Adeno-associated viral (AAV) vectors are the most frequently used gene-transfer tools in the study of the brain and spinal cord, which together are known as the central nervous system (CNS). AAVs are popular tools because: 1) their genomes are easy to manipulate, 2) they have long-term expression; and 3) they have limited toxicity. However, a key challenge of using AAVs for neuroscience research is the lack of a method for genetically manipulating neurons throughout the whole brain. Neurons of the peripheral nervous system (PNS), which connect the heart, lung, gut, and other organs to the CNS, are also an important target for gene delivery, especially for the study of pain. While many new capsids (i.e. the part of the virus that determines tropism) have been developed that increase transduction efficiency, none allow for simple and efficient transduction of both the CNS and PNS.That is until the Gradinaru Lab at Caltech stepped up to the challenge.
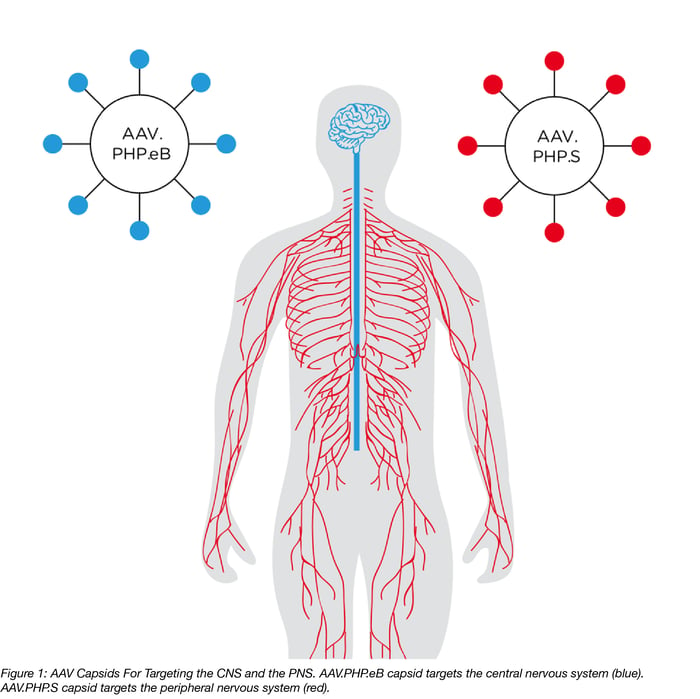
Their Solution: Use the CREATE method to engineer two new IV injectable AAV capsids, AAV.PHP.eB and AAV.PHP.S, that target the CNS or the PNS, respectively.
Read on to learn more about these two new AAV capsids and how they can be useful tools for studying the CNS and PNS!
What is the CREATE method?
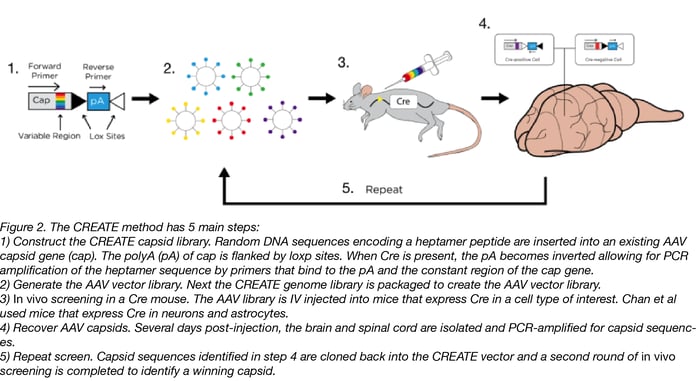
CREATE stands for Cre Recombinase-based AAV Targeted Evolution. In this technique, a pool of mutant AAV capsid (cap) expressing plasmids is generated. Diversity is introduced into the cap plasmid library by inserting short DNA sequences that contain all possible combinations into a region of the cap gene. In this special CREATE cap plasmid, loxP sites flank the downstream polyA sequence. When viruses produced from this plasmid pool are injected into an animal with cell-type-specific Cre, the polyA sequence is inverted only in the Cre-expressing tissues. This inversion creates a template that’s suitable for PCR amplification using pre-designed primers.
By using this library to infect an animal expressing Cre in a cell type of interest, a scientist can isolate DNA from the infected animal and use the pre-designed primers to determine which version(s) of the capsid gene successfully infected the cell type of interest. Repeat rounds of infection and screening by PCR enrich for capsid variants with increased ability to infect the cell type of interest, which, in the case of Chan et al, was neurons and astrocytes.
Why screen for IV deliverable AAVs?
Typically AAVs are directly injected into the brain, but this tends to limit expression to near the site of the injection. IV injection into the bloodstream allows for dispersion of the virus but it has to be able to cross the blood-brain barrier to reach the brain. Delivering the CREATE AAV library via IV screens for capsids that are able to cross this barrier - only those that cross the barrier can be identified by PCR later. The PNS presents a different problem: it’s spread throughout the body and is often difficult to surgically access. Therefore an IV injection is an ideal way to deliver AAV to the PNS. It’s a happy accident that AAV.PHP.S targets the PNS since it was actually engineered to target the CNS.
How well do these AAVs transduce the CNS and PNS?
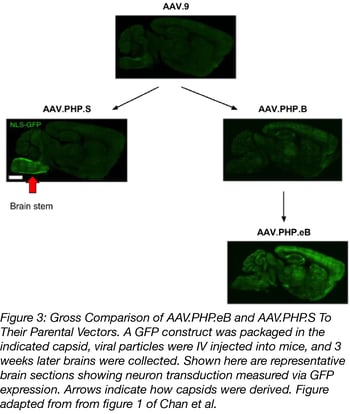
Both plasmids infect the PNS or the CNS much better than the AAV vectors they were derived from. Additionally, AAV.PHP.eB was used at a lower dose than it’s parent AAV.PHP.B: 1x1011 vs 1x1012 viral genomes (vg) per mouse. Figure 3 shows a gross comparison of AAV.PHP.eB and AAV.PHP.S to their parental vectors. To quantify this comparison, capsids were packed with a GFP transgene and GFP intensity and transduction efficiency of neurons were measured. On average, AAV.PHP.eB and AAV.PHP.S had 1.5- to 2-fold higher GFP intensity compared to the strains they were derived from. They also transduced a higher number of neurons overall (GFP+ cells that also stained for a neuronal marker). Check out tables 1 and 2 below for a summary of the quantified transduction efficiency of AAV-PHP.eB and AAV-PHP.S versus their parental AAV vectors.
What can these AAV’s be used for?
1. Multi-color labeling of neurons
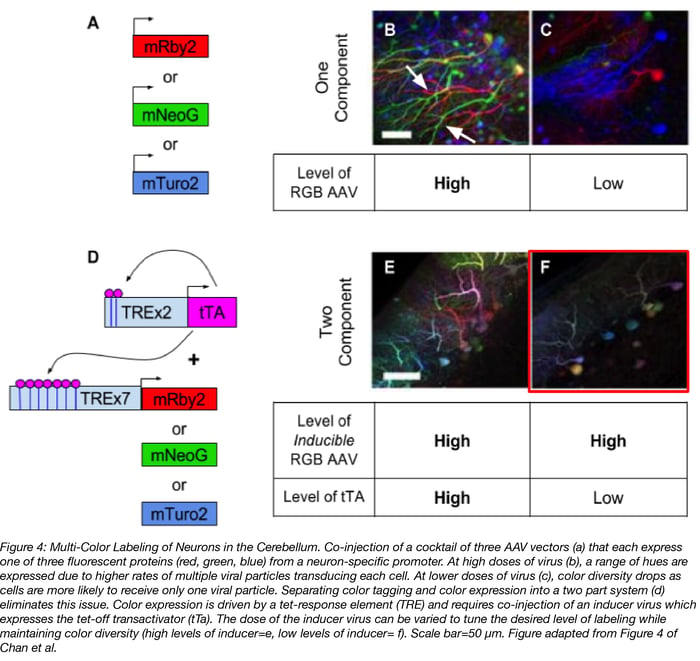
Multi-color labeling of neurons is for more than making pretty pictures (see figure 4). It also enables single-cell anatomical study and neuronal tracing, aka determining where a neuron’s axons and dendrites extend to. The keys to this method are 1) high levels of color diversity so that cells are labeled with their own unique color; and 2) low levels of labeling so that a cell’s projections are easy to trace and not lost in a jumble of fluorescent neurons. A simple but flawed approach to multi-color labeling is to inject a mix of AAV vectors that constitutively express a red, green, or blue (RGB) fluorescent protein (4a-c). When high doses of this viral cocktail are injected, by chance lots of individual cells will be transduced with multiple viral particles and labeled a variety of colors (figure 4b). But notice in figure 4b how difficult it is to trace individual cell’s axons, particularly the two green neurons near the arrows. When lower doses of virus are used in order to decrease the number of neurons that get labeled, neurons are more likely to be transduced by only one RGB AAV and red, green, or blue are the predominate colors expressed (figure 4c).
| AAV Vector | Percent of Neurons Transduced | ||
| Cortex |
Striatum |
Cerebellum | |
| AAV.PHP.B (parental) | 49 +/- 7% | 36 +/- 8% | 37 +/- 10% |
| AAV.PHP.eB | 69 +/- 4% | 55 +/- 7% | 76 +/- 5% |
Table 1: Transduction efficiency of AAV-PHP.B versus AAV-PHP.eB in the CNS. A GFP transgene was packaged into either an AAV-PHP.B or an AAV.PHP.eB capsid. Both viruses were IV injected at a dose of 1x1011 vg per mouse and the level of transduction, measured via GFP expression, was determined 3 weeks later. Table was derived from figure 2e in Chan et al.
| AAV Vector | Percent of Neurons Transduced |
| Dorsal Root Ganglion (DRG) | |
| AAV.9 (parental) | 46 +/- 0.7% |
| AAV.PHP.S | 82 +/- 2% |
Table 2: Transduction efficiency of AAV.9 versus AAV.PHP.S in the PNS. A GFP transgene was packaged into either an AAV.9 or an AAV.PHP.S capsid. Both viruses were IV injected at a dose of 1x1012 vg per mouse and the level of transduction, measured via GFP expression, was determined 3 weeks later. Table was derived from figure 3b in Chan et al.
To achieve the full diversity of colors AND sparse labeling of neurons , Chan et al separated color labeling and color expression into a two part system (figure 4d):
Part 1: A cocktail of three AAV vectors expressing RGB proteins under the control of an inducer, i.e. tet-off transactivator (tTA), is injected at a high dose to maximize the number of cells that are labeled.
Part 2: An AAV expressing the inducer (i.e. tet-off transactivator) is co-injected. The dose of this AAV can be adjusted to achieve sparse levels of labeling, without loss of color diversity.
With this two part system, color diversity is maintained at both high (figure 4e) and low doses (figure 4f) of the tTA inducer virus, but the desired sparsity of labeling is achieved with lose doses of inducer virus. Additionally, this system is less cumbersome than brainbow, another method for multi-color labeling of neurons, because it doesn’t require the generation of a transgenic mouse.
2. Cell-type-specific restricted transgene expression in the CNS and PNS
Previously to broadly introduce a transgene into a specific cell type in the brain, you needed to make a transgenic mouse. Combining AAv.PHP.eB’s or AAV.PHP.S’s great neuronal transduction efficiency, as shown in tables 1 and 2, with a cell-type specific promoter or enhancer now allows for cell-type-restricted expression in both the CNS (figure 5 in Chan et al) and the PNS (supplemental figure 8 of Chan et al). These new AAVs can serve as tools for basic science research (neuronal tracing and morphologic studies), as well as translational applications (mechanistic studies and gene therapy treatments for neurological diseases).
The Gradinaru Lab has a lot going on. Here are the key takeaways:
- Two new AAV capsids (figure 1) were engineered with the CREATE method (figure 2) to allow for efficient IV delivery of virus to the CNS or PNS (figure32, tables 1 & 2).
- These capsids are useful tools for neuronal tracing and morphology studies when used as part of a two part fluorescent tag-then-express system that allows for controlled induction of multi-color labeling of neurons (figure 4).
- These capsids can serve as powerful cell-type-restricted gene vectors for neurons, no transgenic mouse required (see figure 5 and supplemental figure 8 in Chan et al).
References
1. Chan, KY, Jang MJ, Yoo BB, et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nature Neuroscience. 2017; 20(8):1172–1179. PubMed PMID: 28671695.
2. Deverman BE, Pravdo PL, Simpson BP, et al. Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain. Nature biotechnology. 2016;34(2):204-209. PubMed PMID: 26829320. PubMed Central PMCID: 5088052.
3. Lentz TB, Gray SJ, Samulski RJ. Viral Vectors for Gene Delivery to the Central Nervous System. Neurobiology of disease. 2012;48(2):179-188. PubMed PMID: 22001604. PubMed Central PMCID: 3293995.
Additional Resources on the Addgene Blog
- Important considerations when using AAVs
- Learn more about using AAV as a tool for gene expression in mammals.
- Subscribe to Viral Vector Blog Posts
Additional Resources on Addgene.org
- Search for AAV Vectors to use with your research.
- Visit Our AAV Guide Page for info on AAV components, common uses of AAV, and much more!
- Find ready-to-use AAV from Addgene here or check out these protocols if you want to package your own virus.
Topics: Viral Vectors, AAV






Leave a Comment