Your new antibody has arrived, hooray! Before you jump straight into the exciting new experiments that you’ve been planning, it is a good idea to pause and make sure that your antibody has been appropriately validated so that you can have confidence in your results. But what is appropriately validated? This post will go over some of the general considerations you should review before using an antibody.
Antibodies and Reproducibility
First, let’s review why validation is so important. Antibodies are a critical component of the researcher toolkit due to their ability to bind to molecular targets with high affinity and specificity. However, there are countless variables that impact an antibody’s affinity and specificity for a given target and it is all too common for antibodies to bind to off-target molecules or even to not bind to their target molecule at all, leading to inconclusive results, erroneous conclusions, and wasted time. In fact, many have called out antibodies as major culprits in the reproducibility crisis of biomedical research (Baker, 2015; Bradbury & Plükthorn, 2015).
To combat this, a number of groups have come together over the past few years to propose standards and recommendations for improving antibody use and reliability (Roncador, et al., 2015; Taussig, et al., 2018; Uhlen, et al., 2016). The major takeaway from these recommendations is: validate, validate, validate. This doesn’t necessarily mean that you yourself have to perform extensive validation experiments for every antibody that you receive; citing existing validation data can be acceptable. However, it is your responsibility to determine what counts as adequate validation. If you are not convinced by the existing data, you may need to roll up your sleeves and perform some extra validation experiments yourself.
Goal of Antibody Validation
As you assess existing data or plan your own validation experiments, it’s important to know what you are aiming to achieve. In general, these are three core goals of antibody validation:
- Confirm reactivity with the target antigen - Recognition of an immunogen does not guarantee recognition of the target antigen. You want to make sure that your antibody binds to what you expect it to bind to.
- Show specificity for the target antigen - Just because an antibody can bind to its target, does not mean that is all it will bind to - particularly when you start using complex samples such as whole cell lysate or tissue sections. It is important to confirm that your antibody is not binding to proteins outside of your target protein.
- Show suitability for an intended application - Remember, just because your antibody performs as expected in one application does not mean that it will perform as expected in another. You’ll need to validate that points 1 and 2 hold true under your specific experimental conditions.
Strategies for Antibody Validation
There are a number of resources available that provide in-depth discussions on strategies for antibody validation in different applications. Here, I want to highlight the general suggestions given by the International Working Group on Antibody Validation (Uhlen, et al., 2016). This group proposed 5 Pillars of antibody validation which represent five experimental strategies researchers can use to show that their antibody performs as expected. One of the benefits of these pillars is that they are intentionally generalizable and not specific to certain applications or antibody targets. That said, not every strategy will be suitable for every antibody or every application. The goal is to show that your antibody has been validated using at least one of these strategies. Choosing which strategy to use will depend on your intended application and available resources.

|
| Figure 1: Table summarizing the IWGAV five proposed antibody validation pillars. Created with BioRender.com. |
Orthogonal Strategies
This approach compares target expression of antibody-independent approaches to your antibody-dependent approach. Antibody-independent methods may include analysis of transcriptomic or proteomic data from several different tissues or cell types. If your antibody indeed binds to your target then you would expect to see good correlation between the levels of target detected in both methods.
These approaches have relatively low sensitivity, which can be problematic if you are trying to show that your antibody can distinguish between two very similar proteins. Additionally, they require variable expression of your target across samples - it is hard to distinguish between specific, invariable expression and non-specific, background signal, so you’ll need a range of expression strengths to be confident in your data. This is a good option for applications like immunohistochemistry on human tissues, when (1) other strategies may not be feasible and (2) large-scale proteomic and transcriptomic datasets already exist.
Example: You want to use an antibody to examine the levels of your favorite protein in human tissues. You know that there is a wealth of transcriptomic data from this sample type, so you use your bioinformatics skills (or your friendly neighborhood bioinformatician) to look at RNA expression of your gene of interest across a set of human tissues.
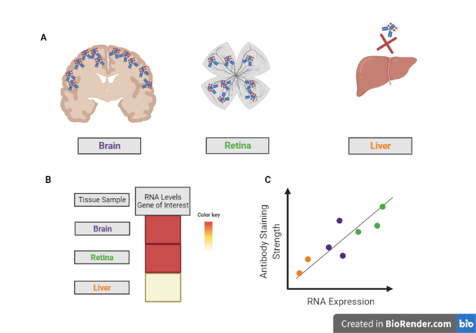
From this data, you see that there is high RNA expression in the retina and in parts of the brain, but almost none in the liver (A). In general, RNA levels tend to correspond to protein levels, so you perform immunohistochemistry with your antibody on these tissues (B). If that antibody works as expected, you will see a strong antibody signal in the retinal sample, but little to no signal in the liver sample, which you do (C), so your antibody is validated!
Genetic Strategies
This approach uses a gene knock-down or knock-out approach to show that an antibody binds specifically to the expected protein. Common approaches include using RNAi to knock-down expression of your target or CRISPR/Cas9 to create mutations that essentially eliminate expression of the target. Removing (or reducing) the protein of interest in your sample, should lead to reduced or eliminated antibody signal. If you see a similar antibody signal between your control and your knock-out sample, you should be concerned that your antibody is binding to something unexpected. This approach has high specificity, but you need to be aware of any possible alternative translation start sites or alternative splicing that could lead to some form of the target protein to be present and recognizable by your antibody despite the manipulation. Additionally, not all sample types are amenable to genetic manipulation, nor can all genes be easily knocked down or out.
Example: You just received an antibody that you want to use to look at the subcellular localization of a protein of interest. Conveniently, you found a cell line in which the gene for your protein has been mutated using CRISPR/Cas9. You check the mutation and it causes a large deletion that removes the entire exon that codes for the epitope recognized by your antibody.
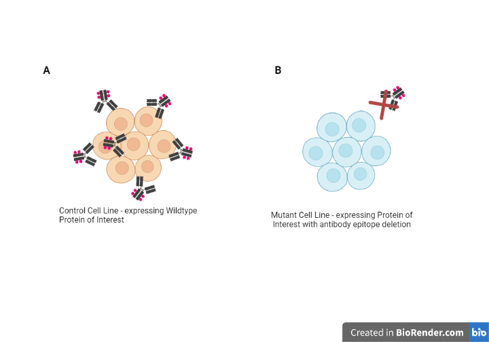
You perform immunocytochemistry on these mutant cells as well as a non-mutant line and see a great antibody signal in the non-mutant line but no signal in the mutant line!
Recombinant Strategies
This approach is essentially the opposite of the Genetic strategy and looks for increased antibody signal in samples that have had the target protein expression experimentally increased. The simplest version would be to use cells that do not express your protein of interest endogenously and transfect them with a plasmid encoding your protein of interest. Then you should expect to see evidence of antibody binding in the transfected cells but not in control cells. But (there is always a ‘but’), you should be aware that overexpression of a protein can also influence antibody binding by altering the relative concentration of potential antibody binders and so this approach may not be best if you are ultimately planning to use your antibody to detect endogenous levels of the target in other cells or tissues.
Example: You want to show that your antibody binds to the expected target protein in a Western blot.
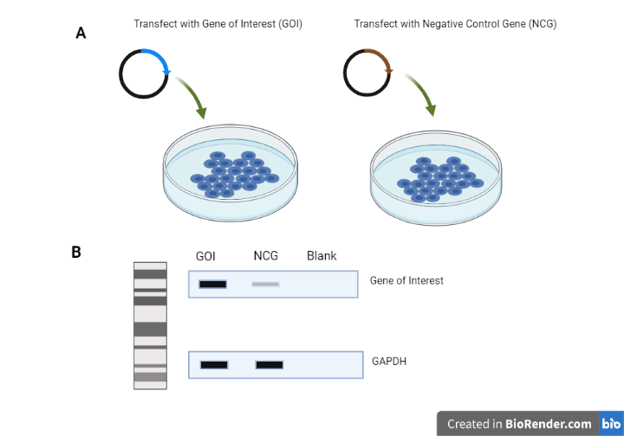
You acquire a plasmid encoding your protein of interest, known as Gene of Interest (GOI), and transfect it into your favorite cell line. In parallel, you transfect cells with another plasmid not encoding this protein, which we will call Negative Control Gene (NCG) (A). Then you lyse the cells and run a Western Blot with your antibody (B). Your results show a lovely, strong band from the cells transfected with your protein of interest and show just the faintest band from the cells transfected with the other plasmid. Your antibody works!
Multiple (Independent) Antibodies Strategies
These strategies use multiple, unique antibodies against the same protein, each targeting a different epitope, and look for correlation between the results. Using antibodies against different epitopes on the same target reduces the likelihood that both antibodies would bind to the same off-target molecules, which gives you more confidence when both antibodies exhibit similar results. Unfortunately, it can be difficult to find epitope information for some antibodies, which makes it hard to ensure that you have unique antibodies. An additional caveat is that different antibodies may or may not perform well under the same conditions.
Example: You want to sort cells based on the expression of a new receptor that is of interest to you. You found a couple of antibodies that are predicted to bind different epitopes of this protein.
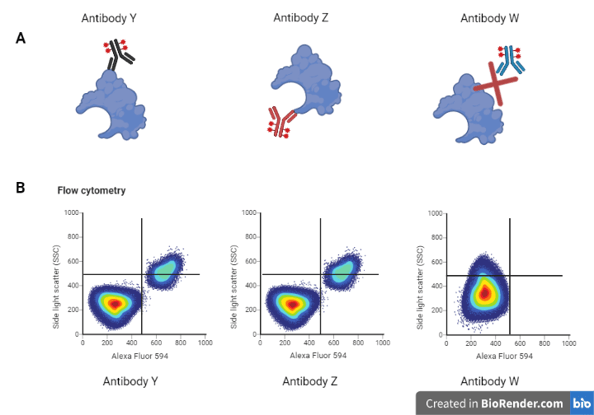
You perform flow cytometry using both antibodies, as well as a third unrelated antibody as a negative control (A). The two antibodies to your protein of interest give you very similar results, while the third looks entirely different (B), giving you confidence that your new antibodies will work for your experiments.
Capture MS Strategies
This approach uses immunoprecipitation to capture proteins from a sample using the antibody and then uses mass spectrometry to identify the captured proteins. If the antibody is specific, then you expect that the bulk of the peptides identified to be from the target protein. This approach is ideal for validating antibodies for immunocapture applications and is amenable to high-throughput analyses, however, it requires access to a mass spec facility and the ability to analyze the resulting data, so it may not be the most accessible option for many laboratories. Additionally, it can be difficult to distinguish between proteins bound by the antibody vs proteins bound to the target protein, so it’s important to understand if protein:protein interactions could be a factor in your analysis.
Example: You are planning an RNA-immunoprecipitation experiment and want to validate that your antibody is specifically capturing your RNA-binding protein of interest.
So you perform an immunoprecipitation with your antibody and send the captured proteins off for mass spec analysis (above). When the results come back it shows that the majority of the protein in the sample was identified as your target protein, allowing you to proceed with confidence.
Share your results
Ultimately, like all of science, validation experiments don’t mean anything unless you share the results with others. If you publish data generated using your antibody, it’s best scientific practice to provide or reference the data that convinced you of your antibody’s trustworthiness. You may also consider sharing your validation attempts by other means, such as through antibody databases like Antibodypedia or through your antibody provider(s) if they accept and share user data on the antibody webpages. These outlets can be particularly useful for sharing negative data. Negative data often goes overlooked, but knowing that an antibody likely won’t work in a given application is just as helpful as knowing when it will. And remember, antibodies are like puppies. Just because an antibody doesn’t work in your application doesn’t mean it’s a bad antibody but more that it’s misunderstood and it could work great in another context! So the next time you test an antibody (perhaps one from Addgene!), consider sharing your results so that others in the community can benefit and we can all help improve antibody reproducibility!
References and Resources
Additional resources on the Addgene blog:
- Still trying to choose an antibody? Check out our Selecting an Antibody blog post!
- Learn more about Antibodies and their many applications in our Antibodies 101 blog series
- New to antibodies? Read our Antibodies 101: Introduction to Antibodies blog post!
Resources on Addgene.org
- Find plasmids encoding antibodies at Addgene
- Stay tuned for ready-to-use antibodies from Addgene
- Designing a recombinant validation experiment? Find plasmids here!
References
- Baker M (2015) Reproducibility crisis: Blame it on the antibodies. Nature 521:274–276. https://doi.org/10.1038/521274a
- Bradbury A, Plückthun A (2015) Reproducibility: Standardize antibodies used in research. Nature 518:27–9. https://doi.org/10.1038/518027a
- Roncador G, Engel P, Maestre L, Anderson AP, Cordell JL, Cragg MS, Šerbec VČ, Jones M, Lisnic VJ, Kremer L, Li D, Koch-Nolte F, Pascual N, Rodríguez-Barbosa J-I, Torensma R, Turley H, Pulford K, Banham AH (2015) The European antibody network’s practical guide to finding and validating suitable antibodies for research. Mabs 8:27–36. https://doi.org/10.1080/19420862.2015.1100787
- Taussig MJ, Fonseca C, Trimmer JS (2018) Antibody validation: A view from the mountains. New Biotechnol 45:1–8. https://doi.org/10.1016/j.nbt.2018.08.002
- Uhlen M, Bandrowski A, Carr S, Edwards A, Ellenberg J, Lundberg E, Rimm DL, Rodriguez H, Hiltke T, Snyder M, Yamamoto T (2016) A proposal for validation of antibodies. Nat Methods 13:823–827. https://doi.org/10.1038/nmeth.3995
Topics: Antibodies






Leave a Comment