Autophagy (Greek for “self-eating”) is a process by which cytoplasmic material, including organelles, are targeted to lysosomes for degradation. Autophagy is a dynamic process which involves autophagosome synthesis, delivery of materials to be degraded to the lysosome, and degradation of autophagic substrates inside the lysosome. Historically, methods for studying autophagy focused on counting the number of autophagosomes. This approach, however, has inherent limitations because it turns a dynamic process into a static measurement and it provides limited information about what materials or organelles are being targeted for autophagy. The development of several fluorescent autophagy reporters now allows for the measurement of autophagic flux, or the changes in autophagic activity, and are a more reliable indicator of autophagic activity. The aim of this post is to provide an overview of four autophagy biosensors currently available from Addgene.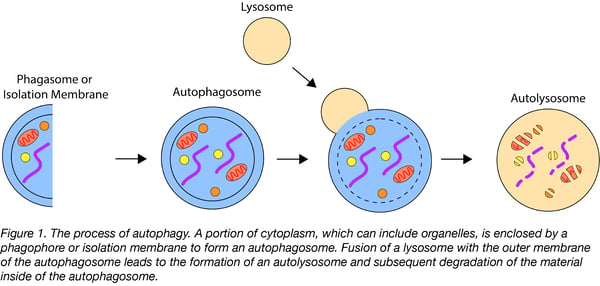
Rosella autophagy biosensor
Rosella is a dual-emission autophagy biosensor that’s comprised of two tandem fluorescent proteins: a relatively pH-insensitive RFP variant, DsRed.T3, that’s connected to a pH-sensitive GFP variant, super ecliptic pHluorin (SEP). In the absence of autophagy, the untargeted Rosella remains in the neutral pH cytoplasm where it emits red and green fluorescence. Following the induction of bulk autophagy, Rosella is targeted to the vacuole in yeast or the lysosome in mammalian cells. This decrease in pH quenches SEP but not DsRed, leading to emission of only red fluorescence. The ratio of red to green fluorescence allows for clear distinction between vacuolar/lysosomal and cytoplasmic Rosella. When fused with a mitochondria-targeting sequence, Rosella can be used to track the targeted autophagy of mitochondria, or mitophagy. Rosella can also be fused to a protein of interest to track the autophagy of that protein.
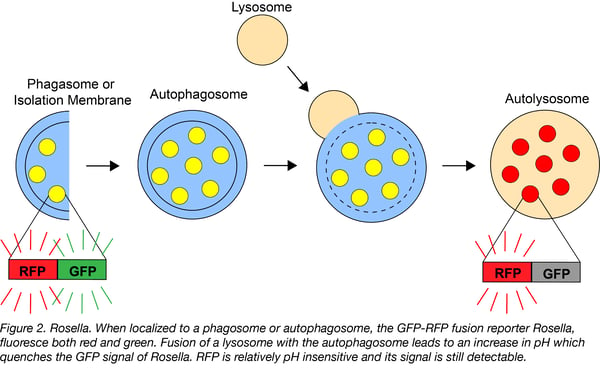
Find the cytosol-targeted and mitochondria-targeted Rosella at Addgene!
Keima autophagy biosensor
Keima is a mammalian dual-emission autophagy biosensor. Its resistance to lysosomal proteases is key to its pH-dependent fluorescence. In neutral environments, dimeric Keima is excited at 440 nm and in acidic environments at 586 nm. Regardless of excitation wavelength, Keima emits at 620 nm. The ratio of signal strength excited at 586 nm to 440 nm serves as a measure of autophagy induction: when Keima is in the cytosol, this ratio is low; but when Keima accumulates in autolysosomes, this ratio is high. When untargeted, Keima can be used to measure bulk autophagy, but the addition of a mitochondria-targeting sequence makes Keima useful for monitoring mitophagy. It’s important to remember that because Keima’s fluorescent relies on lysosomal acidity, this biosensor can’t be used with fixed cells.
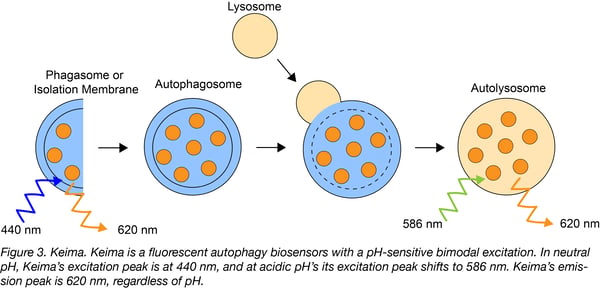
Find the mitochondria-targeted Keima variant at Addgene!
mRFP-GFP-LC3 tandem fluorescent protein
This autophagy biosensor contains mRFP and GFP fused to the autophagy-related microtubule-associated proteins 1A/1B light chain 3B, or LC3. LC3 is a commonly used marker for measuring autophagic flux. mRFP-GFP-LC3 functions as an autophagy bionsensor via the same mechanism as Rosella: GFP is quenched in acidic environments, while mRFP is relatively stable and fluorescences even at the acidic pH found in lysosomes. Therefore, the formation of autophagosomes leads to an increase of yellow puncta (GFF positive/RFP positive), with these puncta turning red (GFP negative/RFP positive) upon fusion of an autophagosome with a lysosome. An increase in autophagy leads to more yellow and red puncta, while a block at fusion of autophagosomes with lysosomes leads to an increase in yellow puncta but a decrease in red puncta.
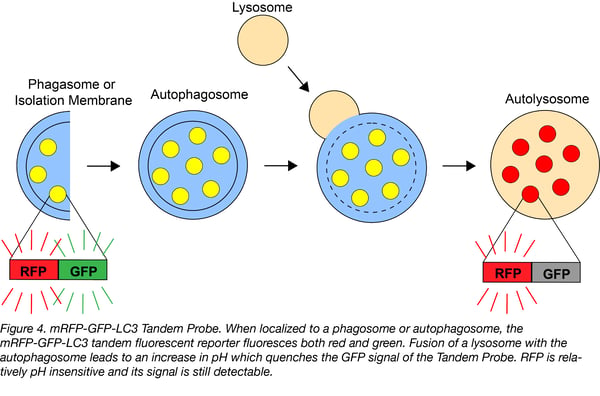
Find the mRFP-GFP-LC3 plasmid at Addgene!
GFP-LC3-RFP-LC3∆G: A fluorescent probe that releases an internal control
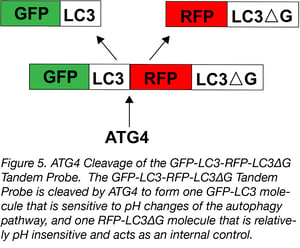
Another autophagy biosensor is the GFP-LC3-RFP-LC3ΔG probe. It’s similar to the mRFP-GFP-LC3 Tandem Fluorescent Protein, but following translation, this fusion protein is cleaved to generate one molecule of GFP-tagged LC3 for every one molecule of RFP-tagged LC3∆G.
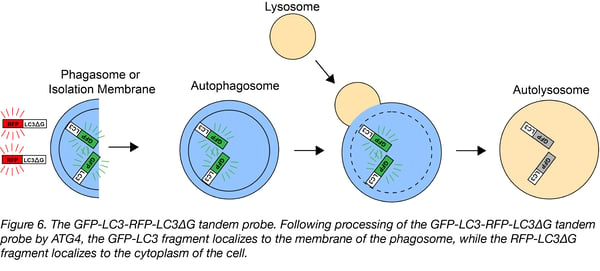
Like the unmodified LC3, GFP-LC3 is conjugated to phosphatidylethanolamine (PE) and recruited to autophagasomal membranes, which leads to its degradation by autophagy. The RFP-LC3∆G, however, can not be conjugated to PE due to a C-terminal glycine deletion, so it remains in the cytosol acting as an internal control. Autophagic flux is estimated by the GFP/RFP ratio: a decrease in GFP fluorescence relative to RFP fluorescence suggests an increase in autophagic flux, while an increase in GFP fluorescence relative to RFP fluorescence suggests a decrease in autophagic flux.
Find the GFP-LC3-RFP-LC3∆G plasmid at Addgene!
References
Klionsky, Daniel J., Ana Maria Cuervo, and Per O. Seglen. "Methods for monitoring autophagy from yeast to human." Autophagy 3.3 (2007): 181-206. PubMed PMID: 17224625.
Mizushima, Noboru, Tamotsu Yoshimori, and Beth Levine. "Methods in mammalian autophagy research." Cell 140.3 (2010): 313-326. PubMed PMID: 20144757. PubMedCentral PMCID: PMC2852113.
Noda, Takeshi, and Daniel J. Klionsky. "The quantitative Pho8Δ60 assay of nonspecific autophagy." Methods in enzymology 451 (2008): 33-42. PubMed PMID: 19185711.
Rosado, Carlos, et al. "Rosella: a fluorescent pH-biosensor for reporting vacuolar turnover of cytosol and organelles in yeast." Autophagy 4.2 (2008): 205-213. PubMed PMID: 18094608.
Sargsyan, A., et al. "Rapid parallel measurements of macroautophagy and mitophagy in mammalian cells using a single fluorescent biosensor." Scientific reports 5 (2015): 12397. PubMed PMID: 26215030. PubMed Central PMCID: PMC4517063.
Yoshii, Saori R., and Noboru Mizushima. "Monitoring and measuring autophagy." International journal of molecular sciences 18.9 (2017): 1865. PubMed PMID: 28846632. PubMed Central PMCID: PMC5618514.
Additional resources on the Addgene blog
- Beware of fluorescent protein oligomerization
- Seeing red: simple GFP photoconversion
- Light sheet fluorescence microscopy
Resources on Addgene.org





Leave a Comment