DNA before being inserted into new target sites. This is similar to how retroviruses, like HIV, replicate. Class I TEs do not encode a transposase enzyme.
Class I TEs are considered replicative since every time they jump they make a copy of themselves. This increases the number of copies of the TE while also increasing the size of their host’s genome.
There are two types of Class 1 TEs: those with long terminal repeats (LTRs) and those without (non-LTR TEs). LTR retrotransposons are like retroviruses both in their structure and mechanism of replication. They contain two genes: gag and pol. The pol polyprotein encodes the reverse transcriptase and integrase enzymes which are required for LTR retrotransposon transposition. Non-LTR retrotransposons contain two open reading frames (ORFs) which often terminate with a poly(A). ORF2 encodes endonuclease and reverse transcriptase activities.
 |
|
Figure 1: Overview of retrotransposon transposition. Retrotransposons are mobilized by a ‘copy-paste’ mechanism. First they copy themselves as RNA transcripts and then use their reverse transcriptase (RT) enzyme to convert back into DNA. Then they can be inserted into new target sites. Created in BioRender.com. |
Class II TEs: DNA transposons
Class II TEs are also known as DNA transposons, since they do not use an RNA intermediary when they move. Most Class II transposons have a non-replicative “cut and paste” mechanism of transposition: they first excise themselves from one location and then insert somewhere else (Fig. 2). Class II TEs have LTRs on both of their ends.
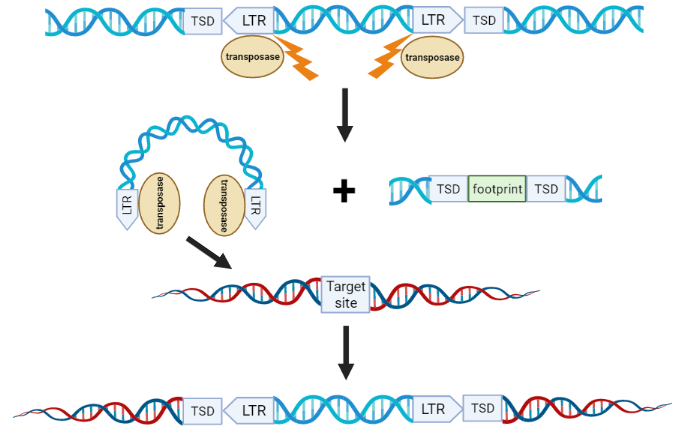 |
| Figure 2: Overview of DNA transposon transposition. Modified from Sandoval-Villegas et al., 2021. To mobilize a transposon, first the transposase enzyme binds the transposon’s long terminal repeats (LTRs), induces a double-stranded break, and excises the transposon from the donor DNA. A DNA footprint is left behind. When the transposon-transposase complex finds its target site, it integrates, which produces a target site duplication (TSD). Created in BioRender.com. |
Autonomous vs. non-autonomous TEs
You can think of autonomous transposons as “complete transposons” since they encode the protein they need to move around, either a transposase or a reverse transcriptase. Non-autonomous transposons, however, need another TE to express a transposase or reverse transcriptase so they can transpose. Both class I and II transposons can be autonomous or non-autonomous.
DNA transposons commonly used in the lab
While there are many different types of transposons, DNA transposons are most commonly used in the laboratory for genome manipulation. When transposons are used in the lab, the transposase gene is provided in trans so that a gene of interest can be inserted between the transposon’s LTRs, similar to when packaging viral vectors.
Let’s take a closer look at three different transposon systems adapted for use as research tools: Sleeping beauty, PiggyBac, and Tol2.
Sleeping beauty
Sleeping Beauty is a synthetic transposable element developed from inactivated Tc1/mariner transposons found in fish (Sandoval-Villegas et al., 2021). Sleeping Beauty’s preferred target site for integration is TA dinucleotides and it leaves behind the CAG DNA footprint from its terminal sequences at the excision site after cleavage by the transposase. It has a cargo capacity >100 kB, although integration efficiency decreases with cargo size. Sleeping Beauty has a close-to-random integration profile in mammalian genomes. SB is active in vertebrates and integrates in human cells at rates similar to retroviral vectors. The hyperactive version of the SB transposase, SB100X, is ~100-fold more efficient compared to first generation SB transposase (Mátés et al., 2009). hySB100x improves on SB100X by having 30% higher transposition activity (Voigt et al., 2016).
piggyBac
Although its name suggests otherwise, piggyBac was discovered in the cabbage looper moth (Potter et al., 1976). Its target site is TTAA and unlike other transposons, it does not leave behind a DNA footprint sequence after excision. piggyBac can mobilize DNA over 100 kB in size and is active in vitro as well as in vivo in yeast, plant, insect, and mammalian cells, including human cells. piggyBac is biased to integrate at transcription start sites, CpG islands, and DNaseI hypersensitivity sites. Like Sleeping Beauty, piggyBac has an integration efficiency in human cells similar to retroviral vectors. The hyperactive PB transposase (hyPB) is ~10-fold more active in mammalian cells compared to the codon-optimized wild-type piggyBac transposase.
Tol2
Tol2 was the first reported active DNA transposon in vertebrates. It was discovered in Japanese medaka fish because its insertion into the fish’s tyrosinase gene caused albinism (Koga et al., 1996). Unlike Sleeping Beauty and piggyBac, Tol2 has a weak consensus sequence for its preferred target integration site: TNA(C/G)TTATAA(G/C)TNA. Tol2 can deliver 10-11 kB to mammalian cells without decreased efficiency, with a maximum cargo capacity of ~200 kB of DNA. Similar to piggyBac, Tol2 also prefers to integrate at transcription start sites, CpG islands, and DNaseI hypersensitivity sites. Tol2 is only active in vertebrates and has lower efficiency of integration in human cells than piggyBac and Sleeping Beauty. Minimal Tol2 or miniTol2 is a truncated version of the original Tol2 and has ~3-fold increase in transposition activity
Applications of transposons
Now that you’ve learned about some of the popular transposon systems, let’s take a look at how they can be used in the lab.
Transposon mutagenesis screens
Transposons, by their nature, are mutagenic elements, which makes them a great tool for mutagenesis screens that detect loss-of-function or gain-of-function mutations. In these screens, the transposons encode reporter genes, mutagenic cassettes, or barcodes. When transposons are delivered to cells or a model organism, they insert into the host’s genome. Then, the transposon insertion sites are detected with next generation sequencing, and analyzed to identify which insertions were positive or negative selected for during the experiment (Sandoval-Villegas et al., 2021). Target site preference and host range are important to consider when selecting a transposon for a mutagenesis screen. For example, piggyBac and Tol2 are best for screening promoters and enhancers since they are biased to insert at these sites.
Transgenic animals
Transgenic animals are often generated by directly injecting DNA into the pronuclei of fertilized eggs, which leads to random incorporation of that sequence into the zygote’s genome, a process that’s highly unpredictable. Transposons, however, efficiently incorporate into the zygote’s genome after injection into the cytoplasm of a fertilized egg, a process which is inefficient when DNA is injected. Sleeping Beauty, piggyBac, and Tol2 have all been used to generate transgenic animals, including zebrafish, mice, rats, and rabbits (Sandoval-Villegas et al., 2021).
Gene transfer
Transposons are an alternative to viral vector delivery of transgenes, like for iPSC reprogramming and gene and cell therapy, and have potential to overcome some of the limitations of viruses. TEs have a large payload, up to 100 kB with Sleeping Beauty and piggyBac, which is a big advantage over viral vectors (~5 kB payload for AAV and ~8 kB payload for lentivirus). Large cargos, such as the ~11 kB cDNA for dystrophin, the gene mutated in muscular dystrophy, require truncation to fit in viral vectors, but easily fit in a TE. Transposons are also less likely than viral vectors to induce an immune response and are also easier and cheaper to generate. Gene disruption due to integration can occur for both delivery methods, but since TEs insert mostly in intergenic regions, gene disruption is less of a concern.
RNA guided transposons insertion
One drawback to DNA transposons is that while they have a specific integration target site, such as TTAA for Sleeping Beauty, they can insert at any number of these sites found in the genome.
To target transposons to a single location in bacterial genomes, the Sternberg lab developed the INTEGRATE (Insertion of Transposable Elements by Guide RNA-Assisted Targeting) system. INTEGRATE is based on a naturally occurring Tn7 transposon found in Vibrio cholerae that encodes a Type I-F CRISPR-Cas system. This system has four major components: 1) a CRISPR RNA (crRNA), 2) four proteins (TniQ, Cas8, Cas7, Cas6) that forms the QCascade DNA-targeting module with the crRNA, 3) a transposase complex made up by three transposase proteins (TnsA, TnsB, TnsC), and 4) the donor DNA or mini-transposon that contains the DNA cargo of interest flanked by ~50-150 bp transposon end sequences (Fig. 3A).
For the INTEGRATE system to transpose, first the QCascade complex surveys the genome and binds its target DNA sequence. The transposase complex binds the mini-transposon, excises it from the donor molecule, and inserts it ~50 bp downstream of QCascade’s genomic target site (Fig. 3B). The INTEGRATE system can achieve ~100% integration of DNA up to 10 kB in size in bacteria.
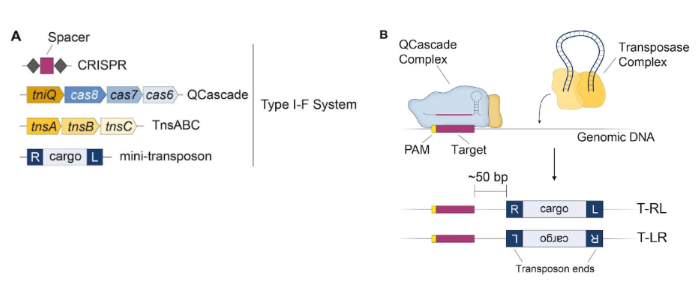 |
| Figure 3: INTEGRATE system for RNA guided transposon insertion in bacterial genomes. A) Components of the INTEGRATE system. B) Mechanisms of RNA-guided transposition with INTEGRATE. The QCascade complex surveys the genome and binds its target DNA sequence. The transposase complex binds the mini-transposon, excises it from the donor molecule, and inserts it ~50 bp downstream of QCascade’s genomic target site. T-RL= transposon right-left orientation. T-LR= transposons left-right orientation. Image from Leo Vo. |
TL;DR
Transposons are another tool to move DNA around or into eukaryotic and prokaryotic genomes. They require just two key components, the transposon’s DNA sequence and its transposition protein, to move DNA. There are three popular systems for use in the lab (Sleeping Beauty, piggyBac, and Tol2), each of which has several different applications (mutagenesis screens, transgenic animals, gene transfer). The discovery of RNA guided transposons in bacteria sets the stage for combining the super powers of CRISPR and transposons to allow for the targeted delivery of large DNA cargos (>100 kB) to specific locations in the genome.
References and Resources
References
Feschotte C, Pritham EJ (2007) DNA Transposons and the Evolution of Eukaryotic Genomes. Annu Rev Genet 41:331–368. https://doi.org/10.1146/annurev.genet.40.110405.090448
Kumar A (2020) Jump around: transposons in and out of the laboratory. F1000Res 9:135. https://doi.org/10.12688/f1000research.21018.1
Munoz-Lopez M, Garcia-Perez J (2010) DNA Transposons: Nature and Applications in Genomics. CG 11:115–128. https://doi.org/10.2174/138920210790886871
Pray, L. (2008) Transposons: The jumping genes. Nature Education 1(1):204. https://www.nature.com/scitable/topicpage/transposons-the-jumping-genes-518/
Sandoval-Villegas N, Nurieva W, Amberger M, Ivics Z (2021) Contemporary Transposon Tools: A Review and Guide through Mechanisms and Applications of Sleeping Beauty, piggyBac and Tol2 for Genome Engineering. IJMS 22:5084. https://doi.org/10.3390/ijms22105084
Additional resources on the Addgene blog
- Learn about the INTEGRATE system for bacterial genome engineering
- Read about the Sleeping Beauty DNA transposon
- Learn about the piggyBac DNA transposon
Topics: Plasmids 101






Leave a Comment