Researchers express genes of interest from plasmids in order to study gene function or to engineer cells for specific purposes. Unfortunately, plasmid copy numbers vary within cell populations and over time resulting in variable gene expression that can impact observed phenotypes. Factors such as the growth medium, growth temperature, and growth rate can all impact plasmid copy number in a cell.
One can attempt to eliminate fluctuations due to copy number by introducing genes of interest into the chromosome. But this has its own set of problems. Rapidly dividing cells initiate genome replication more than once per cell division and thus, genes closer to the origin of replication will be overrepresented compared to those further away.
As a result, variable gene expression can make data hard to interpret or can interfere with the precision needed in metabolic engineering or other synthetic biology circuits. What is a molecular biologist to do? Let’s take a look at how plasmids recently deposited by Chris Voigt’s lab help keep gene expression stable regardless of copy number (Segall-Shapiro et al., 2018).
Designing stabilized promoters
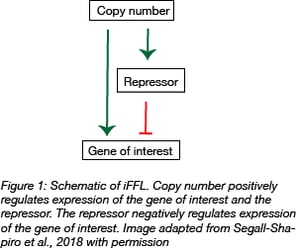 To keep gene expression stable regardless of copy number, the team made use of stabilized promoters. In contrast to unregulated promoters that are always on, stabilized promoters incorporate feedback loops such as the incoherent feedforward loop (iFFL). In general, iFFLs use an input signal that both positively and negatively controls the output. In this case, plasmid copy number is the input and gene expression is the output.
To keep gene expression stable regardless of copy number, the team made use of stabilized promoters. In contrast to unregulated promoters that are always on, stabilized promoters incorporate feedback loops such as the incoherent feedforward loop (iFFL). In general, iFFLs use an input signal that both positively and negatively controls the output. In this case, plasmid copy number is the input and gene expression is the output.
The input, plasmid copy number, directly influences gene expression. By itself, increasing the copy number of a gene increases its expression. However, if you introduce negative regulation by using a promoter that is responsive to a repressor protein, the increase in expression due to copy number is kept in check. In such an iFFL, as plasmid copy number increases, the amount of repressor protein also increases thereby intensifying repression of the gene of interest and keeping its expression level constant.
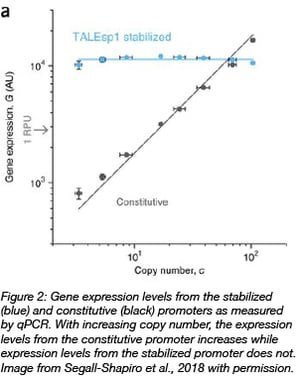 In the case of the stabilized promoters from Segall-Shapiro et al., they chose a TALE protein as the repressor because TALEs can be programmed to bind tightly to arbitrary DNA sequences and have been shown to achieve ~100 fold repression in E. coli. In addition, TALEs bind non-cooperatively to the promoter region, which is predicted to be required to decouple gene expression and plasmid number.
In the case of the stabilized promoters from Segall-Shapiro et al., they chose a TALE protein as the repressor because TALEs can be programmed to bind tightly to arbitrary DNA sequences and have been shown to achieve ~100 fold repression in E. coli. In addition, TALEs bind non-cooperatively to the promoter region, which is predicted to be required to decouple gene expression and plasmid number.
In a suite of plasmids modified from pSC101 with copy numbers ranging from 3-100 per cell, a stabilized promoter buffered against the effects of copy number on reporter gene expression. On the other hand, using a constitutive promoter in the same background resulted in a 20-fold variation of gene expression.
Because pSC101 is not representative of all types of plasmids, the stabilized promoters were also tested in different plasmid backbones that varied in size or carried different genes. In these cases, gene expression also remained stable regardless of plasmid copy number.
Stable gene expression whether growing slowly or quickly
Growth media and growth stage can both affect gene copy number. Moreover, gene copy number is affected differently by growth rates when a gene is expressed from a plasmid as opposed to the chromosome. Let’s take a look at how the stabilized promoters perform during these types of fluctuations.
When cells are dividing rapidly, multiple rounds of replication are initiated before the cell has divided. This means that genes located near the origin will be over-represented compared to those further away. While constitutive promoters drove maximal gene expression near genomic origins of replication, the stabilized promoters created by Segall-Shapiro et al. showed nearly no position-dependent differences in gene expression.
On the other hand, slow growth can also cause changes in plasmid copy number. During the shift from exponential to stationary phase, the number of plasmids per cell increases 4-5 fold (for pUC plasmids) or ~2 fold (for p15a, R6K, and ColE2 plasmids) as cell division slows (Kittleson et al., 2011). Segall-Shapiro et al. showed that the stabilized promoters were able to better modulate gene expression during shifts in growth rate as compared to a constitutive promoter.
To simulate the increase in plasmid copy number during the transition to stationary phase, Segall-Shapiro et al. placed a trans-acting RepA protein under the control of IPTG. Because RepA is a plasmid replication initiator protein, they could alter copy number of the plasmid by altering IPTG concentrations. Higher IPTG concentration leads to higher RepA expression which results in high plasmid copy number. When they increased concentrations of IPTG, a constitutive promoter upstream of GFP increased expression of GFP six-fold, but a stabilized promoter increased expression only two fold. Moreover, gene expression levels returned to the baseline after 3 hours with the stabilized promoter while gene expression from the constitutive promoter did not.
Similar levels of expression between a plasmid and the chromosome
The team also showed that the levels of gene expression from stabilized promoters were equivalent whether the genes were expressed from the genome or a plasmid. Thus, a stabilized promoter can allow a system to be moved from a plasmid to the genome without greatly impacting the level of gene expression.
For example, the synthesis of deoxychromoviridans requires three genes whose proteins are needed sequentially. When Segall-Shapiro et al. placed the genes under the control of a constitutive promoter and then moved them into the genome, deoxychromoviridans production titers decreased. But when the genes were placed under the control of a stabilized promoter, deoxychromoviridans titers were similar regardless of where the genes were expressed.
What’s the use of stable gene expression?
Constant gene expression can enable more confidence in gene expression readouts from assays such as qPCR or microarray. But, aside from facilitating the joys of a p-value < 0.05, having such fine control of gene expression is useful for many metabolic engineering and synthetic biology goals. In metabolic engineering, the expression of enzymes in a pathway will ideally be optimized to promote generation of the desired product while avoiding the accumulation of toxic intermediates or the shuttling of the intermediates into other pathways. In cases where a specific ratio of products is needed, such as within a multi-protein complex, stabilized promoters can eliminate variability between the different components. When your pathway needs precise fine-tuning of gene expression levels, give these stabilized promoters a try.
References
1. Segall-Shapiro, Thomas H., Eduardo D. Sontag, and Christopher A. Voigt. "Engineered promoters enable constant gene expression at any copy number in bacteria." Nature biotechnology 36.4 (2018): 352. PubMed PMID: 29553576.
2. Kittleson, Joshua T., Sherine Cheung, and JChristopher Anderson. "Rapid optimization of gene dosage in E. coli using DIAL strains." Journal of biological engineering 5.1 (2011): 10. PubMed
PMID: 21787416. PubMed Central PMCID: PMC3163176.
- Read the Plasmid 101 blog posts
- Check out CRISPR Meets Synthetic Biology: A Conversation with MIT’s Christopher Voigt
Resources on Addgene.org
- See the synthetic biology special collections page
- Download the Plasmid 101 eBook
- See all plasmids deposited by the Voigt lab
Topics: Plasmid Elements, Other Plasmid Tools, Plasmids






Leave a Comment