Have you ever wanted to selectively kill a subset of cells in your model system? Turns out that with light-inducible photosensitizers and a quick zap of the proper color light, you can do just that. Photosensitizing dyes and proteins have been around for awhile (check out this review), but the Bruchez and Tsang labs recently developed a photosensitizer composed of the protein complex complex and the MG-2I-dL5** fluorogen that can be used to ablate cells in culture and in vivo. Read on to learn more about this killer illumination technique!
What is a photosensitizer?
First, for a few definitions. A photosensitizer is a dye or protein that absorbs light to produce reactive oxygen species (ROS). The ROS produced by photosensitizers inactivate target proteins or cells while having little effect on neighboring cells. The photosensitizer used in this work is composed of two parts: 1) an activatable protein (the FAP - fluorogen activating protein - dL5** in this case), and 2) a fluorogen (the TAP - targeted and activated photosensitizer - MG-2I in this case). The fluorogen is not fluorescent until it binds its protein partner thus forming the full FAP-TAP complex (MG-2I-dL5**). A FAP-TAP complex will light up if excited with light of specific wavelength and also produce ROS - targeted cell ablation via ROS is achieved through localized expression of the FAP.
A new photosensitizer: FAP-TAP MG-2I-dL5**
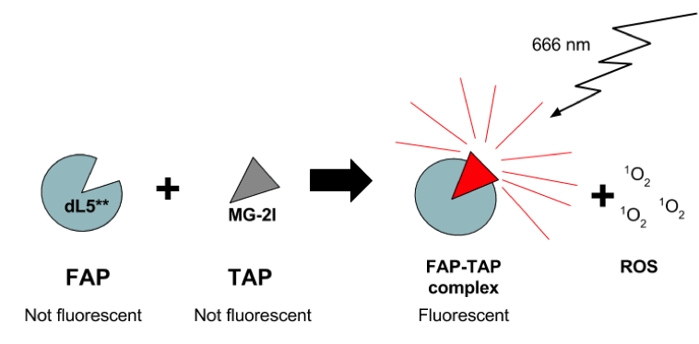
MG-2I-dL5** is a new FAP-TAP complex where dL5** is the FAP. It’s job is to bind MG-2I, a photosensitizer fluorogenic dye. MG-2I is the TAP portion of the complex and is derived from the malachite green (MG) dye, but has been modified with the addition of two iodine residues. Compared to MG, MG-2I has increased ROS generation when bound by the dL5** FAP. The iodine-modifications also shift MG-2I’s excitation wavelength to 666 nm, which is within the near-infrared (NIR) spectrum (>620 nm). Above is a brief schematic of how the MG-2I-dL5** FAP-TAP system works.
Find the Plasmids from the Publication Here
Light-induced cell ablation with MG-2I-dL5**
 He et al tested out light-induced cell ablation with MG-2I-dL5** in two different experimental settings: HEK293 cell culture and in live zebrafish. For cell culture experiments, only dL5**-expressing cells treated with MG-2I dye and exposed to light stained dead within 30 minutes of exposure. 90 seconds of illumination were sufficient for 100% effective killing of dL5** cells without death of neighboring bystander wild-type cells.
He et al tested out light-induced cell ablation with MG-2I-dL5** in two different experimental settings: HEK293 cell culture and in live zebrafish. For cell culture experiments, only dL5**-expressing cells treated with MG-2I dye and exposed to light stained dead within 30 minutes of exposure. 90 seconds of illumination were sufficient for 100% effective killing of dL5** cells without death of neighboring bystander wild-type cells.
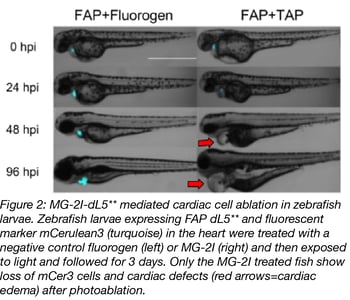
Zebrafish experiments with MG-2I-dL5** were performed with embryos and adult fish. Zebrafish are an ideal model organism for these experiments because developing embryos are transparent, so it’s easy to watch development in real time. In both cases, dL5** was tagged with mCerulean3 (mCer3) for visualization and expressed from a heart-specific promoter. Fish were labeled with MG-2I, either by incubating embryos in a MG-2I solution, or injecting adult fish with the dye. Embryos exposed to just 12 minutes of illumination had fewer mCer3-labeled cells in their hearts 24 hours post treatment and no mCer3 cells present 96 hours post treatment (see figure 2). Induced cell death was confirmed by TUNEL staining. The video below shows what happens to the zebrafish’s heartbeat and blood circulation of following FAP-TAP light activation.
Video: https://www.youtube.com/watch?v=xOtH5bs4HiI
Advantages of MG-2I-dL5** over other photosensitizers
One of the main advantages of MG-2I-dL5** over other genetically encoded photosensitizers is that it works with NIR light. NIR wavelengths have better tissue penetration than shorter wavelengths, which makes it possible to do things like in vivo imaging. Excitation with NIR light also produces less non-specific ROS because NIR light doesn’t excite natural biological chromophores found in cells. He et al also show that MG-2I-dL5** only generates small amounts of non-specific ROS when exposed to YFP and RFP excitation wavelengths, which suggests that it could be used with these fluorescent proteins.
He et al also compared MG-2I-dL5** head-to-head with KillerRed, a genetically encoded red fluorescent photosensitizer protein that generates ROS when illuminated with green light. They found that KillerRed required longer exposure times (5 minutes vs 10 seconds) to achieve the same level of killing as Mg-2I-dL5**. Mg-2I-dL5** also had higher photostability than KillerRed, with KillerRed reaching greater than 75% photobleaching after just 1 minute of exposure. MG-2I-dL5** experienced minor bleaching after longer exposure times. Additionally, KillerRed requires mitochondrial targeting to increase cell ablation efficiency, but dL5** phototoxocity is induced when expressed on the cell surface, in the cytosol, in mitochondria, or in cell nuclei. The two photosensitizers had similar rates of off-target damage to neighboring cells.
TL;DR MG-2I-dL5** is a NIR photosensitizer that allows for precise light-based control of protein inactivation/cell ablation while having limited collateral damage to bystander cells. Are you ready to use MG-2I-dL5** with your research? Let us know about your experiences using photosensitizers in the comments below!
References
1. He, J., Wang, Y., Missinato, M. A., Onuoha, E., Perkins, L. A., Watkins, S. C., . . . Bruchez, M. P. (2016). A genetically targetable near-infrared photosensitizer. Nature Methods,13(3), 263-268. doi:10.1038/nmeth.3735. PubMed PMID: 26808669. PubMed Central PMCID: PMC4916159.
2. Jacobson, K., Rajfur, Z., Vitriol, E., & Hahn, K. (2008). Chromophore-assisted laser inactivation in cell biology. Trends in Cell Biology,18(9), 443-450. doi:10.1016/j.tcb.2008.07.001. PubMed PMID: 18706812. PubMed Central PMCID: PMC4445427.
3. Bulina, M. E., Chudakov, D. M., Britanova, O. V., Yanushevich, Y. G., Staroverov, D. B., Chepurnykh, T. V., . . . Lukyanov, K. A. (2005). A genetically encoded photosensitizer. Nature Biotechnology,24(1), 95-99. doi:10.1038/nbt1175. PubMed PMID: 16369538.
Additional Resources on the Addgene Blog
- Read Other Optogenetics Blog Posts
- Sign Up to Receive the Fluorescent Protein eBook
- Learn about Small Molecule Fluorophores
Resources on Addgene.org
- Visit Our Fluorescent Protein Guide Page
- Check out MiniSOG2 and SuperNova, two other photosensitizers used for cell ablation







Leave a Comment