Hopefully you know that, if you’re an academic researcher at a nonprofit institution, you can order plasmids covering a wide range of fields from Addgene. What you might not know is that Addgene distributes curated collections of plasmids as kits with greatly reduced costs per plasmid. We generally create kits to allow you to easily use a set of plasmids together on a tight budget.
To help popularize our kit resources, we’re highlighting just a few of the many useful kits you can order from the repository:
- FX Cloning Kit
- BIOFAB Plasmid Set
- pCri System
- Cell Free Protein Expression Kit
- FREQ Seq
- PUF Assembly Kit
Below you’ll find information about what these kits contain, how they can be used, and how they’ve been used to date . If you have any questions, please write to us in the comments section.

FX cloning kit
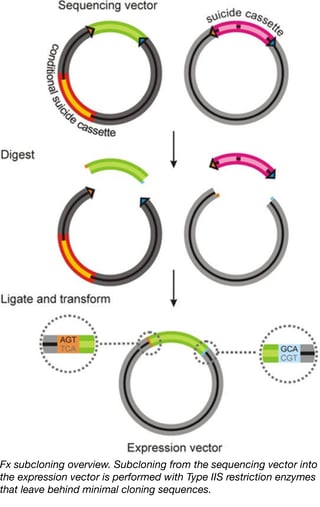 This kit is designed to be used with Type IIS restriction enzymes and standardized primers to clone your gene of interest into both sequencing (pInitial) and protein production (a variety of pExpression vectors) plasmids. The type IIS restriction sites used in these vectors are of low abundance (meaning they are unlikely to be found in your gene of interest) and prevent incorporation of extensive sequences used for cloning into the final translated products. Negative selection markers built into all plasmids in the kit help decrease false positives observed in the cloning process, making it easier to produce the constructs you need. Expression vectors within this kit contain a variety of promoters to facilitate gene expression in bacteria, yeast, mammalian cells, insects, and even Xenopus oocytes. The expression vectors additionally contain various combinations of protein tags (His, Avi, and GFP) as well as protease sites to facilitate protein purification. We hope this kit will allow you to quickly study your gene of interest in a variety of model systems, thereby opening your research to a variety of experimental opportunities.
This kit is designed to be used with Type IIS restriction enzymes and standardized primers to clone your gene of interest into both sequencing (pInitial) and protein production (a variety of pExpression vectors) plasmids. The type IIS restriction sites used in these vectors are of low abundance (meaning they are unlikely to be found in your gene of interest) and prevent incorporation of extensive sequences used for cloning into the final translated products. Negative selection markers built into all plasmids in the kit help decrease false positives observed in the cloning process, making it easier to produce the constructs you need. Expression vectors within this kit contain a variety of promoters to facilitate gene expression in bacteria, yeast, mammalian cells, insects, and even Xenopus oocytes. The expression vectors additionally contain various combinations of protein tags (His, Avi, and GFP) as well as protease sites to facilitate protein purification. We hope this kit will allow you to quickly study your gene of interest in a variety of model systems, thereby opening your research to a variety of experimental opportunities.
The plasmids in this kit have been used to purify a variety of peptides including antibiotic efflux pumps and ankyrin seven-repeat proteins:
BIOFAB plasmid set
 3000 Series
3000 Series
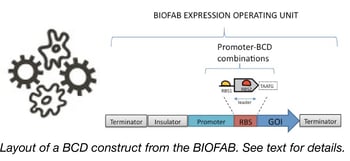
Confronted with the problem of creating vectors for predictable protein expression in bacteria, researchers at the BIOFAB and the Endy Lab created this kit. The plasmids within it contain a variety of different promoters followed by leader sequences (encoding small leader peptides) downstream of which you can clone your gene of interest. Translation of the standardized leader peptide prevents secondary structures in the mRNA encoding your gene of interest from affecting its translation initiation and instead makes initiation heavily dependent upon the core of a Shine-Delgarno sequence found at the 3’ end of the leader sequence. Using a variety of GFP and RFP fusions as their test genes, the researchers at the BIOFAB showed that expression of many different genes was predictable based solely on the promoters and Shine-Delgarno sequences used in this so-called bicistronic design (BCD). The 49 vectors in this kit can be used to drive production of your favorite gene at your level of choice and should be useful for a variety of synthetic biology or protein purification studies in E. coli.
800 and 1900 Series
If you’re expressing two genes from a single transcript, or if you want to prevent read-thru transcription from resulting in the production of a downstream peptide, the transcription terminators in this series are just the tool you need. The plasmids in this portion of the BIOFAB Plasmid Set contain terminators of varying strengths centered within GFP/RFP bicistronic reporters characterized in Cambray et al., 2013. These terminators can be used to precisely tune your polycistronic E. coli expression constructs.
Some applications of plasmids from this kit include the production of a suicide switch in cyanobacteria, synthetic inverters in E. coli, and biosensors in E. coli:
pCri system
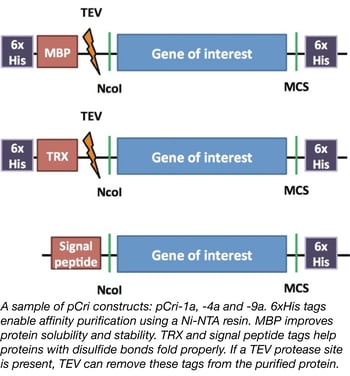 The Gomis-Ruth lab designed the pCri system to simplify the protein purification process by making it easy for a researcher to quickly generate N- and C-terminal epitope tag fusions to a protein of interest. The plasmids used to generate these fusions all use the same restriction enzymes for cloning. Furthermore, some plasmids contain peptides to facilitate protein folding and/or protease sites to remove the epitope tags. Different plasmids in the kit can be used to purify proteins from E. coli, Bacillus subtilis, or Pichia pastoris depending upon your specific protein folding needs.
The Gomis-Ruth lab designed the pCri system to simplify the protein purification process by making it easy for a researcher to quickly generate N- and C-terminal epitope tag fusions to a protein of interest. The plasmids used to generate these fusions all use the same restriction enzymes for cloning. Furthermore, some plasmids contain peptides to facilitate protein folding and/or protease sites to remove the epitope tags. Different plasmids in the kit can be used to purify proteins from E. coli, Bacillus subtilis, or Pichia pastoris depending upon your specific protein folding needs.
The pCri system has recently been used to study a matrix metalloproteinase:
You can find more info on pCri in Mary Gearing’s blog post on the system
Cell-free protein expression kit
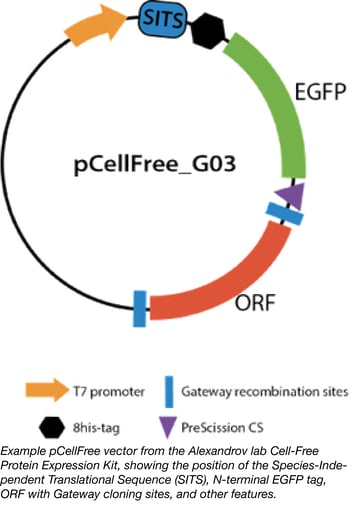 Cell-free expression systems have been improved for both eukaryotic and prokaryotic expression in recent years, with many different systems now available. However, it can be difficult to compare their performance, as most translation initiation sequences are optimised for a particular system. The Alexandrov lab at the University of Queensland’s Institute for Molecular Bioscience has provided a solution for this problem by developing a general translation initiation sequence known as the Species-Independent Translational Sequence (SITS). This sequence bypasses the 5′ mRNA cap required for eukaryotic systems and instead directly engages the ribosome for translational complex assembly.
Cell-free expression systems have been improved for both eukaryotic and prokaryotic expression in recent years, with many different systems now available. However, it can be difficult to compare their performance, as most translation initiation sequences are optimised for a particular system. The Alexandrov lab at the University of Queensland’s Institute for Molecular Bioscience has provided a solution for this problem by developing a general translation initiation sequence known as the Species-Independent Translational Sequence (SITS). This sequence bypasses the 5′ mRNA cap required for eukaryotic systems and instead directly engages the ribosome for translational complex assembly.
PhD student Dejan Gagoski used SITS to create the pCell-Free vectors, Gateway-compatible backbones that enable cell-free expression of proteins in both prokaryotic and eukaryotic cell extracts (Gagoski et al 2015). He has also constructed a library of eGFP-tagged human ORF clones to allow testing and comparison of different cell-free expression systems; the Cell-free expression test kit represents a set of 88 clones in a pCellFree vector that enables protein expression in any in vitro translation system. The gene set was carefully chosen to perform statistically relevant benchmarking of cell-free expression systems, and tested for product integrity, expression level, and aggregation propensity in four expression systems: E. coli, wheat germ, HeLa, and Leishmania (Gagoski, et al 2015[2]). The proteins encoded in this set range in size from 4 to 156 kDa, enabling the user to characterise the efficiency of their cell-free system in correlation to the size of the product. Analysis of protein size and expression level can be conveniently performed through the N-terminal eGFP tag carried on all constructs.
The Cell-Free Protein Expression Kit has been used to produced recombinant proteins for in-vitro assays:
FREQ-Seq
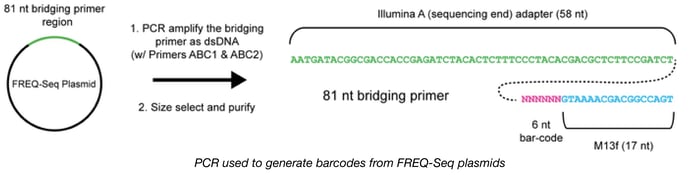 To help scientists track the frequencies of specific alleles in microbial populations through time, Christopher Marx's lab has engineered and validated a new method called FREQ-Seq. This strategy allows scientists to construct barcoded, locus-specific libraries compatible with Illumina next generation sequencing in order to study evolutionary dynamics. By counting DNA sequence reads, FREQ-Seq can quantitatively determine allele frequencies across timepoints or populations. This kit consists of a 48 plasmid adapter library, with each plasmid carrying a unique barcoded Illumina-M13F bridging primer. These bridging primers are amplified and used to generate Illumina sequencing libraries using a 2-step PCR-based protocol and are compatible with single-end or paired-end read flow cells. FREQ-Seq is an open source platform and the libraries can be generated in a cost-effective manner with minimal bias.
To help scientists track the frequencies of specific alleles in microbial populations through time, Christopher Marx's lab has engineered and validated a new method called FREQ-Seq. This strategy allows scientists to construct barcoded, locus-specific libraries compatible with Illumina next generation sequencing in order to study evolutionary dynamics. By counting DNA sequence reads, FREQ-Seq can quantitatively determine allele frequencies across timepoints or populations. This kit consists of a 48 plasmid adapter library, with each plasmid carrying a unique barcoded Illumina-M13F bridging primer. These bridging primers are amplified and used to generate Illumina sequencing libraries using a 2-step PCR-based protocol and are compatible with single-end or paired-end read flow cells. FREQ-Seq is an open source platform and the libraries can be generated in a cost-effective manner with minimal bias.
The FREQ Seq kit has been used in microbiology studies monitoring sequence dynamics under various selective pressures:
PUF Assembly Kit
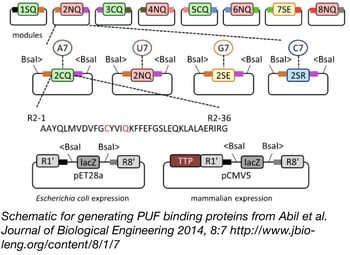 PUF proteins are composed of repeat segments that can be engineered to bind specific RNA bases. By swapping out these different modular repeat segments, you can create a PUF protein that binds your RNA sequence of interest. The Zhao lab has made this easy with their PUF Assembly Kit. The kit contains all the variant PUF segments required to bind any 8 base RNA sequence. The segments are provided in Gateway compatible vectors that allow you to quickly clone your segments of choice into either an E. coli or a mammalian expression vector in a one-pot reaction. Using this system, you can create custom RNA binding proteins in as little as 3 days.
PUF proteins are composed of repeat segments that can be engineered to bind specific RNA bases. By swapping out these different modular repeat segments, you can create a PUF protein that binds your RNA sequence of interest. The Zhao lab has made this easy with their PUF Assembly Kit. The kit contains all the variant PUF segments required to bind any 8 base RNA sequence. The segments are provided in Gateway compatible vectors that allow you to quickly clone your segments of choice into either an E. coli or a mammalian expression vector in a one-pot reaction. Using this system, you can create custom RNA binding proteins in as little as 3 days.
Abil et al., 2014 show that you can fuse custom PUF proteins to a translational repressor to down-regulate a gene of interest. You can generate PUF proteins with a wide variety of functions via different fusions. Some applications of PUF protein fusions reported in the literature include RNA detection, RNA cleavage, and translational repression or activation:
See Kendall Morgan’s blog post on the PUF Assembly Kit for some insights on its potential from its creators.
We hope you’ll find these kits useful in your research. If you’d ever like to deposit a kit, please contact us at depost@addgene.org.
Topics: Other Plasmid Tools, Plasmids






Leave a Comment