We are excited to announce the recipients of this year’s Michael Davidson and Roger Tsien Commemorative Conference Awards. Congratulations to the five winners!
Every two years, we seek applications for awards to early career scientists using fluorescent proteins in their research. These awards help early career researchers attend a conference of their choice while honoring the late Michael Davidson and Roger Tsien who have played enormous roles in the development and dissemination of fluorescent protein tools.
This year, we’ve made some changes. In addition to supporting in-person conference attendance, the award now supports virtual conference attendance. We also opened up applications to research technicians and undergraduate researchers, in addition to postdocs and graduate students as we have done in the past.
Below, read about the five winners and their research.
Abhi Aggarwal: Glutamate sensing reporters
 Abhi Aggarwal is a postbaccalaureate research technician in Kaspar Podgorski’s lab at Janelia Research Campus, HHMI where he works on developing and optimizing biosensors in the lab. He has recently developed a third-generation intensiometric glutamate sensing fluorescent reporter (iGluSnFR3) to study glutamate, a signaling molecule in the brain. “We hope that iGluSnFR3 will enable significant steps forward in our understanding of synaptic and circuit functions in health and disease,” says Aggarwal.
Abhi Aggarwal is a postbaccalaureate research technician in Kaspar Podgorski’s lab at Janelia Research Campus, HHMI where he works on developing and optimizing biosensors in the lab. He has recently developed a third-generation intensiometric glutamate sensing fluorescent reporter (iGluSnFR3) to study glutamate, a signaling molecule in the brain. “We hope that iGluSnFR3 will enable significant steps forward in our understanding of synaptic and circuit functions in health and disease,” says Aggarwal.
This biosensor changes fluorescence based on the release of glutamate from neurons and this new version of the biosensor is faster and more sensitive than its predecessors. Currently, Aggarwal, and colleagues Filip Tomaska and Tim Hanson, are characterizing variants of iGluSnFR3 in vivo and have used it to detect glutamate release from hundreds of synapses at once. “This work has only been made possible due to the collaborative and highly interactive environment that Janelia offers,” says Aggarwal.
In addition to the biosensors, Aggarwal and colleagues are working on developing new surface display vectors to improve expression and trafficking of these biosensors.
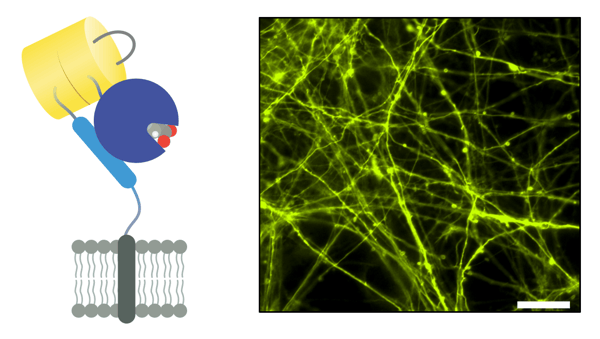 |
| Figure 1: Schematic of iGluSnFR3 (left). GluSnFR3 in pMinDisplay (PDGFR anchoring domain), in electroporated in rat hippocampal cultured neurons, imaged 14 days post-transfection, excited at 505 nm (right). Scalebar represents 100 µm. Image from Abhi Aggarwal. |
Brandán Pedre Pérez: Redox biosensors

Brandán Pedre Pérez is a postdoc at the German Cancer Research Center (DKFZ) who has developed new redox sensitive GFP2 (roGFP2)-based biosensors for live cells. His work focuses on the enzyme hydropersulfide-propagating enzymes 3-mercaptopyruvate sulfurtransferase (MPST) which catalyzes the desulfuration of 3-mercaptopyruvate and generates an enzyme-bound hydropersulfide that is then transferred to thiol-containing acceptors. These hydropersulfides modulate protein activity and have important roles during oxidative stress.
To monitor this activity, Pedre Pérez’s biosensor fuses MPST to roGFP2. Because roGFP2 contains two cysteines near the chromophore that are susceptible to redox modification, the MPST hydrosulfide can be transferred to the roGFP2 cysteine thiols and change its fluorescence properties. The biosensor can also be used along other thiol-containing acceptors in competition assays.
“I believe that improved biosensors or biosensors that detect new redox metabolites are vital for researchers to better understand the processes that happen inside the cell,” says Pedre Pérez.
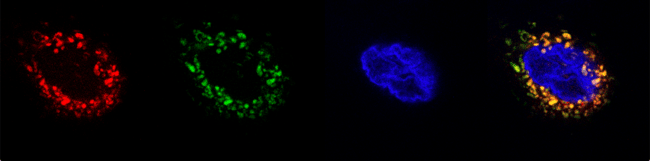 |
|
Figure 2: Micrographs of the biosensor expressed in the human cell line, Hep3B. From left to right: Mitotracker Red, GFP for the biosensor, Hoechst, and overlay). |
Elizabeth Hubbard: Neurons involved in opioid withdrawal
 Elizabeth Hubbard is a fourth year Ph.D. candidate at the University of California, Irvine where she studies opioid use disorder. Her goal is to identify cell populations in the brain that contribute to the development of behavioral signs of opioid withdrawal before they occur.
Elizabeth Hubbard is a fourth year Ph.D. candidate at the University of California, Irvine where she studies opioid use disorder. Her goal is to identify cell populations in the brain that contribute to the development of behavioral signs of opioid withdrawal before they occur.
First, she used a genetic method to tag neurons activated by opioid withdrawal with tdTomato and assessed cFos expression. She will explore the connectivity patterns of these cells and the functional contribution to a variety of behaviors related to addiction and pain. Once these are completed, Hubbard will image these neurons using GCaMP7, measuring calcium signaling related fluorescence during morphine administration and withdrawal to explore how these cells may encode the negative affect associated with drug withdrawal.
“Together, this project will allow us to better understand how neuron activity changes following opioid withdrawal, providing insight into new methods for treating patients in recovery from opioid use disorder,” says Hubbard.
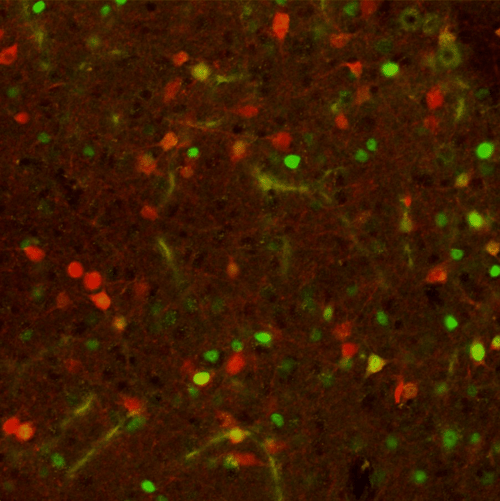 |
| Figure 3: 20x confocal image of the periaqueductal gray (PAG), with red neurons expressing tdTomato and green neurons expressing cFos. Image from Elizabeth Hubbard. |
Haoyi Qiu: Pain response in mechanical allodynia
 Haoyi Qiu is a M.Sc. student at McGill University in the department of Physiology. Her research focuses on the neural circuitry involved in mechanical allodynia, a condition where a light touch creates a painful sensation. The dorsal horn of the spinal cord is the first site of sensory input from our extremities. “Here, painful stimuli are processed by a network of excitatory and inhibitory neurons whose function and connectivity are poorly understood,” says Qiu.
Haoyi Qiu is a M.Sc. student at McGill University in the department of Physiology. Her research focuses on the neural circuitry involved in mechanical allodynia, a condition where a light touch creates a painful sensation. The dorsal horn of the spinal cord is the first site of sensory input from our extremities. “Here, painful stimuli are processed by a network of excitatory and inhibitory neurons whose function and connectivity are poorly understood,” says Qiu.
Qiu developed a dual virus system using both optogenetics and chemogenetics to study the activity of parvalbumin (PV)-expressing neurons, neurons that the lab previously found had a role in perceiving touch inputs. In this system, light activates the neurons (via oChIEF) and a chemogenetic agonist inhibits the neuron (via hM4D). She fused fluorescent protein tags to the receptors to monitor their localization and expression. Qiu saw that inhibition of the PV neurons resulted in mechanical allodynia. With the dual regulation, she could then use pulses of blue light to activate the neurons at specific firing frequencies. She hopes to identify how in vivo firing frequency of PV neurons affect the response to light touch in mechanical allodynia.
-min.png?width=600&name=Haoyi%20Addgene%20Feature%20(1)-min.png) |
| Figure 4: While Qiu’s work on the PV neurons is in progress, she has validated the dual virus method to study CR neurons which are depicted here. (A) Schematic diagram of the construction of the dual virus and the experimental approach. (B) CR-Cre mice demonstrate pain response due to blue light stimulation of dual virus-expressing CR neurons. (C) Duration of painful responses in the left and right paw due to blue light activation and CNO silencing. (D) Immunohistochemistry verification of the dual virus expression in CR neurons of the spinal cord. Created with BioRender.com. Image from Haoyi Qiu. |
He Tian: Directed evolution of biosensors
 He Tian (goes by “Tian”) is a postdoc at Harvard University where she engineers genetically encoded voltage indicators. Tian developed a video-based high-throughput screen for directed evolution of biosensors based on their dynamic properties. Her method takes a pooled library of mutant sensors expressed in a mammalian cell line engineered to be electrically excitable. The mutants are co-expressed with a photoconvertible fluorescent protein mEOS4a. Cells containing a sensor that shows a favorable outcome are then photoconverted, sorted, and sequenced to identify the mutants for further enrichment. Tian notes that this platform can be used to engineer many types of biosensors.
He Tian (goes by “Tian”) is a postdoc at Harvard University where she engineers genetically encoded voltage indicators. Tian developed a video-based high-throughput screen for directed evolution of biosensors based on their dynamic properties. Her method takes a pooled library of mutant sensors expressed in a mammalian cell line engineered to be electrically excitable. The mutants are co-expressed with a photoconvertible fluorescent protein mEOS4a. Cells containing a sensor that shows a favorable outcome are then photoconverted, sorted, and sequenced to identify the mutants for further enrichment. Tian notes that this platform can be used to engineer many types of biosensors.
This screening method allowed Tian to optimize a red‑shifted archaerhodopsin‑derived genetically encoded voltage indicator which had improved performance in live mouse brains. Currently, she is using the improved voltage indicator coming from the screen to understand mouse cortical and hippocampal neural microcircuits. “In the longer term, this screening platform may help us to create high-performance sensors to enable multiplexed imaging of neurotransmitter, calcium, and membrane voltage,” says Tian.
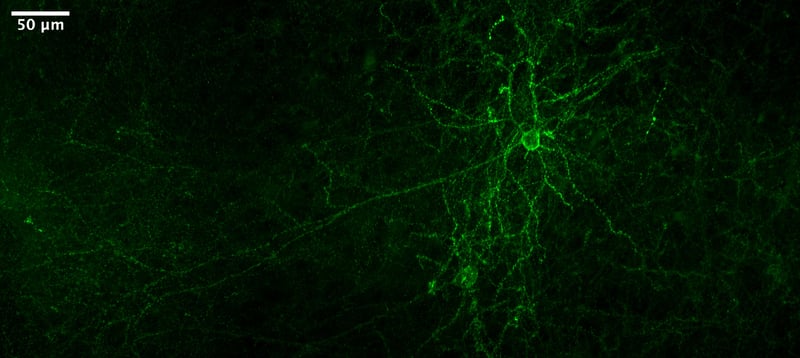 |
|
Figure 5: Citrine-tagged voltage indicator expressed in mouse cortex. Image from He Tian. |
Congrats again to all the recipients!
Topics: Fluorescent Proteins, Addgene News, Awards






Leave a Comment