This post was contributed by guest blogger Behnam Nabet, a postdoctoral fellow at Dana-Farber Cancer Institute.
Targeted protein degradation
In the Bradner and Gray labs, we synthesize compounds that enable selective removal of proteins-of-interest from the proteome. Rather than inhibiting protein function, these so-called “small molecule degraders” recruit the proteasome to destroy targeted proteins. We previously developed small molecule degraders that achieve selective degradation of endogenous proteins (notably, BRD2/3/4, CDK9, TRIM24, FLT3, BTK, and ALK) by linking small molecules that bind these target proteins to other small molecules that bind an E3 ligase. These bifunctional degraders co-opt E3 ligases such as cereblon (CRBN) or von Hippel-Lindau (VHL) to bring the endogenous degradation machinery into close proximity with the target protein, leading to polyubiquitination of the target protein and proteasomal degradation. Remarkably, small molecule degraders provide distinct advantages over pharmacological inhibitors including rapidly depleting a protein-of-interest, increasing target selectivity, overcoming resistance to inhibitors, and inducing prolonged biological effects.
While targeted degradation of endogenous proteins is an exciting strategy, it is difficult to apply this technique to oncoproteins that lack selective small molecule binders. To overcome this limitation, we created a generalizable tag-based strategy, which we refer to as the degradation tag (dTAG) system, to co-opt the endogenous ubiquitin proteasome machinery to degrade any protein. We are particularly excited about the dTAG system as we can use it to evaluate the effects of rapid and selective degradation of protein targets in time scales that are not possible with traditional genetic approaches. In addition, the dTAG system is versatile and can be used in both cell-based assays and mouse models.
The dTAG system
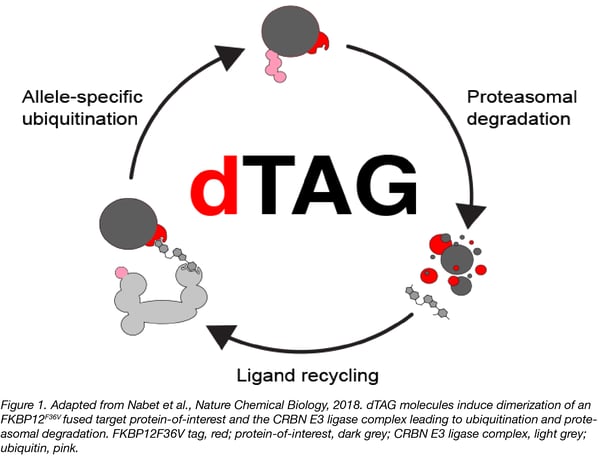
The dTAG system pairs highly selective small molecule FKBP12F36V degraders (dTAG molecule) and expression of FKBP12F36V-tagged proteins. The dTAG molecule induces ternary complex formation between the FKBP12F36V-fusion and the ubiquitin proteasome machinery through binding with cereblon, leading to rapid target degradation (Figure 1).
Plasmids: We generated a series of plasmids that are available from Addgene. These plasmids enable lentiviral expression of N- or C-terminally FKBP12F36V-tagged fusion chimeras or locus-specific knock-in of FKBP12F36V. Detailed protocols describing the use of these plasmids are available here. The Methods section of Nabet et al., Nature Chemical Biology additionally provides detailed information on cloning procedures used to construct the deposited plasmids.
Molecules: We synthesized dTAG molecules by conjugating a CRBN-binding ligand (thalidomide) to an FKBP12F36V binding ligand (ortho-AP1867) via various chemical linkers. Our current lead dTAG molecule is dTAG-13 due its immediate action, potent ligand performance, great depth of degradation, absolute selectivity, broad utility for degrading targets in different cellular compartments, and in vivo utility. dTAG molecules will be commercially available in the near future, and we will provide an update with vendor and catalog numbers. As we are happy to share aliquots of dTAG-13 in the meantime, please address your requests for dTAG molecules and any questions to Behnam Nabet (behnam_nabet@dfci.harvard.edu), Nathanael Gray (nathanael_gray@dfci.harvard.edu), or Jay Bradner (james.bradner@novartis.com).
Considerations when getting the dTAG system up and running in your lab
Confirm the functionality of your fusion chimeras
For each protein-of-interest, it is important to determine which terminus can accommodate the FKBP12F36V tag without disrupting protein function. A literature search can help determine which terminus is favorable for tagging. However, if it is unclear which terminus should be tagged, the dTAG lentiviral plasmids can serve as a rapid means of comparing functionality of N- and C-terminal fusions and degradation in a cell line of interest prior to starting knock-in experiments.
Test dTAG molecule toxicity in new cell lines
Treat parental cells (or cells expressing a control -- for example, an FKBP12F36V-tagged LACZ) with dTAG molecules in your biological assay of choice to ensure that there are no toxicities at the doses used to evaluate degradation of the FKBP12F36V-fusion chimera of interest. In the cell lines we have evaluated to date, we observed little to no toxicity with dTAG-13 at doses as high as 20 µM (see Figures 3 and 5 in Nabet et al., Nature Chemical Biology for examples).
Monitor neo-substrates
Small molecule degraders may recruit CRBN to nonstandard or “neo-substrates.” We advise evaluation of degradation of known cereblon neo-substrates (IKZF1, IKZF3, GSPT1, ZFP91, and CK1a). While expression of these cereblon neo-substrates varies in different contexts (for example, IZKF1 and IZKF3 are cell-type specific transcription factors with important roles in Multiple Myeloma), it is important to ensure lack of degradation in your context. Immunoblotting is recommended, while quantitative proteomics experiments will provide the best confirmation of selective FKBP12F36V fusion chimera degradation. In the cell lines we have evaluated to date, we have not observed off-target neo-substrate degradation with dTAG-13.
Example experiments
See our recent papers noted below for examples of dose-responses and time-courses (Nabet et al., Erb et al., Huang et al., Weintraub et al.), washout experiments (Nabet et al. and Weintraub et al.), chemical and genetic rescue experiments (Nabet et al.), proteomics (Nabet et al., Erb et al., Huang et al.), transcriptional and epigenomic profiling studies (Nabet et al., Erb et al., Weintraub et al.), assessments of fusion chimera functionality (Nabet et al., Erb et al., Huang et al., Weintraub et al.), knock-in (Nabet et al. and Weintraub et al.), and in vivo studies (Nabet et al.).
 Behnam Nabet, PhD is a postdoctoral fellow at the Dana-Farber Cancer Institute. He is a chemical biologist interested in targeted protein degradation, and lead author on the recent paper describing the dTAG technology. Follow him on twitter @behnamnabet.
Behnam Nabet, PhD is a postdoctoral fellow at the Dana-Farber Cancer Institute. He is a chemical biologist interested in targeted protein degradation, and lead author on the recent paper describing the dTAG technology. Follow him on twitter @behnamnabet.
References
Nabet et al. The dTAG system for immediate and target-specific protein degradation.. Nature Chemical Biology. May 2018. 14(5): 431-441. PubMed PMID: 29581585.
- This reference contains the full description of the dTAG system. The dTAG system was employed to evaluate degradation of multiple FKBP12F36V-tagged proteins using our lentiviral expression platform, to assess selective degradation of FKBP12F36V-BRD4 using CRISPR/Cas9-mediated knock-in, and to evaluate in vivo degradation.
- Featured molecules: dTAG-7 and dTAG-13
Erb et al. Transcription control by the ENL YEATS domain in acute leukemia. Nature. March 9 2017. 543(7644): 270-274. PubMed PMID: 28241139. PubMed Central PMCID: PMC5497220.
- The dTAG system was employed to evaluate degradation of ENL. Cell lines were engineered to express ENL-FKBP12F36V and endogenous ENL was inactivated using CRISPR/Cas9 gene-editing.
- Featured molecules: dTAG-7 and dTAG-13
Huang et al. MELK is not necessary for the proliferation of basal-like breast cancer cells. eLife. September 2017. 6: e26693. PubMed PMID: 28926338. PubMed Central PMCID: PMC5605198.
- The dTAG system was employed to evaluate degradation of MELK. Cell lines were engineered to express FKBP12F36V-MELK and endogenous MELK was inactivated using CRISPR/Cas9 gene-editing.
- Featured molecules: dTAG-7, dTAG-13, and dTAG-47
Weintraub et al. YY1 Is a Structural Regulator of Enhancer-Promoter Loops. Cell. December 14 2017. 171(7):1573-1588.e28 PubMed PMID: 29224777. PubMed Central PMCID: PMC5785279.
- The dTAG system was employed to evaluate degradation of YY1. Cell lines were engineered to express YY1-FKBP12F36V using CRISPR/Cas9-mediated knock-in.
- Featured molecule: dTAG-47
Additional Resources on the Addgene Blog
- Easi-CRISPR: Generating Knock-In and Conditional Mouse Models
- CRISPR 101: Cas9 Nickase Design and Homology Directed Repair
- Plasmids for endogenous gene tagging in human cells
Resources on Addgene.org
- Browse all CRISPR plasmids
- CRISPR plasmids for protein tagging
- Find validated gRNAs
Topics: Other Plasmid Tools, Plasmids






Leave a Comment