I love Addgene's new Antibody Guide and not just because I helped create it. But while antibodies, as the OG (as the kids say) affinity reagents deserved to be celebrated, there are many other amazing affinity reagents we just didn't have the space to give proper credit to. Thankfully, as Addgene's resident blog editor, I could take the needed steps to continue where the guide left off - namely, a blog post devoted to the other affinity reagents.
Affinity Reagents
Affinity reagents are molecular tools that have the ability to specifically recognize and bind proteins. They’re used in a number of research and clinical applications. Most of the time, when someone speaks about an affinity reagent, they’re talking about a standard antibody: An ~150kD protein with both light and heavy chains made by immune systems. But here, we're not interested in antibodies. Instead, we're going to focus on what I'm going to call "alternative affinity reagents" - everything from camelid antibodies to antibody fragments to tools completely unrelated to antibodies.
Why use an alternative affinity reagent?
The main advantage of alternate affinity reagents is their size. Antibodies are large enough that their bulk can be a barrier at times. Smaller, slimmer affinity reagents can have deeper penetration into a tissue, useful for applications like IHC, or be able to access an epitope folded into a space that would size-exclude an antibody. In imaging applications, smaller affinity reagents means higher resolution. Many of the below are also small enough to be expressed in bacterial and other cell systems.
Since the majority of these alternative reagents are created in the lab, instead of generated in vivo, they have higher reproducibility and consistency in the final product. Because the constant (non-epitope binding) region is the first thing to go when downsizing an affinity reagent, most of the alternative reagents have reduced immunogenicity. Effector functions are eliminated, as is antibody recognition, since antibody-specific antibodies bind to and recognize the constant region. Nonspecific binding, caused cell receptors for the constant region, is also reduced.
Disadvantages
Every rose has its thorns and every affinity reagent has its downsides. These reduced-in-size options often have reduced binding strength, half-lives, and/or specificity as well. They’re more difficult to source, may not be readily available for a large number of targets, and can require specific conjugates. And, of course, once you have them, you’ll need to either re-develop or re-optimize your assay for an entirely new affinity reagent.
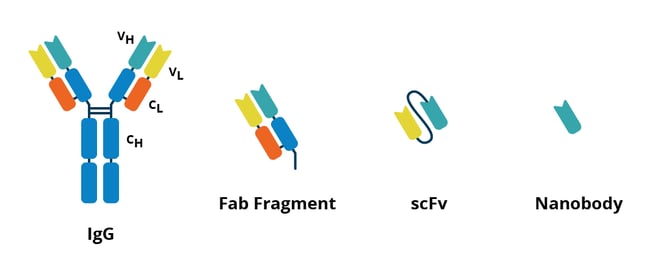 |
| Fig. 1: A comparison of an antibody (left) to several alternative affinity reagents. |
Types of Affinity Reagents
(Listed in no particular order.)
Fab fragments are created through enzymatic cleavage of the variable region (the antibody region that binds to the antigen) from existing antibodies. Each cleaved antibody will produce two ~50kD Fab fragments, one for each arm of the antibody’s ‘Y’. They are often used in clinical applications. Fab fragments can be created in lab in situations where a suitable antibody has been identified but a smaller or less immunogenic version is needed.
Single-chain fragment variables (scFvs) are ~27kD engineered fusion proteins similar to a Fab fragment but created through engineering instead of cleavage. They are comprised of the variable heavy and light chain regions, connected by a short linker peptide. The lack of constant region makes them less immunogenic, but they have lower affinity and longevity compared to antibodies, along with a higher likelihood of aggregation (Bates, 2019). They are small enough to be produced in bacteria and are generally cheaper to make than antibodies.
Minibodies are created by binding together two scFvs with a CH3 linker, resulting in an ~50kD protein. Minibodies are bispecific, meaning that each scFv can target a different antigen; binding of each can be either simultaneous or independent of the other (Shahied, 2004). They are often used for clinical applications, including imaging.
Diabodies are ~60 kDa proteins formed by two Fab fragments bound by short peptide linkers. They are bispecific, with a lot of flexibility in the orientation of the two fragments. However, rigidity can be introduced through mutation, allowing diabodies to be used for assembling protein nanostructures. Diabodies readily co-crystalize, a major advantage over antibodies (Kwon, 2019).
Single-domain antibodies (sdAbs), sold under the commercial name nanobodies, are derived from a camelid antibody lacking a light chain, called a heavy chain antibody (Muyldermans S., 2013). They have the unique distinction of being the only affinity reagent on this list derived from nature. Single-domain antibodies are extremely small at 12-15 kD but can have weak signals due to their monovalent nature. They are stable up to 80°C and can refold correctly after denaturing.
Designed ankyrin repeat proteins, or DARPins, are genetically engineered small affinity proteins, around 14-15 kDs in size. These are antibody mimics and are not closely related to antibodies. Instead, they are derived from a class of binding proteins known as ankyrin repeat proteins. DARPins typically consist of 3-4 repeats, have high specificity and binding affinity, can be produced in E. coli, and are both soluble and easily engineered.
Monobodies are ~10kD synthetic binding proteins which use the Fibronectin type III domain as their scaffolding instead of the antibody constant region. They are generated using combinatorial libraries, and high specificity has been reported for a number of targets. Due to their size and lack of disulfide bonds, monobodies can be easily expressed in transfected eukaryotic cells, allowing for them to be used as intracellular inhibitors.
Aptamers are not proteins at all. These high-affinity RNA molecules can bind to a wide variety of targets, including proteins, peptides, amino acids, drugs, metal ions, and cells. They can be used either with or as fluorophores in a wide variety of colors.
To antibody or not to antibody?
That is the question… or is it? This post isn’t meant to convert anyone to downsize their affinity reagents, trendy as that may sound. Validated antibodies are still an excellent option for many applications, and frankly, if ain't broke, don't fix it!
That being said, it's important to know all your options when planning and running experiments. You never know what problems will arise in your research or what data you may need to collect. Understanding available alternatives to antibodies can allow you to be flexible in your approach to affinity reagent applications. Whatever affinity reagents you use, one thing is clear: it’s an exciting time to be a scientist looking for that perfect fit!
References and Resources
Additional resources on the Addgene blog
Plasmids 101: Aptamer Fluorophores
Antibodies 101: scFVs
Antibodies 101: Introduction to Antibodies
Additional resources on addgene.org
Antibody Guide
Recombinant antibodies available from Addgene
Browse our recombinant antibody plasmid collection
References
Grumezescu, A. M. (2018). Drug targeting and stimuli sensitive drug delivery systems. Elsevier Science & Technology Books.
Bates A, Power CA. David vs. Goliath: The Structure, Function, and Clinical Prospects of Antibody Fragments. Antibodies (Basel). 2019 Apr 9;8(2):28. doi: 10.3390/antib8020028. PMID: 31544834; PMCID: PMC6640713.
Shahied, L., et. al. (2004). Bispecific Minibodies Targeting HER2/neu and CD16 Exhibit Improved Tumor Lysis When Placed in a Divalent Tumor Antigen Binding Format. Journal of Biological Chemistry. https://doi.org/10.1074/jbc.M407888200
Kwon, N., Kim, Y., Lee, J. (2019). Structural diversity and flexibility of diabodies. Methods. 10.1016/j.ymeth.2018.09.005
Muyldermans S. (2013). Nanobodies: natural single-domain antibodies. Annu Rev Biochem. doi: 10.1146/annurev-biochem-063011-092449.
Topics: Antibodies






Leave a Comment