 Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
A retroviral mouse CRISPR knockout library
Article contributed by Alycia Bittner
CRISPR pooled libraries are popular tools for genetic screens, enabling genome-wide interrogation of the genes required for cellular processes. For a successful screen, it is necessary to identify the gRNAs that elicit the phenotype, and this depends on the method used to introduce the gRNAs into cells. Most CRISPR pooled libraries have been created in lentiviral systems for mammalian screens, which enables researchers to maximize the chance of transducing a only a single gRNA per cell. The viral DNA containing the gRNA then integrates into the genome. Thus the gRNAs responsible for a phenotype can be identified by next-generation sequencing of the genomic DNA. However, some cell types are poorly transduced by lentivirus, and thus screening with available CRISPR libraries was not possible.
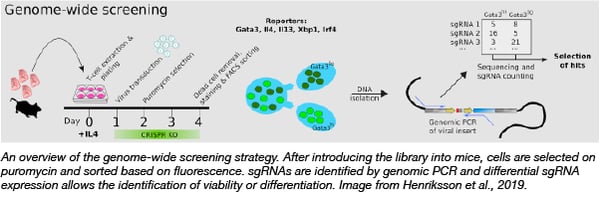
Primary murine T cells cannot be transduced by lentivirus and have been difficult to manipulate genetically. To solve this issue, Sarah Teichmann and colleagues created a retroviral CRISPR knockout library using gRNAs designed by the Kosuke Yusa Lab. This retroviral library was used to transduce T helper type 2 (Th2) cells from mice that constitutively express Cas9 and screen for genes that affect Th2 differentiation and activation, using expression of differentiation factors as the screen readout. This retroviral mouse CRISPR pooled library can be used for screening of cells that are refractory to lentiviral transduction, now enabling forward genetic screens in cells with few genetic tools.
Find the mouse CRISPR knockout library at Addgene.
Henriksson et al., Cell, 2019. PubMed PMID: 30639098.
LipoGlo: A reporter for measuring ApoB bound lipoproteins
Article contributed by Leah Schwiesow
A leading cause of death worldwide, cardiovascular disease is driven by atherogenic lipoproteins that accumulate within the vascular wall. ApoB, the main structural component of atherogenic lipoproteins, shuttles lipoproteins through the circulatory systems, but can also lead to the formation of atherosclerotic plaques. ApoB bound lipoproteins (ApoB-LPs) are indirectly measured through their triglyceride or cholesterol contents (what is commonly called bad cholesterol). However, size distribution and concentration of ApoB-LPs has been shown to have a strong correlation with cardiovascular disease and indirect measurement of ApoB-LPs through triglyceride/cholesterol does not provide a clear view of either of these metrics (Sniderman et al, 2012; Lawler et al, 2017).
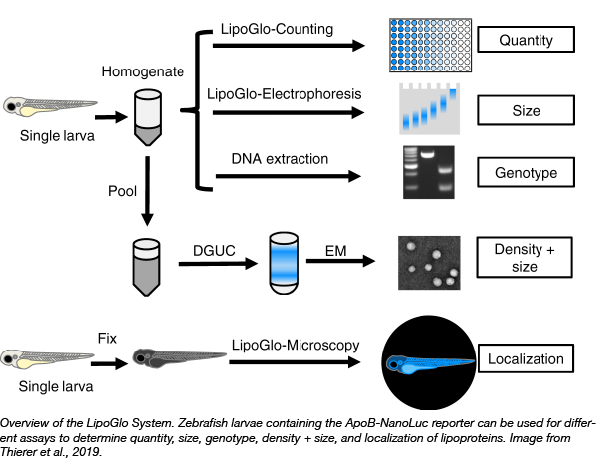
In an effort to measure ApoB-LPs directly and develop a potentially high throughput in vivo model to identify drug candidates that modulate ApoB-LP size and concentration, Steven Farber and colleagues at the Carnegie Institute for Science have developed the LipoGlo system in zebrafish (Thierer et al, 2019). Zebrafish are a great model for high throughput study of ApoB-LPs because their ApoB homologes are functionally conserved, they develop rapidly, they are small, and their transparency allows for whole organism imaging. They created an endogenous ApoB-NanoLuc reporter by inserting the NanoLuc luciferase downstream of the apoB gene. Using this reporter, the researchers explore many aspects of ApoB-LP biology, including quantification of abundance, size, and localization in vivo. The LipoGlo system represents a new, intriguing approach to expand our understanding of ApoB-LPs and accelerate the discovery of additional genes and small-molecule probes to aid in the understanding and treatment of cardiovascular disease.
Find the LipoGlo plasmids at Addgene!
Lawler et al, QJM: An International Journal of Medicine 2017. Pubmed PMID: 16371404
Sniderman et al, Athersclerosis 2012. Pubmed PMID: 23068583
Thierer et al, Nature Communications 2019. Pubmed PMID: 31366908
Adapting sigma54-dependent transcription for CRISPR activation in bacteria
Article contributed by Will Arnold
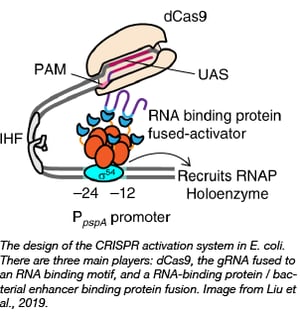 Baojun Wang’s lab recently developed a CRISPR tool for the specific activation of transcription in bacteria by co-opting σ54-based transcription. Transcription from σ54 promoters requires binding of bacterial enhancer binding proteins (bEBPs) at a sequence upstream of the promoter called the upstream activating sequence (UAS). The Wang Lab created a tripartite system consisting of dCas9, a gRNA fused to an RNA binding motif, and a RNA-binding protein-bEFP fusion. Thus, the bEFPs bind the UAS via the dCas9-gRNA complex which allows transcription from the promoter.
Baojun Wang’s lab recently developed a CRISPR tool for the specific activation of transcription in bacteria by co-opting σ54-based transcription. Transcription from σ54 promoters requires binding of bacterial enhancer binding proteins (bEBPs) at a sequence upstream of the promoter called the upstream activating sequence (UAS). The Wang Lab created a tripartite system consisting of dCas9, a gRNA fused to an RNA binding motif, and a RNA-binding protein-bEFP fusion. Thus, the bEFPs bind the UAS via the dCas9-gRNA complex which allows transcription from the promoter.
They found that the system is active in E. coli and closely related species, is amenable to use with other bEBP/UAS pairs, and displays high dynamic range. However, the promoter in question must be σ54-dependent and the system may not be adaptable to distantly related species from E. coli. This tool should serve as both a powerful tool for generating complex regulatory systems and further the study of natural σ54-dependent promoter systems.
Find these CRISPR activation plasmids at Addgene.
Liu et al., Nature Communications, 2019. PubMed PMID: 31451697.
Genetically encoded voltage indicators for use in worms
Article contributed by Brook Pyhtila
 Genetically encoded voltage indicators (GEVIs) change fluorescence in real time when neurons or muscle cells transmit electrical information. Such voltage imaging can be performed in intact animals and offers many advantages over traditional measurements such as patch clamp recordings. However, technical difficulties such as small fluorescence changes or a requirement for very high excitation light intensities have made these indicators difficult to work with.
Genetically encoded voltage indicators (GEVIs) change fluorescence in real time when neurons or muscle cells transmit electrical information. Such voltage imaging can be performed in intact animals and offers many advantages over traditional measurements such as patch clamp recordings. However, technical difficulties such as small fluorescence changes or a requirement for very high excitation light intensities have made these indicators difficult to work with.
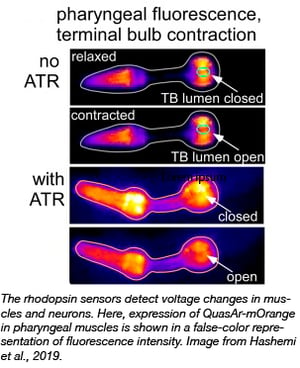
The Gottschalk lab has surveyed a range of microbial rhodopsin GEVIs in excitable cells of the nematode Caenorhabditis elegans. They have found several voltage biosensors and conditions that can be used to monitor real time voltage changes in worms and have made these constructs available at Addgene. These include Arch(D95N) and QuasAr, as well as the eFRET sensors, QuasAr-mOrange and MacQ-mCitrine. All of these sensors can detect action potentials in worms with fluorescence changes of up to 128% per 100 mV in the pharynx, and 78% in body wall muscle cells. These changes were larger than previously reported for other organisms, and the Gottschalk group hypothesizes that this may be due to different lipid compositions of C. elegans membranes, which might beneficially influence the rhodopsin properties.
Find these voltage biosensors at Addgene.
Hashemi et al., PNAS. 2019, PubMed PMID 31371514.
Topics: Other Plasmid Tools, Plasmids






Leave a Comment