Promoters control the binding of RNA polymerase and transcription factors. Since the promoter region drives transcription of a target gene, it therefore determines the timing of gene expression and largely defines the amount of recombinant protein that will be produced. Many common promoters. like CMV, EF1A, and SV40 promoters, are always active and thus referred to as constitutive promoters. Others are only active under specific circumstances. In this post, we’ll discuss inducible promoters, which can be switched from an OFF to an ON state, and how you might use these in your research. When you're done with this post, check out our follow up post on repressible promoters.
 How are inducible promoters regulated?
How are inducible promoters regulated?
Inducible promoters can be regulated by positive or negative control.
Positive inducible
In the OFF state, the promoter is inactive because the activator protein cannot bind. After an inducer binds to the activator protein, the activator protein can bind to the promoter, turning it ON and initiating transcription.
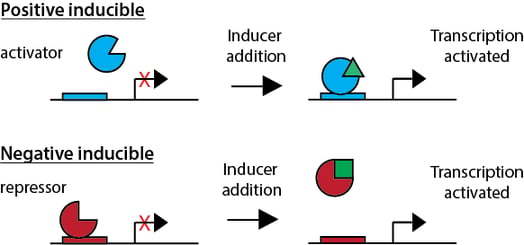
Negative inducible
In the OFF state, the promoter is inactive because a bound repressor protein actively prevents transcription. Once an inducer binds the repressor protein, the repressor protein is removed from the DNA. With the repressor protein absent, transcription is turned ON.
Types of inducible promoters
Chemical agents, temperature, and light are all examples of factors that can lead to the induction of a promoter. Below, you’ll find a short description of these three types of inducible promoters, and examples of each type. Many of these promoter systems are available at Addgene!
Chemically inducible promoters
Chemically regulated promoters are among the most common inducible promoters. The positive inducible tetracycline ON (Tet-On) system, a versatile tool developed for use in prokaryotes and eukaryotes, works via direct activation. In this system, the activator rtTA (reverse tetracycline-controlled transactivator) is normally inactive and cannot bind the tetracycline response elements (TRE) in a promoter. Tetracycline and its derivatives serve as inducing agents to allow promoter activation.
One of the most commonly used prokaryotic promoters is the negative inducible pLac promoter. This promoter requires removal of the lac repressor (lacI protein) for transcription to be activated. In the presence of lactose or lactose analog IPTG, the lac repressor undergoes a conformational change that removes it from lacO sites within the promoter and ceases repression of the target gene. A simplified lac inducible system is found in many bacterial expression vectors.
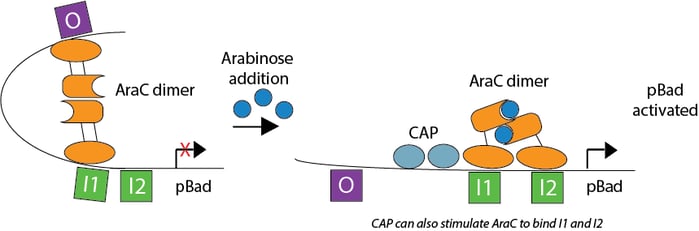
Negative inducible promoter pBad is another popular prokaryotic promoter often used for bacterial protein purification. When arabinose is absent, regulatory protein AraC binds O and I1 sites upstream of pBad, blocking transcription. The addition of arabinose causes AraC to bind I1 and I2 sites, allowing transcription to begin. In addition to arabinose, cAMP complexed with cAMP activator protein (CAP) can also stimulate AraC binding to I1 and I2 sites. Supplementing cell growth media with glucose decreases cAMP and represses pBad, decreasing promoter leakiness.
Other examples of chemically induced promoters include positive inducible alcohol and steroid regulated promoters commonly used in plant research.
| Promoter Subtype | Promoter Example | Activator | Addgene Plasmid/Kit |
| Alcohol-regulated | AlcA promoter | AlcR |
pGGA008 (AlcA) and pGGC011 (AlcR) from red flame GreenGate kit |
| Steroid-regulated | LexA promoter | XVE (synthetic) |
pFZ19 (yellow flame) |
Temperature inducible promoters
Temperature sensitive expression systems are typically less leaky than chemically induced promoters; they show near-zero expression at regular temperatures but can be induced by heat or cold exposure. Examples include the heat shock-inducible Hsp70 or Hsp90-derived promoters, in which a gene of choice is only expressed following exposure to a brief heat shock. In the case of Hsp70, the heat shock releases heat shock factor 1 (HSF-1), which subsequently binds to heat shock elements in the promoter, thereby activating transcription. Addgene depositors have developed heat shock-inducible Cre and Cas9 for easy genome engineering in species like C. elegans and Drosophila.
Light inducible promoters
Light is another way to activate gene expression, and two-component systems used in synthetic biology use light to regulate transcription. Red flame plasmid pDawn contains the blue-light sensing protein YFI. When light is absent, YFI phosphorylates FixJ, which binds to the FixK2 promoter to induce transcription of the phage repressor cI. Repressor cI inhibits transcription from phage promoter pR, preventing expression of a reporter gene. When light is present, YFI is inactive, preventing repressor cI synthesis and allowing reporter gene transcription to take place. Addgene also has popular light-regulated two component systems designed by the Tabor lab.
Which inducible promoter is right for me?
Here’s a list of some important variables to consider when choosing an inducible promoter.
Experimental System
Because transcription machinery differs between cell types or organisms, promoters are similarly variable. Bacterial promoters only work in prokaryotic cells and typically only in the same or closely related species from which they were derived. Similarly, the various eukaryotic cell types (mammalian, yeast, plants, etc.) require unique promoters and there is very little crossover. The induction mechanism must also be compatible with your experimental system.
Leakiness
If you’re expressing a gene that may be toxic, it’s important that your inducible promoter not be too leaky. For inducible prokaryotic promoters, pLac is known to be slightly leaky, and pBad is likely a better option since expression can be repressed by glucose. Temperature inducible promoters are also known for their low leakiness.
Inducibility level
How much do you want to induce transcription of your target gene? The strength of inducible promoters can vary a lot. If you’re looking for high inducibility, the Tet-On system may be a good choice, as it’s documented to induce transcription >1,000-fold when activated.
Lag time
The time needed to induce a given promoter varies. Promoters that require only the addition of an external inducer, like Tet systems, may be activated very rapidly. In contrast, promoter systems that require transcription of a repressor/inducer will have higher lag time due to the time needed for transcription/translation.
Many thanks to Nicole Zurcher who contributed to the writing of this article.
Additional Resources on the Addgene Blog:
- Plasmids 101: The Promoter Region
- Plasmids 101: Cre-lox
- Light-Switchable Two-Component Systems for E. coli
- Browse other blog posts from our Plasmids 101 series
Resources on Addgene.org:
- Inducible Promoters for Bacteria
- Tetracycline (Tet) Inducible Systems at Addgene
- Cre-lox Guide and Plasmids
Topics: Plasmid Elements, Plasmids 101, Plasmids






Leave a Comment