 This post was contributed by guest blogger, Eugenia Rojas.
This post was contributed by guest blogger, Eugenia Rojas.
A question worthy of a PhD: How do you visualize protein turnover within a neuron?
For my PhD I studied a synaptic protein that is linked to neurodegeneration. The level of this protein is decreased in Alzheimer’s disease patient’s brains. However, it is not known why or how this happens. Therefore, I set out to study how protein turnover is regulated in neurons.
Protein turnover is the combined effect of protein synthesis and degradation. These processes control the abundance of proteins in cells. Homeostatic regulation of protein abundance is essential for normal cell function, and, because I was studying a synaptic protein, I wanted to measure it’s levels in its normal neuronal context. In other words, I needed a method for visualizing protein turnover in situ in neurons.
The classic approach for studying protein turnover is radioactive pulse-chase experiments. However, these experiments are problematic because they require radiolabeled amino acids. The use of radioactive materials is highly regulated, requires designated work areas and come with health risks. Importantly, with this method it is not possible to follow turnover of individual proteins in situ.
For the visualization of specific proteins in situ immunocytochemistry is perhaps the most common approach. With a good and specific antibody and the proper microscope setup we can observe the whole population of one particular protein in neurons or in any other cell type. However, we cannot distinguish new from old proteins within that pool. Therefore, it is not possible to visualize the localization of protein synthesis.
Alternatively, fluorescent protein tags can be used to facilitate the visualization of specific proteins in cells. Tagged protein can easily be overexpressed from a plasmid but overexpression may interfere with the system. Another method for studying protein synthesis in situ is the knock-in of fluorescent tags into the gene locus of interest. An advantage of this alternative is the possibility of real-time imaging. Robert Singer’s lab at Albert Einstein College of Medicine has recently published techniques to follow translation in living single cells. However, before CRISPR-Cas9 these techniques were quite tedious and, even here, there is the question of whether or not the tags influence behavior.
A conference and a breakthrough!
During my graduate studies I attended a scientific symposium called Horizons in Molecular Biology. This annual symposium is organized by international PhD students at the Max Planck Institute for Biophysical Chemistry, Göttingen in Germany. I attended this symposium seeking inspiration to resolve the technical hurtle of monitoring protein turnover of specific proteins in situ. I was in luck when I attended a very interesting talk by Prof. Erin M Schuman, a former Caltech researcher, now Director at the Max Planck Institute for Brain Research in Frankfurt am Main, Germany. She presented a novel technology, back then still unpublished, that allowed the visualization of specific de novo synthesized proteins in situ. I was privileged to be invited to learn from them at the MPI in Frankfurt am Main and perform similar experiments.
Visualizing newly synthesized proteins with FUNCAT
Professor Schuman’s technique applies metabolic labeling using a noncanonical amino acid, azidohomoalanine (AHA). In the technique, AHA is incorporated into newly synthesized proteins instead of methionine. The labeled proteins can be detected using click-chemistry that covalently links the azido containing AHA to a fluorescent alkyne tag. The newly synthesized and tagged proteins can then be observed with fluorescence microscopy. This application of AHA labeling is known as fluorescence noncanonical amino acid tagging (FUNCAT, Dieterich et al. Nature Neuroscience, 2010) and allows the visualization of the whole de novo synthesized proteome. However, FUNCAT alone cannot be used to determine the turnover of one specific protein. Professor Schuman’s lab showed that FUNCAT’s usefulness can be extended to determine the turnover of a specific protein by combining it with the proximity ligation assay (PLA).
The clever combination of FUNCAT and PLA
PLA was originally published by the Söderberg lab in Sweden as a method for the visualization of protein-protein interactions in situ. In the standard PLA assay, primary antibodies directed towards two endogenous proteins of interests, suspected to interact, are added to a cellular sample. Both antibodies have a short antibody specific DNA strand added to them. Only if the proteins of interest are in close proximity to each other can the DNA strands take part in rolling circle DNA amplification. The amplified DNA segment is labeled with fluorescent probes that enable visualization of the location of the protein-protein interaction.
To study protein levels in situ, the Schumann lab used a modified version of PLA where one antibody was directed toward a protein of interest but the second antibody was directed toward AHA. Now the rolling circle amplification could only take place if the two antibodies recognized an endogenous protein of interest that also had incorporated the AHA tag after pulse labeling of a determined period of time. Thus, only newly synthesized proteins of interest were detected and later visualized by FUNCAT-PLA.
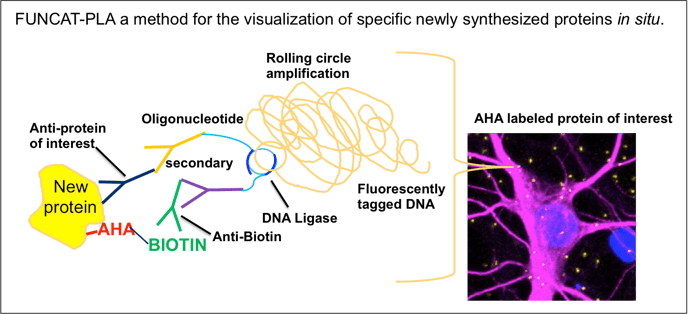
Before these methods were combined, it was not possible to visualize specific newly synthesized endogenous proteins in situ. Now FUNCAT-PLA provides both quantitative and spatio-temporal information about the turnover of specific endogenous proteins in any cell type. Exciting!
Many thanks to our guest blogger, Eugenia Rojas!
 Eugenia Rojas, PhD is a neuroscientist who has always been deeply interested in neurobiology. She is passionate about communicating science to all audiences. Right now, she is a science instructor at The Innovation Institute in Boston.
Eugenia Rojas, PhD is a neuroscientist who has always been deeply interested in neurobiology. She is passionate about communicating science to all audiences. Right now, she is a science instructor at The Innovation Institute in Boston.
References
1. Dieterich, Daniela C., et al. "In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons." Nature neuroscience 13.7 (2010): 897-905. PubMed PMID: 20543841. PubMed Central PMCID: PMC2920597.
2. Nathans, Daniel. "Puromycin inhibition of protein synthesis: incorporation of puromycin into peptide chains." Proceedings of the National Academy of Sciences 51.4 (1964): 585-592. PubMed PMID: 14166766. PubMed Central PMCID: PMC300121.
3. Söderberg, Ola, et al. "Direct observation of individual endogenous protein complexes in situ by proximity ligation." Nature methods 3.12 (2006): 995-1000. PubMed PMID: 17072308.
4. Dieck, Tom, et al. "Direct visualization of newly synthesized target proteins in situ." Nature methods 12.5 (2015): 411-414. PubMed PMID: 25775042. PubMed Central PMCID: PMC4414919.
5. Yarmolinsky, Michael B., and L. Gabriel. "Inhibition by puromycin of amino acid incorporation into protein." Proceedings of the National Academy of Sciences 45.12 (1959): 1721-1729. PubMed PMID: 16590564. PubMed Central PMCID: PMC222790.
6. Buxbaum, Adina R., Bin Wu, and Robert H. Singer. "Single β-actin mRNA detection in neurons reveals a mechanism for regulating its translatability." Science 343.6169 (2014): 419-422. PubMed PMID: 24458642. PubMed Central PMCID: PMC4121734.
Additional Resources on the Addgene Blog
- Fluorescent microscopy techniques - Which is best for me?
- Learn how small molecule fluorophores can be used for cell labeling
- Check out other protein tagging strategies
Additional Resources on Addgene.org
- Fluorescent protein plasmids
- Use FRET to study protein-protein interactions
- Deposit plasmids with Addgene!
Topics: Fluorescent Proteins, Fluorescent Imaging





Leave a Comment