A similar genetic code is used by most organisms on Earth, but different organisms have different preferences for the codons they use to encode specific amino acids. This is possible because there are 4 bases (A, T, C, and G) and 3 positions in each codon. There are therefore 64 possible codons but only 20 amino acids and 3 stop codons to encode leaving 41 codons unaccounted for. The result is redundancy; multiple codons encode single amino acids. Evolutionary constraints have molded which codons are used preferentially in which organisms - organisms have codon usage bias.
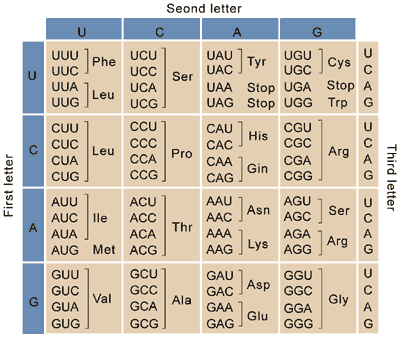 You can find many codon tables showing which codons encode which amino acids (see example to the right). With such simple rules, you might think it’s easy to come up with a workable DNA sequence to encode your peptide of interest and produce that peptide in your organism of choice. Unfortunately, codon preferences make it so you cannot choose among the possible codons at random and expect your sequence to express well in any organism.
You can find many codon tables showing which codons encode which amino acids (see example to the right). With such simple rules, you might think it’s easy to come up with a workable DNA sequence to encode your peptide of interest and produce that peptide in your organism of choice. Unfortunately, codon preferences make it so you cannot choose among the possible codons at random and expect your sequence to express well in any organism.
So what are the evolutionary constraints that lead to these preferences and what can we do about them? Read on to find out!
Why do organisms have different codon usage biases?
The reasons for varied codon preferences among organisms aren’t completely understood, but some possible reasons include:
- Metabolic pressures - it takes cellular resources to produce tRNAs that recognize different codons, modify the tRNAs correctly, and charge the tRNAs with the appropriate amino acids. If an organism uses only a subset of codons, it only needs to produce a subset of charged tRNAs and therefore may need fewer resources for the entire translation process. For example, during high growth rate conditions, E. coli preferentially upregulates production of tRNAs that recognize codons found in highly expressed genes (Emilsson and Kurland, 1990).
- Controlling gene expression through gene sequence - Proteins that are encoded by codons with low abundance or poorly charged tRNAs may be produced at a lower rate than proteins encoded by highly abundant, charged tRNAs. For example Tuller et al. found that translation efficiency is well correlated with codon bias in both E. coli and S. cerevisiae.
- Protein folding - If a protein is encoded by a mixture of codons with highly and poorly charged tRNAs, different regions of the protein may be translated at different rates. The ribosome will move quickly along regions calling for abundant, charged tRNAs but will stall at regions calling for low abundance, poorly charged tRNAs. When the ribosome stalls, this may give the swiftly translated regions a chance to fold properly. For example Pechmann and Frydman found that tracts of non-optimal codons are associated with specific secondary structures in 10 closely related yeast strains.
- Adaptation to changing conditions - Organisms often need to express genes at different levels under different conditions. With varied codon usage, an organism can change which proteins are highly expressed and which are poorly expressed by producing and charging specific tRNA pools. For example, tRNAs used in genes encoding amino acid biosynthetic enzymes may be preferentially charged during amino acid starvation thus resulting in higher production of amino acid biosynthetic enzymes (Dittmar et al., 2005).
How does codon usage bias affect my experiments?
While codon preferences can be very useful for organisms, they can be problematic for researchers trying to express proteins in heterologous hosts. If you simply amplify a gene of interest from the human genome, for instance, it may not express at all in E. coli (you can find a variety of databases showing various organisms’ codon preferences online). Even if the gene is translated, it may not function properly. This is the result of a mismatch between human and E. coli codon preference. Some codons commonly used in humans are not at all common in E. coli and vice versa. When translating these codons, the ribosome may therefore stall at inappropriate locations or fail to make it through the entire transcript resulting in the production of nonfunctional proteins and protein fragments respectively.
Solving the problem of codon usage bias - codon optimization and the expression of alternative tRNAs
Codon optimization
With low cost DNA synthesis, one of the primary ways researchers solve the problem of codon choice is to resynthesize genes in such a way that their codons are more appropriate for the desired expression host. This is known as “codon optimization.” Though simple in theory, this is not as easy as it sounds. Even for relatively short peptides, there can be many possible ways to encode them and what constitutes the “appropriate” codon is not necessarily obvious.
You might think, “Nonsense! I should just choose the codon with the most abundant pool of charged tRNAs in my host organism for every amino acid I’d like to encode,” but, as described above, not every region of a protein should necessarily be translated rapidly to produce a protein that functions properly.
You might then think, “Okay, I’ll just make sure the abundances of the codons I choose for the host match the abundances of codons used in the native organism.” This is possibly a better idea and has been used successfully in the past (Angov et al., 2008), but there are still many more features to consider when designing a full gene. A non-exhaustive list includes:
- Codon abundance relative to cognate tRNA abundance
- Repetitive sequences
- Restriction sites
- Sequences prone to create secondary structures in RNA transcripts
- Effects on transcription (remember, it’s not all about translation - e.g. codon choice may interrupt transcription factor binding sites)
As you might imagine, it’s not easy for humans to balance all of these factors on their own. Luckily, many researchers have created codon optimization algorithms and DNA synthesis companies such as IDT and GenScript host online codon optimization tools. Keep in mind that, just because you optimize a gene with one of these tools, it doesn’t necessarily mean the gene is going to express well. If you do get good expression, you should also functionally analyze the protein produced to ensure that it has folded properly.
You may be able to avoid getting your genes of interest codon optimized by ordering plasmids containing them from Addgene. If a plasmid at Addgene contains a gene that’s been codon optimized for a particular organism, this will sometimes (but not always) be noted in in the “mutation” field on the plasmid page (see plasmid 87904 for example). As many plasmids available from Addgene now have full sequence data, we recommend directly analyzing gene sequences for codon optimization and suitability for your expression host before using them in your experiments.
Expression of alternative tRNAs
If you don’t have the time or the funds to synthesize a codon optimized version of your gene of interest, it’s possible to overexpress low abundance tRNAs in your expression host and thereby increase their abundance. For example, the commercial Rosetta E. coli strains express a variety of tRNAs that are normally found at low abundance in E. coli.
The advantage of producing additional tRNAs is that you can use the same expression system for many different genes without having to create new constructs. However, due to problems such as mismatched translation rates and potential effects on cell growth, even hosts producing alternative tRNAs may not express sufficient amounts of your protein of interest.
Regardless of which method you choose to overcome the problems surrounding codon choice, you should have some method to make sure the proteins you produce function properly. Overexpression can result in the production of insoluble, nonfunctional globs of protein known as inclusion bodies that will generally segregate with the cell pellet during purification procedures. Even if you produce a large amount of protein in your expression host of choice, you should perform a functional assay to make sure your protein isn’t forming inclusion bodies and is folding properly.
References
1. Angov, Evelina, et al. "Heterologous protein expression is enhanced by harmonizing the codon usage frequencies of the target gene with those of the expression host." PloS one 3.5 (2008): e2189. PubMed PMID: 18478103. PubMed Central PMCID: PMC2364656.
2. Dittmar, Kimberly A., et al. "Selective charging of tRNA isoacceptors induced by amino‐acid starvation." EMBO reports 6.2 (2005): 151-157. PubMed PMID: 15678157. PubMed Central PMCID: PMC1299251.
3. Emilsson, Valur, and Charles G. Kurland. "Growth rate dependence of transfer RNA abundance in Escherichia coli." The EMBO journal 9.13 (1990): 4359-4366. PubMed PMID: 2265611. PubMed Central PMCID: PMC552224.
4. Gustafsson, Claes, Sridhar Govindarajan, and Jeremy Minshull. "Codon bias and heterologous protein expression." Trends in biotechnology 22.7 (2004): 346-353. PubMed PMID: 15245907.
5. Maertens, Barbara, et al. "Gene optimization mechanisms: A multi‐gene study reveals a high success rate of full‐length human proteins expressed in Escherichia coli." Protein Science 19.7 (2010): 1312-1326. PubMed PMID: 20506237. PubMed Central PMCID: PMC2970903.
6. Pechmann, Sebastian, and Judith Frydman. "Evolutionary conservation of codon optimality reveals hidden signatures of cotranslational folding." Nature structural & molecular biology20.2 (2013): 237. PubMed PMID: 23262490. PubMed Central PMCID: PMC3565066.
7. Quax, Tessa EF, et al. "Codon bias as a means to fine-tune gene expression." Molecular cell 59.2 (2015): 149-161. PubMed PMID: 26186290. PubMed Central PMCID: PMC4794256.
- This review provides a great overview of codon usage bias
8. Tuller, Tamir, et al. "Translation efficiency is determined by both codon bias and folding energy." Proceedings of the National Academy of Sciences 107.8 (2010): 3645-3650. PubMed PMID: 20133581. PubMed Central PMCID: PMC2840511.
Additional resources on the Addgene blog
- Plasmids 101: protein expression
- Plasmids 101: E. coli strains for protein expression
- Plasmids 101 featured topic page
Resources on Addgene.org
Topics: Plasmids 101, Plasmids






Leave a Comment