 Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Every few months we highlight a subset of the new plasmids in the repository through our hot plasmids articles. These articles provide brief summaries of recent plasmid deposits and we hope they'll make it easier for you to find and use the plasmids you need. If you'd ever like to write about a recent plasmid deposit please sign up here.
Listen to this episode of Hot Plasmids!
Optogenetic control of CRISPR-Cas9 by CASANOVA
Article contributed by Cary Valley
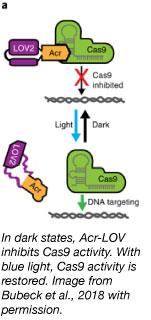 Anti-CRISPR (Acr) proteins potently inhibit type II CRISPR systems including the popular S. pyogenes Cas9 (Rauch et al., 2017, Hynes et al., 2017). However, Acr inhibition is spatially and temporally ubiquitous, making them a rather blunt instrument for controlling CRISPR. To bring greater control to Acr activity as a biological tool, Niopek and colleagues turned to optogenetics. Through the use of light-sensitive protein domains, they have developed a system to control CRISPR-Cas9 activity using blue light (Bubeck et al., 2018).
Anti-CRISPR (Acr) proteins potently inhibit type II CRISPR systems including the popular S. pyogenes Cas9 (Rauch et al., 2017, Hynes et al., 2017). However, Acr inhibition is spatially and temporally ubiquitous, making them a rather blunt instrument for controlling CRISPR. To bring greater control to Acr activity as a biological tool, Niopek and colleagues turned to optogenetics. Through the use of light-sensitive protein domains, they have developed a system to control CRISPR-Cas9 activity using blue light (Bubeck et al., 2018).
They accomplished this by inserting a photo-sensitive LOV2 domain from A. sativa into a loop region of the Acr protein AcrIIA4. In the dark, AcrIIA4 inhibits CRISPR activity. With the addition of blue light, the terminal helices of the LOV2 domain unfold, disrupting the protein conformation and thus, inhibiting function of Acr. This allows CRISPR-Cas9 to function on its target. They name their most potent Acr-LOV fusion protein inhibitor CASANOVA for “CRISPR-Cas9 activity switching via a novel optogenetic variant of AcrIIA4.” CASANOVA responds to blue light quickly. Within minutes of inactivating the CASANOVA complex, fluorescent CRISPR complexes were observed binding telomeric DNA. What’s great about CASANOVA? It works with the S. pyogenes Cas9, dCas9-effector fusions, and xCas9, and requires no additional chemicals or modifications. This versatility opens up the potential for spatial and temporal control in many genetic targeting and editing experiments.
Find the CASANOVA plasmids here!
Bubeck F et al., Nature Methods 2018. PubMed PMID: 30377362.
Studying neuronal activity using the fluorescent voltage indicator Archon
Article contributed by Shreya Vedantam
The neurons in the brain compute using electrical impulses to coordinate emotions, thoughts, and behavior. Historically, scientists study the electrical activity in the brain by inserting electrodes into the brain. But this process is time consuming and expensive. Thus scientists have turned to genetically encoded voltage indicators.
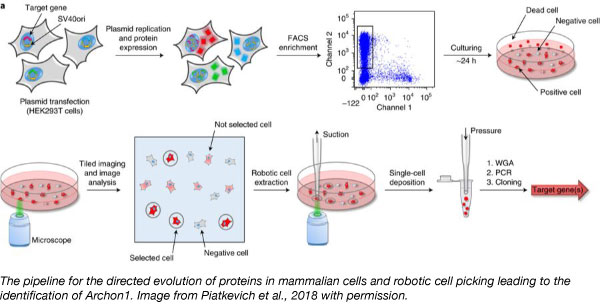
To expand the voltage indicator toolbox, Ed Boyden’s lab developed a way to rapidly screen thousands of proteins for one that can report electrical activity via imaging. They took the light-sensitive protein QuasAr2 and systematically mutated it. Using a robotic picking approach to select cells with variant characteristics they wanted, the team identified Archon1, a novel opsin-based fluorescent voltage indicator. Archon1 can embed itself into the cell membrane where cell voltage can then be measured. Archon1 is bright, exhibits good localization, has high signal:noise ratio, high sensitivity, responds quickly, is photostable, and is compatible with optogenetic control. The researchers also showed that Archon1 is functional in a wide range of animal models such as mice, C.elegans, and Zebrafish.
Find Archon1 plasmids at Addgene!
Piatkevich KD et al, Nat Chem Biol 2018. PubMed PMID:29483642.
Making protein-protein interaction studies easier with a 2in1 vector cloning system
Article contributed by Michelle Cronin
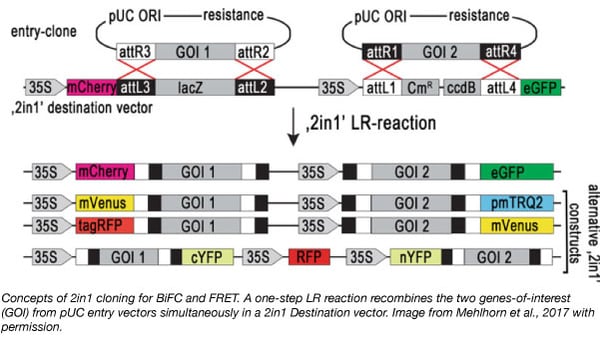
Recently, the Grefen lab introduced the “2in1” vector cloning system - a Gateway compatible approach to simultaneously clone two genes-of-interest (GOI) into two independent expression cassettes on a single plasmid (Grefen and Blatt, 2012). Similar to traditional Gateway™ cloning, the GOI first needs to be cloned into an entry vector. The 2in1 cloning system contains two entry vectors (pUC57-L1L4 and pUC57-L3L2) that offer the advantage of using restriction digest cloning to introduce the GOIs into the 2in1 vector instead of the GatewayTM cloning BP reaction, where insertion of the GOI occurs through recombination.
The 2in1 cloning system is well suited for protein-protein interaction (PPI) studies in plants that rely on transient transfection systems using Agrobacterium (such as Nicotiana benthamiana or Arabidopsis thaliana (Xing et al., 2016)). During Agrobacterium-mediated DNA transfer, each GOI is usually expressed from a different plasmid, which may lead to unequal gene dosage and high variability of protein coexpression. By introducing a single plasmid containing both GOIs, expression levels are more evenly controlled. The Grefen Lab has utilized the 2in1 cloning system to generate a series of plasmids to study PPI using Förster Resonance Energy Transfer (FRET, Heckler et al., 2015) or ratiometric Bimolecular Fluorescence Complementation (rBiFC, Grefen and Blatt, 2012). The 2in1 Plasmid Toolkit consists of 4 plasmid toolkits generated with 4 different, binary 2in1 vectors with all possible combinations of tag orientation (NN, NC, CN, and CC). The 4 kits also contain the two entry vectors pUC-L1L4 and pUC-L3L2.
Find the 2in1 plasmid kits at Addgene!
Mehlhorn DG et al, Methods Mol Biol, 2018. PubMed PMID: 29043675.
Grefen C and MR Blatt. Biotechniques 2012. PubMed PMID: 23066669.
Hecker A et al, Plant Physiol 2015. PubMed PMID: 25971551.
Discovery of new protein binder tools for photoswitchable proteins cPYP and AsLOV2
Article contributed by Angela Abitua
Reversible control of photoswitchable proteins, which change in protein conformation in response to light, offers a way to spatially and temporally control cellular processes in vivo. Many photoactive proteins exist in nature, but developing them into optogenetic tools can be difficult because this requires detailed structural knowledge of the proteins. To overcome this challenge, the Uppalapati and Woolley lab developed a phage display technique using small scaffold proteins to identify novel binding proteins that bind to either the light- and dark- states of the photoswitchable cPYP, which changes in conformation in response to blue light (445nm). The discovery and applications of protein partners that bind to only the light- or dark- state of the photoswitchable proteins offers a powerful approach to control protein-protein interactions in cells because the protein binders can be customized for protein visualization, relocation, degradation, modification, and scaffolding.
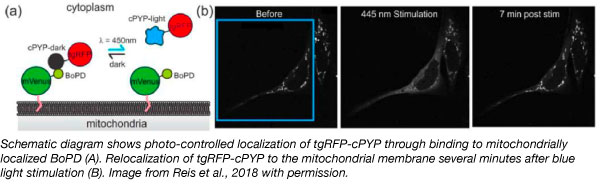
In their study, the Woolley lab showed that these photo-reversible protein-protein interactions can control subcellular localization processes in vivo. They deposited the resulting optogenetic plasmid tools which includes plasmids encoding a fusion between cPYP and tgRFP (PLL7.0 tgRFP cPYP), and plasmids that encode the protein binders (which bind to either the light- or dark- state of the photoswitchable proteins) fused to mVenus linked to a mitochondrial localization sequence TOM20 (pTriEX Tom20 mVenus BoPD). In dark adapted cells expressing these plasmids, tgRFP fluorescence localizes to the mitochondria, because cPYP binds to BoPD. A similar set of newly identified light- and dark- binder plasmid tools were also deposited for the photoswitchable AsLOV2.
Find the cPYP and AsLOV2 protein binder plasmids here!
Reis JM, et al. ACS Synthetic Biology. 2018 PubMed PMID: 30203962.
Topics: Hot Plasmids






Leave a Comment