This post was contributed by guest blogger Joachim Goedart, an assistant professor at the Section of Molecular Cytology and van Leeuwenhoek Centre for Advanced Microscopy (University of Amsterdam).
GFP is the most popular, most widely used genetically encoded fluorescent probe. Several factors contribute to the popularity of GFP including (i) fast and complete maturation to functional, fluorescent protein in almost all organisms and cell types, (ii) no need to add a co-factor, (iii) easy visualization with standard filter sets on a fluorescence microscope, and finally (iv) good toleration in fusion proteins.
Since GFP is such a well-validated, all-round good performing probe, it is the first choice when selecting a genetically encoded fluorescent tag. There are, however, a number of limitations that you may run into if you choose to use it. Several of these limitations and possible solutions are discussed below.
There are numerous GFP variants. Here I use the name ‘GFP’, which refers to mEGFP. This variant was made by introducing two mutations (F64L, S65T) in the original jellyfish GFP (AvGFP) to create enhanced GFP (EGFP), a variant with improved brightness (Tsien, 1998). Another mutation (A206K) is necessary to generate the strictly monomeric EGFP variant, mEGFP (Zacharias et al., 2002).
GFP needs oxygen
Chromophore formation requires a fully folded beta-barrel structure, followed by an intramolecular reaction that generates the chromophore (Tsien, 1998). Importantly, this reaction requires molecular oxygen and as a consequence GFP remains non-fluorescent under anaerobic conditions. This holds true for any of the AvGFP and coral derived fluorescent proteins.Therefore, GFP is not suitable for applications that require anaerobic conditions.
In experiments requiring anaerobic conditions, proteins that bind endogenous fluorescent co-factors can be used. Examples include the LOV domains that bind flavin (Buckley et al., 2015) or the UnaG protein that binds bilirubin (Kumagai, 2013). Next to these naturally occurring co-factor binding proteins, systems wherein a protein binds a synthetic, fluorescent analogue have been engineered (Thorn, 2017). These fluorescent analogues are generally added to the bathing solution and require uptake by the cells. Which of these strategies works best for a specific biological system under anaerobic conditions needs to be examined on a case-by-case basis.
GFP is sensitive to acid
The chromophore of GFP can exist in a protonated and a deprotonated state (Tsien, 1998). The deprotonated state has an absorbance maximum around 488 nm and it emits light that peaks at 508 nm. The protonated state, however, does not absorb light at 488 nm. The ratio between the two states is reflected by the pKa, which is around 6.0 for GFP. This number implies that at pH 6 only 50% of the available green fluorescent proteins emit light.
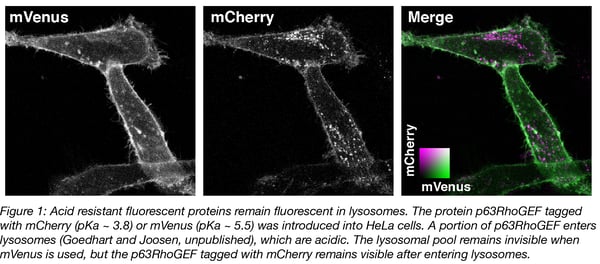
The pKa of fluorescent proteins can be accurately determined in vitro by measuring the fluorescence intensity of purified protein at different acid concentrations. Overviews of the pKa values of different fluorescent protein are available (Cranfill et al., 2016 and FPbase.org). A number of variants remain fluorescent under acidic conditions, with pKa values below 4. Examples include mTurquoise2 (pKa = 3.1), tagRFP (pKa = 3.1) and mCherry (pKa = 3.8). We have observed that mCherry (and mTurquoise2, not shown) remains fluorescent in acidic organelles (Figure 1), showing that its acid tolerance is maintained in cells.
GFP is big
GFP is a 28 kDa protein that resembles a cylinder with a length of 4.2 nm and a diameter of about 2.4 nm (Hink et al., 2000). The complete beta-barrel is necessary for its fluorescence and therefore GFP cannot be downsized by deleting residues. Only about 10 amino acids can be deleted from the C-terminus and about 5 from the N-terminus, but this hardly reduces its size. The substantial size of GFP may interfere with the activity or the localization of the fusion protein. Also, in cases where the protein of interest is relatively small, adding GFP may alter diffusion rates.
Smaller genetically encoded probes are based on cofactor binding proteins as described above. The smallest tags are peptides that are engineered to bind specifically and with high affinity to fluorophores. Examples are the FlAsH tag, the His tag and several others. Another peptide-based strategy uses an enzyme to covalently label the peptide with a fluorophore. Both systems are a hybrid between a genetically encoded tag and exogenously added components. For a review see Lotze et al. (2016). To date, small probes that are entirely genetically encoded do not exist, but the development of non-natural fluorescent amino acids and their incorporation into proteins are steps in that direction (Chatterjee et al., 2013; Hilaire et al., 2017).
GFP can only be used for tagging proteins
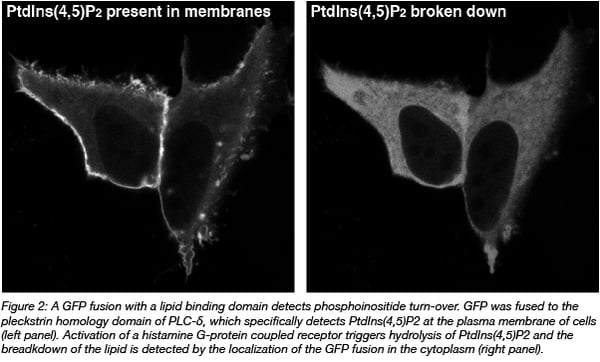
GFP fusion proteins are made by connecting its DNA code to the cDNA encoding for another protein. Effectively, it equips the protein of interest with an additional protein module that can report on location. This approach enables the study of proteins in cells, but not other biomolecules of interest e.g. DNA, RNA, and lipids. Still, these molecules can be indirectly visualized by using protein domains that specifically bind the molecule of interest. For instance, pleckstrin homology domains can be used to detect specific phosphoinositides (Varnai and Balla, 2008) (Figure 2).
RNA production can be detected by using a MS2 coat protein that specifically detects a specific RNA stem loop-structure (Querido and Chartrand, 2008). Finally, a nuclease-deficient Cas9 can bind loci in the genome that are defined by a gRNA. Since Cas9 is a protein, a fusion with GFP allows for detection of genomic locations (Chen et al., 2013).
GFP fluorescence overlaps with autofluorescence
The cyan light that is used to excite GFP, may also excite several components that are naturally present in cells or media. These sources of ‘autofluorescence’ include riboflavin in the medium or flavin containing proteins in mammalian cells. Generally, autofluorescence is less of a problem when the excitation wavelength is shifted to longer wavelengths. Therefore, the use of bright red fluorescent proteins (e.g. mScarlet) or infrared fluorescent proteins (Chernov et al., 2017) may improve detection. Since longer wavelengths have better tissue penetration, (infra)red fluorescent proteins are also a better choice for thicker samples.
Another solution to avoid autofluorescence is offered by bioluminescent proteins. These probes generate light by a chemical reaction and hence, there is no need to excite the sample. Although bioluminescent probes are usually dim relative to fluorescent probes, recent engineering efforts have increased the brightness of these genetically encoded light emitters (i.e. Takai, 2015 and Iwano et al., 2018).
GFP is not tolerated in my fusion
It is standard practice to attach the GFP to the N- or C-terminus of another protein. This may, however, interfere with the function of the protein. This is clearly the case when proteins undergo post-translational modification with fatty acyl chains. For instance, myristoylation requires an N-terminal consensus sequence (MG-) and prenylation requires a C-terminal consensus sequence (-CaaX). Both modifications are disrupted when GFP is fused to these sequences.
When N- and C-terminal fusions are not tolerated the alternative is to insert the GFP into the coding sequence of the protein of interest. One of the reasons that this works well is that the N- and C-terminus of GFP itself are relatively close, thereby minimizing the disruption of the target protein. In this way, we (and others) have successfully generated several functional heterotrimeric G-alpha subunit fusions that do not tolerate N- and C-terminal fusions (Adjobo-Hermans, 2011). Structural information facilitates the choice of the proper site, although it still is worthwhile to generate several variants and check the functionality of the fusion (Mastop, 2018).
Summary
GFP is an all-round useful genetically encoded probe, but there are limitations. Being aware of the limitations is important for the successful use of GFP or any of the other fluorescent proteins. Luckily there are often solutions for many of the limitations, and the number of alternatives to GFP is expected to grow with the ever-expanding genetic toolbox generated by the continuous effort of numerous scientists world-wide.
Many thanks to our guest blogger, Joachim Goedhart!
 Joachim Goedhart is an assistant professor at the Section of Molecular Cytology and Van Leeuwenhoek Centre for Advanced microscopy (University of Amsterdam). He develops, characterizes, and uses genetically encoded fluorescent probes. You can follow him on twitter: @joachimgoedhart.
Joachim Goedhart is an assistant professor at the Section of Molecular Cytology and Van Leeuwenhoek Centre for Advanced microscopy (University of Amsterdam). He develops, characterizes, and uses genetically encoded fluorescent probes. You can follow him on twitter: @joachimgoedhart.
References
1. Zacharias, David A., et al. "Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells." Science 296.5569 (2002): 913-916. PubMed PMID: 11988576.
2. Tsien, Roger Y. "The green fluorescent protein." (1998): 509-544. PubMed PMID: 9759496.
3. Buckley, Anthony M., et al. "LOV-based reporters for fluorescence imaging." Current opinion in chemical biology 27 (2015): 39-45. PubMed PMID: 26087123.
4. Kumagai, Akiko, et al. "A bilirubin-inducible fluorescent protein from eel muscle." Cell 153.7 (2013): 1602-1611. PubMed PMID: 23768684.
5. Thorn, Kurt. "Genetically encoded fluorescent tags." Molecular biology of the cell 28.7 (2017): 848-857. PubMed PMID: 28360214. PubMed Central PMCID: PMC5385933.
6. Cranfill, Paula J., et al. "Quantitative assessment of fluorescent proteins." Nature methods 13.7 (2016): 557. PubMed PMID: 27240257. PubMed Central PMCID: PMC4927352
7. Hink, Mark A., et al. "Structural Dynamics of Green Fluorescent Protein Fused with a Single Chain Fv Protein." Journal of Biological Chemistry (2000). PubMed PMID: 10748019.
8. Lotze, Jonathan, et al. "Peptide-tags for site-specific protein labelling in vitro and in vivo." Molecular BioSystems 12.6 (2016): 1731-1745. PubMed PMID: 26960991.
9. Chatterjee, Abhishek, et al. "A genetically encoded fluorescent probe in mammalian cells." Journal of the American Chemical Society 135.34 (2013): 12540-12543. PubMed PMID: 23924161 PubMed Central PMCID: PMC3783214.
10. Hilaire, Mary Rose, et al. "Blue fluorescent amino acid for biological spectroscopy and microscopy." Proceedings of the National Academy of Sciences 114.23 (2017): 6005-6009. PubMed PMID: 28533371. PubMed Central PMCID: PMC5468623.
11. Várnai, Péter, and Tamas Balla. "Live cell imaging of phosphoinositides with expressed inositide binding protein domains." Methods 46.3 (2008): 167-176. PubMed PMID: 18930153. PubMed Central PMCID: PMC2644460.
12. Querido, Emmanuelle, and Pascal Chartrand. "Using fluorescent proteins to study mRNA trafficking in living cells." Methods in cell biology 85 (2008): 273-292. PubMed PMID: 18155467.
13. Chen, Baohui, et al. "Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system." Cell 155.7 (2013): 1479-1491. PubMed PMID: 24360272. PubMed Central PMCID: PMC3918502.
14. Chernov, Konstantin G., et al. "Near-infrared fluorescent proteins, biosensors, and optogenetic tools engineered from phytochromes." Chemical reviews 117.9 (2017): 6423-6446. PubMed PMID: 28401765.
15. Takai, Akira, et al. "Expanded palette of Nano-lanterns for real-time multicolor luminescence imaging." Proceedings of the National Academy of Sciences (2015): 201418468. PubMed PMID: 25831507 PubMed Central PMCID: PMC4394297.
16. Iwano, Satoshi, et al. "Single-cell bioluminescence imaging of deep tissue in freely moving animals." Science 359.6378 (2018): 935-93. PubMed PMID: 29472486.
17. Adjobo-Hermans, Merel JW, et al. "Real-time visualization of heterotrimeric G protein Gq activation in living cells." BMC biology 9.1 (2011): 32. PubMed PMID: 21619590. PubMed Central PMCID: PMC3129320.
18. Mastop, Marieke, et al. "A FRET-based biosensor for measuring Gα13 activation in single cells." PloS one 13.3 (2018): e0193705. PubMed PMID: 29505611. PubMed Central PMCID: PMC5837189.
Additional Resources on the Addgene Blog
- Read our Introduction to GFP
- Read our Fluorescent Proteins Blog Posts
- Download the Fluorescent Proteins 101 eBook
Resources on Addgene.org
- Browse the Fluorescent Protein Collection
- Find empty backbones for fluorescent protein fusions
- Browse the Fluorescent Proteins Plasmid Kits






Leave a Comment