Remember the game show “The $25,000 Pyramid” where one player tries to get the other to guess a category by listing off things that fall into that category? Okay, let’s play. I’ll list the examples and you try to guess the category:
 ELISA...
ELISA...
qPCR...
Digital droplet PCR...
DNA dot blot...
Transduction assay...
SDS-PAGE...
Electron microscopy…
Any guesses?
That's right... ways to titer AAV!
The point is there are many ways to quantify the composition of a viral vector solution, and generally these methods measure different characteristics of the solution. These characteristics give us information on three important factors.
What can AAV titers tell you?
- Physical titer: the concentration of viral particles containing viral genomes. Physical titers are measured by quantifying the concentration of viral genomes (by qPCR or other DNA quantification methods - see below), since each viral particle typically contains one viral genome. Other than a theoretical maximum value, physical titers don’t necessarily give any indication of the infectious titer. The infectious titer of a viral vector stock can vary based on transduction target and can also be altered by freeze-thaw of the viral solution (Lock et al., 2010).
- Infectious titer: the concentration of viral particles that can transduce cells. Infectious titers are typically quantified by cell transduction assays. Wild-type AAV2 has been reported to have a near-perfect physical-to-infectious particle ratio of 1:1 (Zeltner et al., 2010). However, for recombinant AAV2, the same study reported a physical-to-infectious particle ratio of 50:1 (Zeltner et al., 2010). The specific infectivity of viral preparations is defined by the ratio of physical viral particles to infectious viral particles.
- Ratio of full to empty viral capsids: the ratio of genome-containing viral particles relative to the total number of viral capsids (which can include empty capsids that are devoid of a viral genome). Empty viral capsids can be formed during viral production depending on the experimental conditions used to generate AAV. One study showed that approximately 50% of viral capsids were genome-containing (“full”) (Zeltner et al., 2010), while another reported that only 20% of recombinant AAV2 particles were full, compared to 50% of wild-type AAV2 stocks (Grimm et al., 1999). The percentage of genome-containing viral capsids is typically quantified by electron microscopy of a viral vector solution. Since this technique is intensive and requires an electron microscope, it is not routinely performed on all new viral vector preparations. Addgene has used this method to generally validate our viral vector preparation protocol, and we are pleased to report that over 95% of viral capsids in representative viral vector preparations are genome-containing (Figure 1).
 |
|
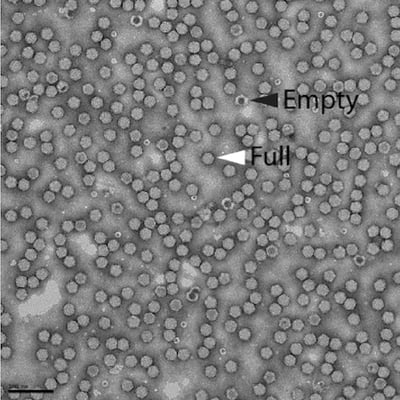
Figure 1: Electron micrograph of an Addgene AAV viral vector preparation after negative staining. Empty vector particles can be identified after negative staining and appear darker than full vector particles. This image shows that the vast majority of the vectors consist of full particles (white arrowhead) relative to empty particles (black arrowhead). Scale bar = 100nm. Image courtesy of David Bell and Svetla Stoilova-McPhie, Center for Nanoscale Systems, Harvard University. |
Which titer do people usually report?
Addgene and other viral vector manufacturing facilities report the physical titer of a viral solution (Figure 2). Because physical titers are used for dosing purposes in preclinical studies, it is important to understand what these values mean and how they can be compared.
 |
| Figure 2: Label that comes on Addgene’s AAV aliquots. Our titer values (circled in red) report physical titer, which is measured by qPCR. |
Physical titer is typically calculated by two popular PCR-based methods:
- Quantitative PCR (qPCR)
- Digital droplet PCR (ddPCR)
Historically, a quantitative DNA hybridization method (DNA dot blotting) had been used to titrate AAV, but this method is not widely used today (Fagone et al., 2012).
PCR-based methods are robust, easy, fast, and convenient. However, these methods are not necessarily precise or accurate for quantifying AAV because PCR can be affected by many experimental factors. To list some complicating factors for quantifying AAV:
- qPCR efficiency can be affected by second-order structure of the AAV genome, caused by repeats and self-complementarity of the AAV genome and the ITRs.
- qPCR efficiency can be affected by the temperature parameters. For example, we found that changing the annealing temperature of the PCR from 60° C to 61° C improved assay reliability.
- Different primers can have different annealing efficiencies (Wang et al., 2013). Although you can optimize your primers for each individual sample, this will reduce convenience and comparability because each sample with be quantified with a different primer pair.
- Ct values vary based on the amount of starting material in the PCR because the sample must fall within the linear region of the standard curve. In addition, high and low-concentration samples can also behave differently in PCR, even if both samples fall within the linear region of the assay. This may be due to competition for binding sites between the primers and the AAV genome repeats.
- Primer annealing can be affected by the presence of protein contaminants, which is a factor in AAV titration since the starting template is the purified viral vector solution (which contains viral capsid proteins) as opposed to purified DNA. This may be mitigated by using purified viral DNA as the template instead of the intact AAV particle.
- Finally, the precision of the assay can vary up to two fold for the final titer value, which is simply due to the noise of the assay. In other words, repeating the exact same PCR twice can give titer measurements that vary up to two-fold.
How can we get more reliable AAV titering?
For our viral service, we currently titrate AAV vector preparations by qPCR. To ensure accurate and reliable results, we have optimized our assay in a few ways:
- Absolute quantification by qPCR requires generating a standard curve of known concentration. We regularly make and validate fresh plasmid standards.
- We validate our absolute quantification by using the universal AAV Reference Standard Material (AAV SFM), which is an AAV sample that has been quantified by 16 labs around the world and can be used by researchers to validate their qPCR assay (Lock et al., 2010). In addition to the AAV RSM, we include a second AAV reference sample of known titer, which is at a higher titer than the AAV RSM. By using these samples in all our qPCR assays, we ensure that the standard curve generated is reliable, and can thus be used to accurately infer titer values of new AAV samples.
- We also typically quantify our AAV sample by two qPCR assays and compare the values to determine accuracy and reliability. When doing this, we make a judgement call on whether the two titer values are within the error of the assay and thus are deemed reliable, or if the sample needs to be quantified again.
Some of the issues with titrating by qPCR are addressed with ddPCR technology (Hindson et al., 2011, Hindson et al., 2013), which has been shown to be more precise and more repeatable than qPCR (Taylor et al., 2017). However, because purified AAV preps contain similar and low levels of background protein and chemical components, ddPCR may not be an improvement over qPCR (Taylor et al., 2017).
What do Addgene AAV titers mean for your experiments?
Because (physical) AAV titers are approximations and they depend highly on the experimental conditions used, titer values of samples from different sources should not be compared. If using AAV from different sources in the same experiment, consider retitering both AAV in your lab. The absolute titer values may not be as important as the relative titer values. Finally, because physical titer does not give an indication of infectious titer, consider titrating each lot of AAV in vivo to determine the optimal dosage. At Addgene, we don’t reserve lots, but if you’re reordering the same virus feel free to email us and we can do our best to send you a particular lot (if we still have it available).
References
1. Fagone, Paolo, et al. "Systemic errors in quantitative polymerase chain reaction titration of self-complementary adeno-associated viral vectors and improved alternative methods." Human Gene Therapy, Part B: Methods 23.1 (2011): 1-7. PubMed PMID: 22428975. PubMed Central PMCID: PMC3640491.
2. Grimm, D., et al. "Titration of AAV-2 particles via a novel capsid ELISA: packaging of genomes can limit production of recombinant AAV-2." Gene therapy 6.7 (1999). PubMed PMID: 10455443.
3. Hindson, Benjamin J., et al. "High-throughput droplet digital PCR system for absolute quantitation of DNA copy number." Analytical chemistry 83.22 (2011): 8604-8610.4. Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, Vessella RL, Tewari M. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods. 2013 Oct;10(10):1003-5. PubMed PMID: 22035192. PubMed Central PMCID: PMC3216358.
4. Lock, Martin, et al. "Characterization of a recombinant adeno-associated virus type 2 Reference Standard Material." Human gene therapy 21.10 (2010): 1273-1285. PubMed PMID: 20486768. PubMed Central PMCID: PMC2957240.
5.Taylor, Sean C., Genevieve Laperriere, and Hugo Germain. "Droplet Digital PCR versus qPCR for gene expression analysis with low abundant targets: from variable nonsense to publication quality data." Scientific Reports 7 (2017). PubMed PMID: 28546538. PubMed Central PMCID: PMC5445070.
6. Wang, Feng, et al. "A reliable and feasible qPCR strategy for titrating AAV vectors." Medical science monitor basic research19 (2013): 187. PubMed PMID: 23828206. PubMed Central PMCID: PMC3706409.
7. Zeltner, Nadja, et al. "Near-perfect infectivity of wild-type AAV as benchmark for infectivity of recombinant AAV vectors." Gene therapy 17.7 (2010): 872-879. PubMed PMID: 20336156. PubMed Central PMCID: PMC2900506.
Additional Resources on the Addgene Blog
- Important Considerations When Using AAVs
- Addgene's AAV Quality Control Methods
- Check Out Our Viral Vector Featured Topic Page
Resources on Addgene.org
- Find All Available AAV
- Learn More about Virus Production
- Find Viral Vector Protocols
Topics: Viral Vectors, Viral Vectors 101, AAV








Leave a Comment