 Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Listen this episode of Hot Plasmids!
Light controls the activity of a microtubule-associated protein to regulate cell growth dynamics
Article contributed by Klaus Wanisch
![]() Listen to the optogenetics podcast segment
Listen to the optogenetics podcast segment
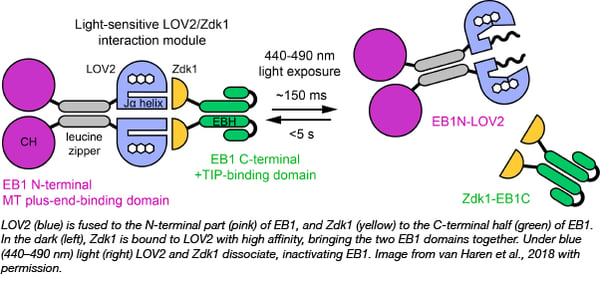
LOVTRAP is an optogenetic system that was recently developed by Klaus Hahn's lab at the University of North Carolina (plasmids at Addgene). It consists of two domains, LOV2 and Zdk1, where Zdk1 binds LOV2 with high affinity in the dark but dissociate under blue light. This tool has now been modified by Torsten Wittmann’s lab at UCSF to modulate cellular dynamics with light by modifying EB1. EB1 normally sits on microtubules to attract other proteins involved in their growth and extension. Now, they split EB1 into two halves (Fig. 1): one containing the microtubule binding domain and the second containing the growth promoting domain. Each half was fused with one of the LOVTRAP domains. When both halves of the modified EB1 protein are overexpressed in cells, microtubule growth can be switched off with blue light which causes LOV2 and Zdk1 to dissociate due to conformational changes, and as a consequence, EB1 protein function is inactivated by the separation of its synergistic domains.
This paper demonstrated that the migration of a cancer cell line equipped with photodissociable EB1 could be stopped in areas with blue light - a finding which opens novel ways in manipulating and studying cellular dynamics. The photoinducible control of protein function is more specific and much faster than any genetic or pharmacological manipulations on cells, and has potential use for a wide range of applications in cellular biology.
Van Haren J et al. Nature Cell Biology 2018 Pubmed PMID 29379139.
The “Riboglow” platform for visualizing RNA production and localization
Article contributed by Tyler Ford
![]() Listen to the riboglow podcast segment
Listen to the riboglow podcast segment
Researchers from the Palmer lab at the University of Colorado Boulder recently developed “Riboglow,” a set of tools that can be used to visualize RNA localization in mammalian cells. These tools consist of cobalamin-binding riboswitches that bind to cobalamin-fused fluorescent dyes; binding of the cobalamin fusion to the riboswitch results in increased fluorescent signal. Prior to the development of these probes, fluorophore binding aptamers and MS2-binding fluorescent proteins were the state of the art in RNA imaging in vivo. These new “Riboglow” probes contain a smaller RNA tag, are less prone to photobleaching, and achieve better signal in vivo as compared to imaging with broccoli aptamers or MS2 tags using the same number of fluorophores. You can find plasmids containing the cobalamin binding riboswitches used in the Riboglow platform at Addgene.
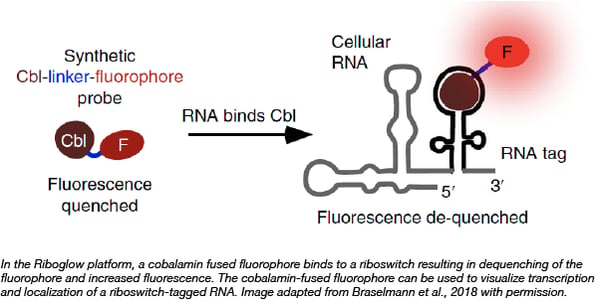
Braselmann E, et al. Nature Chemical Biology. 2018 PubMed PMID: 30061719.
New tools for developing new FPs - the mCarmine story
Article contributed by Caroline LaManna
![]() Listen to the mCarmine podcast segment
Listen to the mCarmine podcast segment
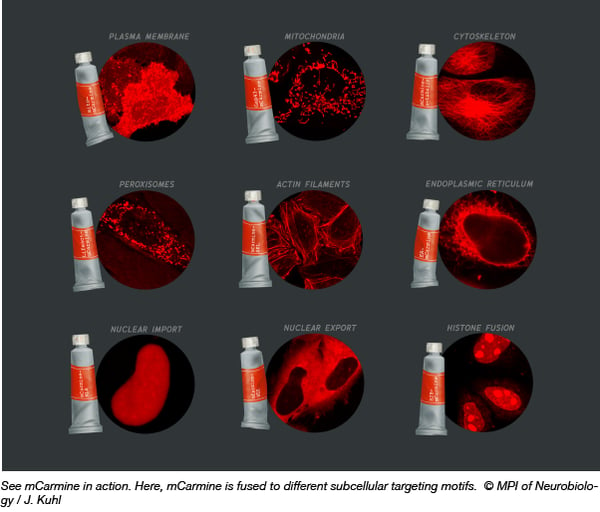
Scientists are always on the lookout for better, brighter fluorescent proteins (FP) with improved tissue penetrance and live cell imaging capabilities. Upon developing a new imaging-based bacterial colony screening platform, Oliver Griesbeck’s lab set out to engineer a new FP. Their aim - to red-shift the boundaries of FP technology by shifting the emission maxima toward the red end of the spectrum (above 600 nm) and increasing the brightness. Arne Fabritius and colleagues started with the FP mNeptune684 (originally described by Li ZY, et al. PLoS One, 2016) and tested a series of mutations. Through this process mCarmine was developed (mNeptune684 with C65S, T107K, A108V, T109K, H161Y, P163T, I175Q, C176T and N177F mutations). Its excitation maximum is reported as 603 nm, while the emission maximum is 675 nm. mCarmine is monomeric, 4-5 fold brighter than mNeptune684, and has a favorable pKa of 5.6. To test out this new FP in your own experiments, mCarmine is available in a mammalian expression vector (pcDNA3) and a bacterial expression vector (pRSETB).
Fabritius A, et al. Cell Chemical Biology 2018. PubMed: PMID 30220597.
ACE system: A versatile in vivo fluorescent reporter for rapid quantification of base-editing and Cas9 cleavage activity
Article contributed by Angela Kaczmarczyk![]() Listen to the base editing podcast segment
Listen to the base editing podcast segment
The increasing popularity of single base-editors has resulted in the rapid expansion of the genome editing toolkit. However, a challenge in optimizing base editing technologies across several cell types is the lack of a rapid and quantitative editing assay.
To help accelerate the expansion of genome editing tools, the Reuben Harris lab has deposited a reporter plasmid that allows for rapid fluorescence-based quantification of base editors that utilize dCas9 fused to cytidine deaminases like APOBEC. To achieve this they created a lentiviral vector system that monitors both APOBEC- and Cas9-mediated Editing (ACE) in real-time. The reporter constitutively expresses eGFP to indicate reporter abundance, and mCherry fluorescence is activated when APOBEC-Cas9n-UGI-mediated activity edits out a frame-shift mutation within the mCherry reporter. Quantification of editing efficiencies simply involves tracking the ratio of mCherry-positive to eGFP positive cells through fluorescence microscopy or flow cytometry. Using the ACE system, the Harris lab showed that human APOBEC3A-Cas9n-UGI and APOBEC3B-Cas9n-UGI base editing complexes are more efficient than the BE3 plasmid containing rat APOBEC1. Finally, they show that the ACE reporter can also quantify Cas9 cleavage, thus exhibiting its versatile utility in quantifying multiple gene editing activities.
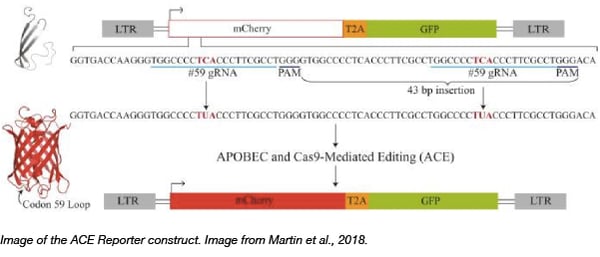
Click here to find the ACE reporter plasmid!
Martin et al., Nucleic Acids Research 2018 PubMed: PMID 29746667.
Topics: Other Plasmid Tools, Plasmids









Leave a Comment