Blugene and I represented Addgene at the recent Keystone meeting on Precision Genome Editing with Programmable Nucleases. Check out #KSgenome on Twitter if you missed our live updates!
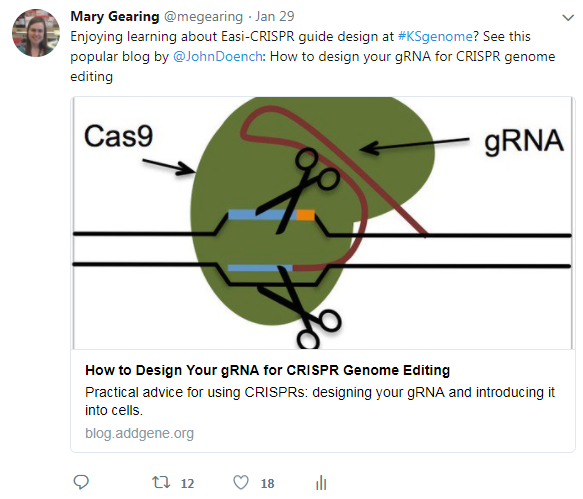
My most popular tweet from the conference is shown above. The blog post I mentioned, John Doench’s How to Design your gRNA for CRISPR Genome Editing, gets over 4,000 views a month. To me, these numbers show that this basic concept of gRNA design is still challenging for researchers using CRISPR, and that standardizing and improving the gRNA design process and rules for various applications is necessary going forward.
The standardization of CRISPR methods was a common thread at the meeting. The new Somatic Cell Genome Editing program, run through the NIH Common Fund, emphasizes that we need more standardized methods for evaluating CRISPR technologies, including the ever important on/off-target efficiency. Likewise, the NIST Genome Editing Consortium hopes to evaluate genome editing pipelines and generate a common lexicon for genome editing studies. These complementary goals are aimed at improving translation of CRISPR technology, but they would also bring benefits to researchers conducting basic or early pre-clinical research. However, standard terms and techniques are not enough to move the CRISPR field forward. To do that, we need to learn much more about cellular DNA repair pathways.
An emphasis on cellular repair post-CRISPR cleavage
When I first looked at the conference agenda, I was surprised to see that it would open not with an introduction to CRISPR nucleases, but with a session on DNA repair. What stuck most in my mind from that session were words from Eric Hendrickson, loosely paraphrased in the tweet below.
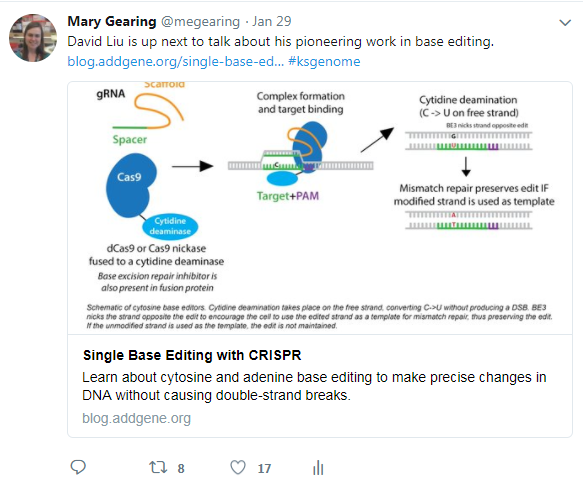
The complexity and challenges of homology directed repair (HDR) are echoed in the amount of interest in base editors, which alter genomic DNA without the need for cellular HDR.
Having had time to digest my notes, since the conference, I think I may have finally grasped just how complicated DNA repair pathways are. I fully admit to being a DNA repair newbie - diagrams of resection and strand invasion are like a foreign language to me. But coming to #KSgenome and seeing the intricacies of DNA repair has emphasized how essential these studies are to improving genome editing efficiency. There is so much more to working with CRISPR than just designing a repair template that contains your desired edit, and I want to make sure that I (and Addgene) don’t forget this in the future.
Here’s a list of blog posts to help you tackle the challenges of DNA repair (with bonus gRNA design resources.) Subscribe to CRISPR updates on our blog to keep current with the latest and greatest!
Addgene resources
3 Tips to Improve HDR Efficiency for CRISPR Editing in Human Cells
Hands-on tips for repair template design from guest bloggers Dominik Paquet and Dylan Kwart, including a detailed look at how cut-to-mutation distance affects editing efficiency.
Optimizing Donor DNA for Enhanced CRISPR Genome Editing
Chris Richardson and members of Jacob Corn’s lab describe how a better understanding of Cas9 biophysics can help enhance HDR.
CRISPR 101: Ribonucleoprotein (RNP) Delivery
The RNP delivery method consistently leads to high on-target editing and low off-target editing rates across multiple systems. Read our introductory blog post, then check out this RNP protocol on Protocols.io.
The PAM Requirement and Expanding CRISPR Beyond SpCas9
The closer your edit is to the PAM site, the higher the efficiency of HDR repair. Check out this post to see if an engineered Cas9 variant might work for your experiment.
PITChing MMEJ as an Alternative Route for Gene Editing
Michohomology-mediated end joining (MMEJ), also known as alternative non-homologous end joining (Alt-NHEJ), has been harnessed for CRISPR knock-in applications.
CRISPR 101: Targeting RNA with Cas13a (C2c2)
Maybe you’d like to avoid editing the genome and just stick to RNA? Cas13 may be able to help you do that!
How to Design your gRNA for CRISPR Genome Editing
Other resources and publications
Researchers at the Keystone meeting had great things to say about CRISPOR, a tool to help you design, pick, and close guides for Cas9 and Cpf1. This tool supports over 250 genomes, and they just released a brand new CRISPOR manual to kick off 2018!
"Mechanisms of precise genome editing using oligonucleotide donors". Kan, Yinan, Brian Ruis, Taylor Takasugi, and Eric A. Hendrickson. Genome Res 27(7) (2017):1099-1111. PubMed PMID: 28356322 PubMed Central PMCID: PMC5495063
Kan et al. use a GFP-BFP conversion system to assess precision genome editing patterns with ssDNA repair templates. Their work demonstrates that different repair template strategies activate different HDR sub-pathways like synthesis-dependent strand annealing (SDSA) and single-strand DNA incorporation (ssDI).
"Easi-CRISPR for creating knock-in and conditional knockout mouse models using long ssDNA donors". Miura, Hiromi, Rolen M. Quadros, Channabasavaiah B. Gurumurthy, and Masato Ohtsuka. Nat Protoc. 13(1) (2018):195-215. PubMed PMID: 29266098
Creating knock-in mouse models with CRISPR has been challenging, with protocols using dsDNA templates achieving ~1-10% efficiency. Easi-CRISPR uses a long ssDNA template to increase efficiency of insertion and gene replacement.
Care to share your favorite CRISPR tool or resource? Let us know in the comments section!
Topics: CRISPR





Leave a Comment