If you’re into cloning, you’re probably aware that there are several methodologies currently available for approaching it. These include the traditional restriction enzyme/ligase-mediated method, the more recently developed Gibson Assembly Cloning and Gateway® cloning technologies, as well as several others. Each method is unique and relies on specific components that are key to the cloning reaction. Understanding the specific components is essential for choosing the correct cloning method for your own experiments, and here we will focus on a unique gene that makes the popular GatewayTM method possible: ccdB. But what is ccdB, what role does it play in modern cloning, and why should you learn more about it? Read on to find out how ccdB can make your cloning experiments a little easier.
One of the most time-consuming aspects of traditional cloning is the identification of clones that actually contain your insert of interest. Simply stated, ccdB makes cloning easier by selecting against vectors that did not take up your insert. But exactly how does ccdB accomplish this? Let’s start with a brief history of the gene and how molecular biologists have harnessed it to evolve cloning technology.
CcdB: A potent toxin...
The ccdB gene, located on the F sex factor plasmid of E. coli, is part of a toxin-antitoxin system encoded by the ccd operon, which is responsible for plasmid maintenance during cell division. ccdB codes for the toxic protein (CcdB) that acts as a DNA gyrase poison, locking up DNA gyrase with broken double stranded DNA and ultimately causing cell death. ccdA, another gene found in the ccd operon, codes for the antitoxin protein (CcdA) that protects the cell against the toxic CcdB. Cells that lose ccdA through the loss of the F plasmid, succumb to the toxicity of CcdB.
...Becomes a powerful cloning tool
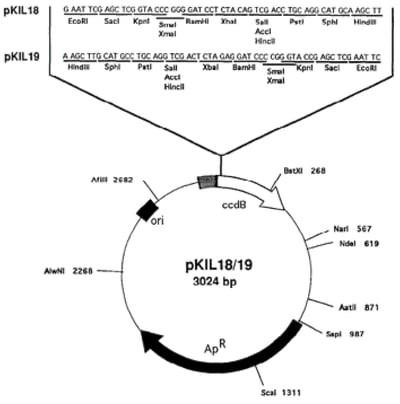
Molecular biologists first saw the potential of this system for enhancing cloning efficiency about 20 years ago and developed cloning vectors to harness it. These vectors, called pKIL18 and pKIL19, contained the ccdB gene in frame with an MCS. E. coli that were transformed with the empty vectors expressed the ccdB gene and were therefore unable to propagate because CcdA wasn’t available to counteract the toxin. If, however, an investigator were to successfully clone an insert into the vector, the ccdB reading frame would be disrupted allowing cells expressing the recombinant plasmid to propagate. Any cells that contained non-recombinant vectors (re-ligated empty vectors, for instance) would still express ccdB and therefore would die. This procedure dramatically reduces the number of clones that do not contain the recombinant plasmid and therefore makes the cloning process much more efficient, as one does not have to thoroughly screen colonies for the insert.
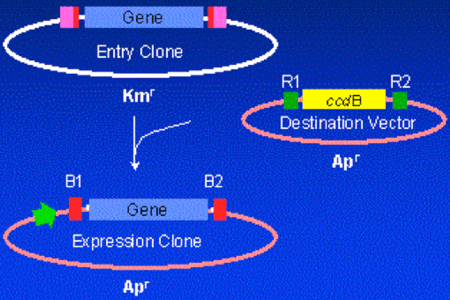
Gateway® technology (developed by InvitrogenTM) is essentially a more modern version of this older system, with added advantages that will be discussed in detail in a separate post. Focusing on the ccdB aspect, Gateway takes advantage of the same principle that cells will not propagate while expressing the gene. Briefly, the vector “backbone” in this system contains ccdB. A successful insertion will completely replace ccdB with the investigator’s insert of interest. Hence correct clones are identified much more efficiently, as those that do not contain the desired insert should not grow.
 |
|
Figure 2: Image from http://teachline.ls.huji.ac.il/72682/tutorials/gateway/introduction.html demonstrating the principal behind the “swapping out” of ccdB with the investigator’s gene of interest. |
CcdB resistant E. coli strains complete the system
But ccdB alone isn’t the only key to the system – how can one work with a plasmid/gene that kills the cells expressing it? The answer lies in special strains of E. coli that tolerate the expression of the toxin gene. One such strain is DB3.1, which contains a mutant version of DNA gyrase (gyrA462) that is resistant to the toxic effects of CcdB. Another commercially available, CcdB-resistant strain is ccdB Survival™ from InvitrogenTM. Using either DB3.1 or ccdB SurvivalTM, one can efficiently propagate and prep plasmids containing the ccdB gene, which can then be used for downstream cloning applications. While these two strains ultimately perform the same function, there is some evidence that ccdB Survival™ can be more difficult to transform, depending upon the specific plasmids being used.
We would also like to note that while DB3.1 and ccdB Survival strains have been developed specifically with ccdB-containing vectors in mind, any plasmid that contains the F plasmid (F’ strains) will also be resistant to CcdB, as the native ccdA will be present. Most common cloning strains of E. coli do not contain the F plasmid (and are considered F-); however, there are a few popular lab strains, such as NEB Stable, JM109, XL1 Blue or XL10 Gold, that are F’. These strains could possibly be used to propagate and prep your ccdB-containing empty backbones but should never be used when selecting for recombinant plasmids. Check out our blog post on Common Lab E. coli strains for more strain info.
Additional cloning resources
As cloning continues to evolve, understanding all of the players becomes ever more important to ensure that you choose the most appropriate method for your own experiments. Please check out Addgene’s Choosing a Molecular Cloning Technique page and the Addgene Blog for a variety of useful resources and protocols! We invite feedback from all members of the research community to help us develop these resources further. Feel free to offer your comments and suggestions, and even to submit a guest blog entry of your own!
References
1. Bernard, P. “Positive Selection of Recombinant DNA by CcdB.” Biotechniques. 1996 Aug;21(2):320-3. PubMed PMID: 8862819.
2. Bahassi, EM., et al. “F plasmid CcdB killer protein: ccdB gene mutants coding for non-cytotoxic proteins which retain their regulatory functions.” Mol Microbiol. 1995 Mar;15(6):1031-7. PubMed PMID: 7623659.
3. Bernard, P., et al. “Positive-selection vectors using the F plasmid ccdB killer gene”. Gene. 1994 Oct 11;148(1):71-4. PubMed PMID: 7926841.
4. Bernard, P., et al. “The F plasmid CcdB protein induces efficient ATP-dependent DNA cleavage by gyrase.” J Mol Biol. 1993 Dec 5;234(3):534-41. PubMed PMID: 8254658.
5. Bernard, P. and Couturier, M. “Cell killing by the F plasmid CcdB protein involves poisoning of DNA-topoisomerase II complexes.” J Mol Biol. 1992 Aug 5;226(3):735-45. PubMed PMID: 1324324.
6. Tam, JE. and Kline, BC. “Control of the ccd operon in plasmid F.” J Bacteriol. 1989 May;171(5):2353-60. PubMed PMID: 2651399. PubMed Central PMCID: PMC209908
Resources on the Addgene Blog
- Plasmids 101: Common Lab E. coli Strains
- Plasmids 101: TOPO Cloning
- Plasmids 101: Golden Gate Cloning
- Read all of our Plasmid Cloning Posts
Resources on Addgene.org
- Choose a Molecular Cloning Technique
- Find Plasmid Cloning Protocols
- Check Out All of Our Protocols
Additional Web Resources
Topics: Plasmids 101, Plasmid Cloning, Plasmids






Leave a Comment