 80,000+ reagents and 700+ educational resources — including blog posts, protocols, guides, and collections — is a lot of material to sift through. Fortunately, we’ve just made it easier for you to find what you need quickly.
80,000+ reagents and 700+ educational resources — including blog posts, protocols, guides, and collections — is a lot of material to sift through. Fortunately, we’ve just made it easier for you to find what you need quickly.
Through the years, we have perfected processes to curate the data our depositors share and enrich it with sequencing data; as a result, the materials in our catalog are very well-characterized. During this time, we’ve also created a variety of educational resources in an effort to make scientific knowledge broadly available.
The ability of our users to discover all this information, however, did not improve at the same pace as the quantity and quality of the information. As a result, resources on the website were hard to find and many remained undiscovered.
To remedy this, throughout the year we rolled out multiple changes to our website, including the recent homepage and search tool updates. Now, let’s take a look at the least visible part behind these changes: the backend technology that powers the new website search.
Tackling findability from multiple angles
We assembled a cross-team team, with members from the Customer Service, Data Curation, Outreach, Product, Scientist, and Software teams. We knew we had to improve our website search function. We also quickly realized that we needed to improve more than that: the general “findability” of the information on the site. Different users have different strategies for finding what they need, and we wanted to support all of them — searching, browsing, drilling down, and zooming out — with an intuitive user interface.
The team approached the issue from multiple angles: the overall user experience of our website, the different website navigation paths, the organization and completeness of our information, and the accuracy and relevance of our search results. The footer and improved menus we launched earlier in the year, as well as our redesigned homepage and our new search tool, are the results of this multidisciplinary effort and cross-team collaboration (thanks to all Addgenies involved).
The new homepage and navigational elements help users who prefer to browse and discover items in our catalog or educational resources through our main dropdown menu. For users who prefer to use search, our updated search engine is now more powerful, intuitive, and searches across all of our content, whether educational resources, blog posts or reagent items.
The technology behind Addgene’s new search
Like millions of other websites, we built our new search engine atop Elasticsearch, a powerful open-source search engine that can scale to search millions of web resources and handle millions of queries. In an iterative process driven by our user experience experts, our software team tailored Elasticsearch to support the different usage patterns the larger team identified.
The basic function of a search engine is to find the website resources that best match a user’s query, and return the matching resources in an order that makes sense to the user. The devil is in the details of course: what does “best” mean? and what “makes sense” to a user? Moreover, there are many ways a user can input their query — by directly typing terms in a search bar, by ticking some boxes, by selecting a suggestion — how should these be supported and combined?
Luckily, we had the advantage of a very specialized domain of information so we were able to fine tune our approach much more than if we were dealing with general purpose information. Below are a few of the strategies we used in our software design. In each case, we leveraged the team’s knowledge of the scientific domain in general and of our data in particular and our deep understanding of Addgene’s customer’s needs.
Categories
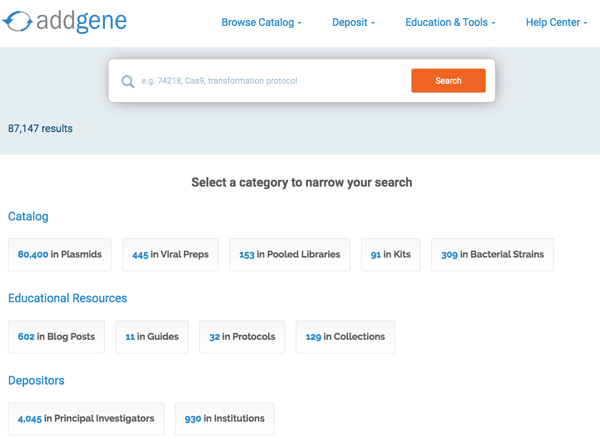
We created a categorization of our information in order to more granularly customize the search engine, and a scoping mechanism that allows category-specific, narrow searches. We defined three top level categories — catalog, educational resources, and depositors — each with subcategories such as plasmids and viral preps for catalog, blog posts and protocols for educational resources, and labs and institutions for depositors (see below). Even if the user experience is similar, the way the software searches through our catalog of materials and services is quite different from the way it searches through our educational resources.
Filters
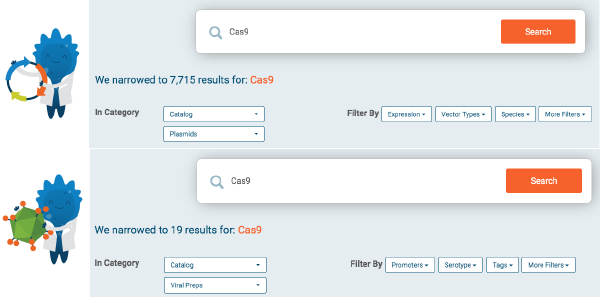
We identified the most useful kinds of information per resource category to expose as filters and designed the search engine to return only the filters that are relevant to a specific query and category. For example, a search scoped to plasmids shows filters for expression and vector type, while a search scoped to viral preps shows filters for promoter and serotype, among others. We also created a normalization mechanism to “collapse” variations and synonyms of the same term that appear in our data into a single filter value. For example, there is only one viral prep filter for promoter “Syn”, that, when selected, will match variations present in the data such as synapsin, hsyn synapsin-1, human synapsin, hSynap.
Matching
We capture multiple variations of the same information in our search indices in order to support different kinds of matches: exact words, partial words, phrases, partial phrases, or numerical identifiers. We also combine the different terms in a query in a way that allows users to narrow down search results by adding more terms.
Weighting
We determined the relative importance of different bits of information for a given resource, and designed the search engine to weight search results differently according to where a match occurred, and whether it was a full or partial match. For example, matches of a name in the depositor field will rank higher than matches of the same name in the publication field.
Suggestions
We provide as-you-type suggestions in different ways. Keyword suggestions inform the user that the terms they are typing have matches in a given category, and serve as a shortcut to a search scoped to that category. Catalog ID suggestions provide shortcuts to specific catalog items as well as related items (for example, a kit that contains a plasmid with the catalog ID). Depositor suggestions provide shortcuts to matching PI labs and institutions.
There are many more technical details that make our new search engine unique. Most importantly, with this new implementation, we laid the foundations for many improvements to come. Ultimately, we would like our search engine to be so specialized that it appears to “understand” biology.
We hope you will give our new search engine a try to find reagents and educational material. Let us know what works for you and what doesn’t.
Try out the new Addgene search engine!
Additional resources from the Addgene blog
- Learn about other features on Addgene's website
- Find a step-by-step guide to using Addgene's new search
- Read to learn more about why we redesigned our homepage and mascot
Topics: Addgene News, Using Addgene's Website





Leave a Comment