This post was contributed by guest bloggers Becky Kucera, M.Sc. and Eric Cantor, Ph.D. from New England Biolabs.
Golden gate assembly limitations
Embraced by the synthetic biology community, Golden Gate Assembly is commonly used to assemble 2–10 DNA fragments in a single “one-pot” reaction to form complex, multi-insert modular assemblies that enable biosynthetic pathway engineering and optimization. However, current best practices for assemblies of more than 10 modules often rely on two-step hierarchical approaches using different Type IIS restriction enzyme specificities at each step. Factors such as enzyme efficiency, stability, and buffer compatibility have placed practical limits on single- or two-step assemblies.
We have reduced these limitations through the use of engineered Type IIS restriction enzymes and careful choice of junction sequences guided by experimentally-derived DNA Ligase fidelity data. Our work demonstrates that it is now possible to achieve 20+ fragment assemblies with both robust efficiency and accuracy.
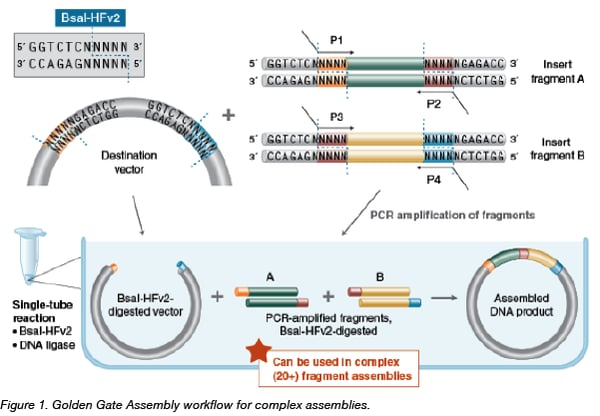
Golden Gate assembly involves the simultaneous activities of a Type IIS restriction enzyme such as BsaI, and a DNA ligase such as T4 DNA ligase. Inserts, either precloned or in amplicon form (shown above) are flanked by BsaI sites in an orientation that results in 5’ four base overhangs after cutting. The destination vector receives the assembly after similar Type IIS restriction enzyme cutting results in 5’ four base overhangs complementary to the overhangs of the assembled inserts. After ligation of the insert into the vector, the BsaI site is no longer present, allowing multiple fragments to be assembled in an ordered, pre-determined manner. If the original insert religates into its original vector, it will subsequently be re-digested and "eligible” for ligation into the destination construct. Reactions are often cycled between 37°C and 16°C to provide optimal reaction temps for both the restriction enzyme and ligase, thereby enhancing the ordered assembly of the target construct.
Development of Golden Gate Assembly test systems
Our research focused on three different levels of assembly:
- High efficiency and accurate single-insert assembly
- Intermediate 5- or 12-fragment assembly
- More complex 24-fragment assembly
Single insert cloning efficiencies were measured based on the acquisition of antibiotic selectable markers by the destination plasmid. Antibiotic selection allowed for high throughput cloning efficiency measurements. We performed additional efficiency measurements using similarly sized Lambda amplicon inserts to control for any suppression of background by antibiotic selection. Screening of transformants by colony PCR confirmed the insertion of the Lambda insert at the same high frequencies as the antibiotic selection markers.
The 5-, 12- and 24-assembly systems were based on the ability to correctly assemble a lacI/lacZ cassette (designed by NEB for use in optimization of assembly systems) to produce a blue colony phenotype upon growth on LB/Cam/X-gal/IPTG agar plates, indicating successful reconstruction of the coding sequence for beta-galactosidase in the lacI/lacZ cassette. Additional confirmation of accurate assemblies was achieved by the sequencing of plasmids isolated from blue or white colonies. Sequencing of blue colonies showed the expected complete sequence for the lacI/lacZ genes (1), while sequencing of white colonies showed a mixture of mis-assemblies and very occasional uncut or cut/re-ligated pGGA destination plasmids.
A final validation of the 5-, 12- or 24-fragment test systems was performed by setting up assembly reactions in which a single component was purposefully omitted. Since any assembly is dependent on the presence of every module, destination construct and functioning Type IIS restriction enzyme and DNA ligase, any single omission should block the formation of a complete assembly that would result in a blue phenotype. Indeed, this was seen in all lacI/ lacZ assembly test systems; no blue colonies were obtained if any single component was omitted.
Screening for successful Golden Gate Assembly
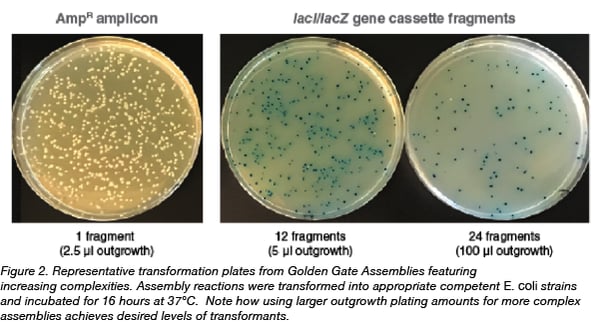
All Golden Gate assemblies feature an inverse proportionality between the complexity of the assembly (number of inserts or modules) and the resulting efficiency of assembly (number of transformants); the greater the number of inserts, the lower the number of transformants. To compensate for this, researchers can plate larger volumes of the outgrowth containing the transformants onto the selection plate. Figure 2 shows representative transformation plates obtained from 1-, 12- and 24-fragment assemblies of the lacI/lacZ cassette, and illustrates how the volume of the 1 ml outgrowth spread on each transformation plate can be manipulated to result in appropriate levels of colony plating densities.
Ligase fidelity and assembly efficiency
Five fragment lacI/lacZ cassette assembly was easily achievable with high levels of transformants and low backgrounds – so much so that there was little range for detectable improvements in the methodology. To further push the test system, we redesigned for 12 and 24 fragments.
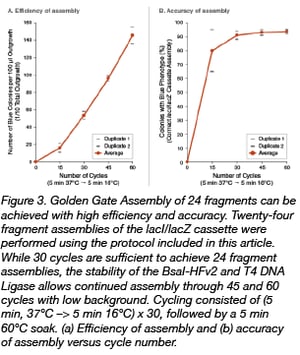 This redesign was guided by both advances in the re-engineering of the original BsaI-HF® Type IIS restriction enzyme and the completion of T4 DNA Ligase fidelity studies conducted by Potapov et al. at NEB (1, 2). While T4 DNA Ligase, the mainstay of most biotechnological cloning efforts for over 50 years, prefers ligation of Watson-Crick base pair substrates, it demonstrates significant activity on some mismatch-containing pairings. During Golden Gate assembly, ligation of mismatched pairs of overhangs can lead to incorrect assemblies, so care must be taken to minimize this possibility. Recently, NEB researchers profiled the comprehensive fidelity of cohesive end ligation by this enzyme for all 3- and 4-base overhang sequences under standard reaction conditions. This data set allows quantitation of sequence-dependent ligation efficiency and identification of mismatch-prone pairings. Using these 4 bp overhang observations, accurate “high-fidelity” junction sets for both the 12- and 24-fragment versions of the lacI/lacZ cassette were designed and synthesized. In conjunction with BsaI-HFv2 (NEB #R3733), re-engineered to provide improved Golden Gate performance, a series of optimization experiments for these more complex assemblies were performed. It was found that high efficiencies and accurate assembly levels were indeed possible, with correct, in-frame assembly proceeding in 99% of 12-fragment assemblies and over 90% for 24-fragment assemblies (Figure 3, Table 1). These improvements (5- to 12-fold increases in the number of transformants while maintaining high levels of correct assemblies) translate to faster assembly workflows that require less screening to confirm the accuracy of assembly. The fidelity data can be applied to derive similar high-fidelity overhangs for any Golden Gate assembly design.
This redesign was guided by both advances in the re-engineering of the original BsaI-HF® Type IIS restriction enzyme and the completion of T4 DNA Ligase fidelity studies conducted by Potapov et al. at NEB (1, 2). While T4 DNA Ligase, the mainstay of most biotechnological cloning efforts for over 50 years, prefers ligation of Watson-Crick base pair substrates, it demonstrates significant activity on some mismatch-containing pairings. During Golden Gate assembly, ligation of mismatched pairs of overhangs can lead to incorrect assemblies, so care must be taken to minimize this possibility. Recently, NEB researchers profiled the comprehensive fidelity of cohesive end ligation by this enzyme for all 3- and 4-base overhang sequences under standard reaction conditions. This data set allows quantitation of sequence-dependent ligation efficiency and identification of mismatch-prone pairings. Using these 4 bp overhang observations, accurate “high-fidelity” junction sets for both the 12- and 24-fragment versions of the lacI/lacZ cassette were designed and synthesized. In conjunction with BsaI-HFv2 (NEB #R3733), re-engineered to provide improved Golden Gate performance, a series of optimization experiments for these more complex assemblies were performed. It was found that high efficiencies and accurate assembly levels were indeed possible, with correct, in-frame assembly proceeding in 99% of 12-fragment assemblies and over 90% for 24-fragment assemblies (Figure 3, Table 1). These improvements (5- to 12-fold increases in the number of transformants while maintaining high levels of correct assemblies) translate to faster assembly workflows that require less screening to confirm the accuracy of assembly. The fidelity data can be applied to derive similar high-fidelity overhangs for any Golden Gate assembly design.
| Number of fragments assembled | Golden Gate Assembly protocol* (Volume of 1mL Outgrowth plated) | Correct assemblies per plate | Fidelity of assembly (percent correct) | Calculated colony totals | ||||||||||||||||||||
| Per 2 μl assembly reaction | Per full assembly reaction** | |||||||||||||||||||||||
| 1 | 5 min., 37 ºC (2.5 μl) | 687 | 100% | 274,200 | 2,742,000 | |||||||||||||||||||
| 1 | 60 min., 37 ºC (2.5 μl) | 1,623 | 100% | 649,200 | 6,492,000 | |||||||||||||||||||
| 12 | (5 min., 37 ºC -> 5 min., 16 ºC) x 30 (5 μl) | 245 | 99.5% | 48,900 | 489,000 | |||||||||||||||||||
| 24 | (5 min., 37 ºC -> 5 min., 16 ºC) x 30 (100 μl) | 78 | 90.7% | 783 | 9,792 | |||||||||||||||||||
Table 1. Yields and fidelities for Golden Gate Assemblies with BsaI-HFv2 and T4 DNA ligase. Efficiency of assemblies per plate using outgrowth volumes described in Figure 3, with calculated yields from entire outgrowth built from 2 µl of the assembly reaction, and from the entire assembly reaction. All assembly reactions had a 5 minute terminal soak after either 37 ºC incubations or cycling.
*Assembly reaction volumes were 20 μl (1, 12 fragments) or 25 μl (24 fragments).
The future of assembly
DNA assembly methods are important tools for many areas of science, and researchers continue to test the limits of DNA assembly approaches with increasingly complex experimental conditions. The ability to construct more complex, multi-fragment assemblies, as shown in this work, will fuel additional efforts to push the technique forward.
Many thanks to our guest bloggers Becky Kucera, M.Sc. and Eric Cantor, Ph.D. from New England Biolabs.
 Becky Kucera is a Principal Development Scientist of Applications & Product Development focusing on DNA cloning and assembly, New England Biolabs, Inc.
Becky Kucera is a Principal Development Scientist of Applications & Product Development focusing on DNA cloning and assembly, New England Biolabs, Inc.

- Eric Cantor is a molecular biologist and a Development Group Leader overseeing a team bringing new products to market, New England Biolabs, Inc.
References
1. Vladimir, Potapov, et al. "Optimization of Golden Gate assembly through application of ligation sequence-dependent fidelity and bias profiling." bioRxiv (2018): 322297.
2. Potapov, Vladimir, et al. "A single-molecule sequencing assay for the comprehensive profiling of T4 DNA ligase fidelity and bias during DNA end-joining." Nucleic acids research (2018). PubMed PMID: 29741723. PubMed Central PMCID: PMC6061786.
Additional resources on the Addgene blog
- Learn the basics of Golden Gate assembly
- Check out NEB's work on ligase efficiency
- Learn about other plasmid cloning techniques
Resources on Addgene.org
- Find basic molecular biology protocols
- Find plasmid cloning protocols
- Check out our molecular biology reference pages
Topics: Plasmid Cloning, Molecular Biology Protocols and Tips, Plasmids







Leave a Comment