This post was contributed by Max W. Shen from MIT, Alvin Hsu Harvard University, and David R. Liu from the Broad Institute and Harvard University.
Over the course of the last six months, COVID-19 has had a tremendous impact on our world -- as of June 4, 2020, COVID-19 has caused an estimated total of 380,000 deaths worldwide (statistics provided by Google on 6/4/20). Many countries entered lockdown for weeks to months, pausing or terminating employment for a significant fraction of the workforce.
Wet lab scientists are no exception to this effect -- meticulously designed experiments in labs around the world were put on hold indefinitely due to COVID-19. In our lab, we were interested in examining the effect of COVID-19 on worldwide scientific activity. However, this type of open-ended question is difficult to answer quantitatively without broad, unbiased data. Therefore, we chose to examine a dataset of Liu lab plasmid requests from Addgene, as a proxy for global activity in our particular scientific subfields.
Plasmid requests correlate with paper publication
The dataset includes 11,426 plasmid requests over 6 years for 35 papers (data was provided by Addgene on 5/26/20 and contains requests from 5/22/14 to 5/19/20). Each paper has between 1–32 plasmids and a total of 2–2,590 requests. We received plasmid requests from 56 countries, though the data is heavily dominated by the U.S. (42%), Europe (23%), and China (14%). Spikes in plasmid requests correspond to publications of popular papers.
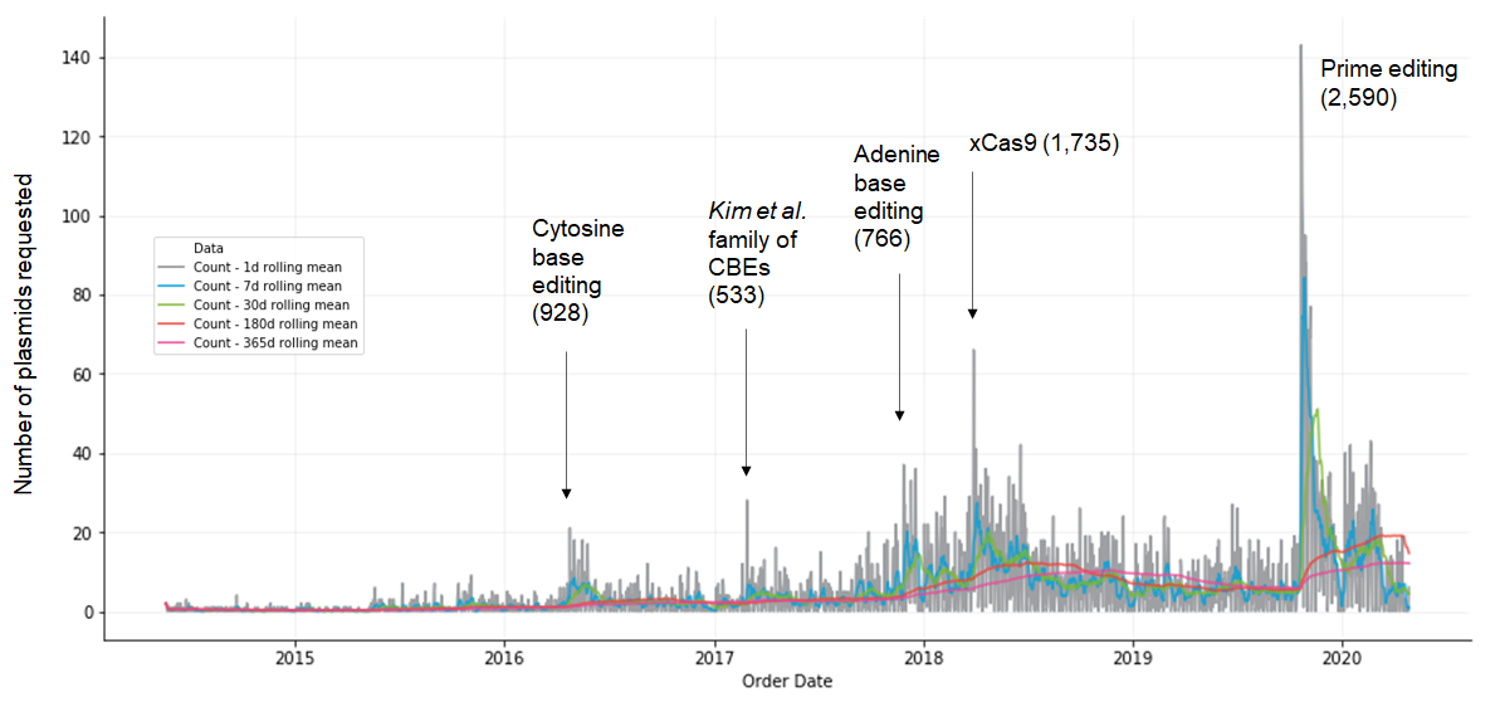 |
| Figure 1: Plasmid requests by date. Parentheses depict the total number of plasmids ordered for each paper. |
Rough estimation of COVID-19 effect on plasmid requests
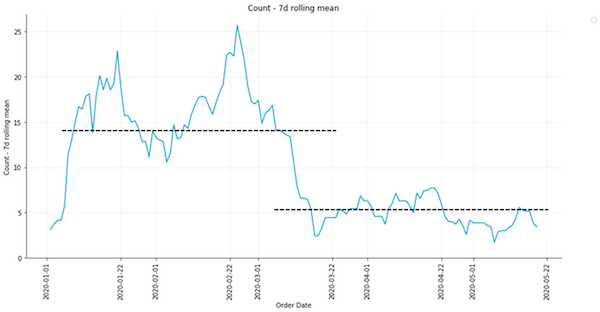
Before we began any sophisticated analyses, we first visualized our data in various ways. Smoothing our plasmid order data with a 7-day rolling mean gave us a crude insight into the magnitude of the effect COVID-19 had on our data. Because a majority of our plasmid orders come from the U.S. and Europe, we see a corresponding dip in plasmid request activity around March. A rough estimate shows that activity level in March was around 33% of that observed between Jan. 1 and Mar. 1, 2020.
 |
|
Figure 2: A simple approximation of the effect of COVID-19 on data smoothed with a 7-day rolling mean. |
While this global approximation already shows us a rough "COVID-19 effect" on order data, it does not capture the fact that different parts of the world encountered COVID-19 at different times. Most notably, China was affected by COVID-19 earlier than the rest of the world, from mid-January to mid-March.
To obtain a better view of how plasmid orders were affected by the COVID-19 pandemic, we compared the number of plasmid orders and new COVID-19 cases for each week by location. We observed that as expected, plasmid orders (left, blue) drop dramatically when there are many new COVID-19 cases being reported (right, red), likely due to quarantining
| Figure 3. An animated map showing the plasmid orders and new COVID-19 cases each week. |
Statistical Modeling
Note -- If statistical modeling isn’t up your alley, feel free to skip directly to the results.
To start modeling our data, we first chose to re-examine our global data at a finer temporal granularity. For each paper, we observed that the demand for the corresponding plasmids tended to follow an exponential decay model, with dips during weekends and significant holidays.
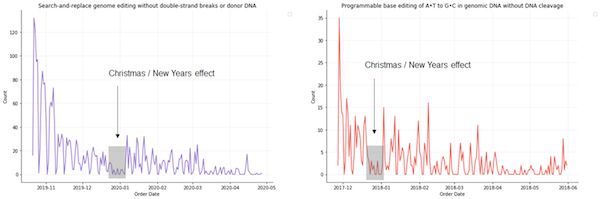 |
| Figure 4: Plasmid request trajectories suggest an exponential decay demand model with weekend and holiday effects. |
Based on these observations, we considered the following (noiseless) model for our data:
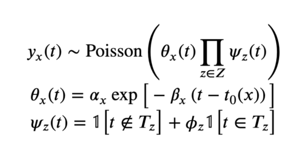 We choose to let yx(t) represent the number of unique PIs or labs placing orders for paper x on date t. We cannot use individual plasmid orders, because when a PI or lab places an order, they are likely to order several plasmids at once, breaking the statistical independence assumption of the Poisson process. This modification also eases the interpretation of the parameters we infer, since scientific productivity is better interpreted as the rate of scientists or labs placing orders.
We choose to let yx(t) represent the number of unique PIs or labs placing orders for paper x on date t. We cannot use individual plasmid orders, because when a PI or lab places an order, they are likely to order several plasmids at once, breaking the statistical independence assumption of the Poisson process. This modification also eases the interpretation of the parameters we infer, since scientific productivity is better interpreted as the rate of scientists or labs placing orders.
Our model factors the likelihood in our Poisson process into two components: 𝜃x(t) and 𝜓z(t). 𝜃x(t) describes the exponential decay process we observed, while 𝜓z(t) multiplies the likelihood by the corresponding factor 𝜙z if t is during event z. The "events" we considered for our model were weekends, Christmas/New Years, and COVID-19 lockdown. To allow our model to give uncertainty estimates for each parameter, we added log-normal noise to each of the terms which make up our distribution, resulting in the following hierarchical model:
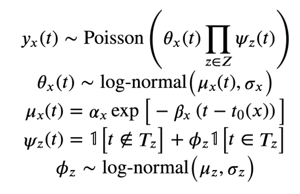
Because the U.S., Europe, and China account for over 75% of all of our plasmid orders, we will focus the rest of our discussion on these three regions. Each of these regions was affected at a different time by COVID-19, so we defined the COVID-19 event windows as 01/15/20 -- 03/15/20 for China and 03/11/20 -- 05/19/20 (the last date in our dataset) for the U.S. and Europe. To improve the stability of our model, we considered only the 10 papers with more than 10 unique order dates in 2020.
We fit our model for each region using stochastic gradient descent in Pytorch, which gave us the maximum likelihood estimates of each parameter. We approximated the data likelihood with multivariate Gauss-Hermite quadrature, since the analytical expression for the data likelihood contains an intractable integral of the product of a Poisson and multivariate log-normal likelihoods.
Effects of weekends, winter break, and COVID-19 on plasmid requests
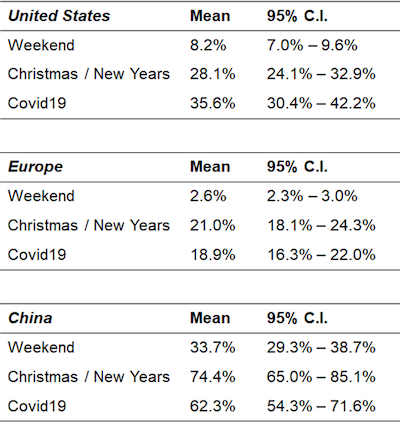 |
| Figure 5: Inferred effects. Values are the inferred percentage of normal activity (the average rate of unique PIs/labs placing orders per day) on dates impacted by each effect. |
The inferred parameters suggest that COVID-19 induced a 2x–5x reduction in the rate of scientists placing orders per day across regions, with Europe most impacted and China least impacted. The inferred weekend effect is, in retrospect, generally more dramatic than the COVID-19 effect, though we remind the reader that this is not a causal interpretation -- the effect would generally not be the same if weekends were somehow imposed on scientists.
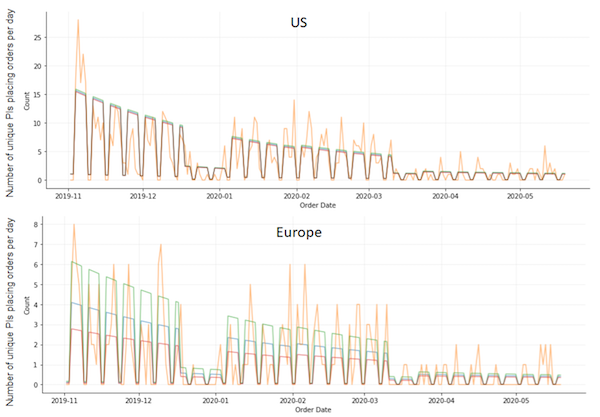
|
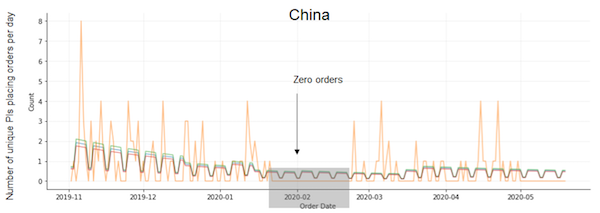
| Figure 6: Model fit. Orange line depicts observed data. Blue line is the sum of the mean rate for each paper. Green and red lines depict the sum of mean +1 and -1 std for each paper, respectively. |
The fitted model captures weekend, Christmas and New Years, and COVID-19 effects fairly well. However, we observed one complication in the China data – there is a period of 1 month, from 1/22/20–2/23/20, where there were zero plasmid requests. Afterwards, from 2/24/20–3/15/20 (the supposed end of lockdown), request activity appears to return to normal. The model fit these two discrete phases with an estimate of 62.3% for the COVID-19 effect in China. However, it is evident that if we believe that 1/22/20–2/23/20 are the more accurate dates of lockdown in China, then the observed data is compatible with an infinity-fold reduction in activity due to COVID-19 to zero. Thus, depending on when we believe the China COVID-19 date range is, they could be the most impacted or least impacted. These results highlight the instability and uncertainty in estimating parameters from relatively sparse data from China.
An important additional caveat is that our dataset contains plasmid orders from just one lab, which limits how broadly one can interpret our analysis of COVID-19’s effect on scientific activity. The effects of COVID-19 on many other scientific fields could be different.
Conclusion
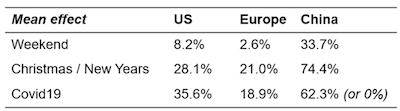 |
| Figure 7: Mean inferred effects by region. |
In conclusion, our model infers that COVID-19 induced a 2x–5x reduction in the rate of scientists placing orders per day across regions. Our investigation of model fit revealed some potential instabilities in the China data, so we recommend interpreting the model results for China with more uncertainty.
Many thanks to our guest bloggers Max W. Shen from MIT, Alvin Hsu Harvard University, and David R. Liu from the Broad Institute and Harvard University.
 Max Shen is a Ph.D. candidate at MIT. His research uses applied machine learning and statistical methods for fundamental scientific discovery and high-impact applications.
Max Shen is a Ph.D. candidate at MIT. His research uses applied machine learning and statistical methods for fundamental scientific discovery and high-impact applications.
 Alvin Hsu is a graduate student at Harvard University. He is interested in using selection, evolution, and machine learning to solve difficult problems in chemistry and chemical biology.
Alvin Hsu is a graduate student at Harvard University. He is interested in using selection, evolution, and machine learning to solve difficult problems in chemistry and chemical biology.
 David R. Liu is Director of the Merkin Institute and Vice-Chair of the Faculty at the Broad Institute; Professor of Chemistry and Chemical Biology at Harvard University; and Howard Hughes Medical Institute Investigator. Liu’s research integrates chemistry and evolution to illuminate biology and enable next-generation therapeutics. Prime editing, base editing, PACE, and DNA-templated synthesis are four examples of technologies pioneered in his laboratory. Learn more here.
David R. Liu is Director of the Merkin Institute and Vice-Chair of the Faculty at the Broad Institute; Professor of Chemistry and Chemical Biology at Harvard University; and Howard Hughes Medical Institute Investigator. Liu’s research integrates chemistry and evolution to illuminate biology and enable next-generation therapeutics. Prime editing, base editing, PACE, and DNA-templated synthesis are four examples of technologies pioneered in his laboratory. Learn more here.
Additional resources on the Addgene blog
- Learn more about plasmid sharing during COVID-19
Resources on Addgene.org
- Find plasmids from David Liu's lab
- Visit our COVID-19 plasmids and resources





Leave a Comment