Apoptosis or “programmed cell death" plays a pivotal role in an array of biological processes including development, the immune system, and cell turnover. Apoptosis is a highly controlled process that is triggered by internal and external signals such as developmental cues and DNA damage. These signals activate a cascade of caspases, protease enzymes that cleave proteins. Executioner caspases are activated last in the cascade and are responsible for the degradation of over 600 cellular components ultimately leading to cell fragmentation and death (Elmore, 2007).
To study apoptosis in vivo, scientists have used genetically encoded fluorescent executioner caspase reporters. Several apoptosis reporters have been developed, such as the Forster resonance energy transfer (FRET)-based executioner caspase reporters. In these reporters, the FRET pair is linked by a caspase cleavage site. Cutting the linker by a caspase results in a change in fluorescence emission. FRET and fluorescence based reporters however have several limitations. First, FRET signals are often difficult to detect, and thus quantify, due to small fluorescence changes of the donor and acceptor after proteolytic cleavage. Secondly, fluorescence imaging of living organisms can be challenging due to issues with tissue autofluorescence, cell heterogeneity, and/or cell position changes (To et al., 2016, Zhang et al., 2019). Thus to effectively study apoptosis in vivo scientists need a protease reporter that has a large dynamic range and high brightness.
The Shu Lab from UCSF designed an improved apoptosis reporter, ZipGFP, which utilized the self-assembling split GFP (GFP1-10 and GFP11) to increase fluorescence 10-fold after protease activity (To et al., 2016). However, the lab wanted to increase the fluorescence range to better study the spatio-temporal dynamics of apoptosis in living animals. To do this they modified a different self-assembling split GFP (GFP1-9 and GFP10-11) by flipping a single β-strand of GFP, hence the name FlipGFP. FlipGFP increases fluorescence 100-fold compared to other reporters (Zhang et al., 2019).
How does FlipGFP work?
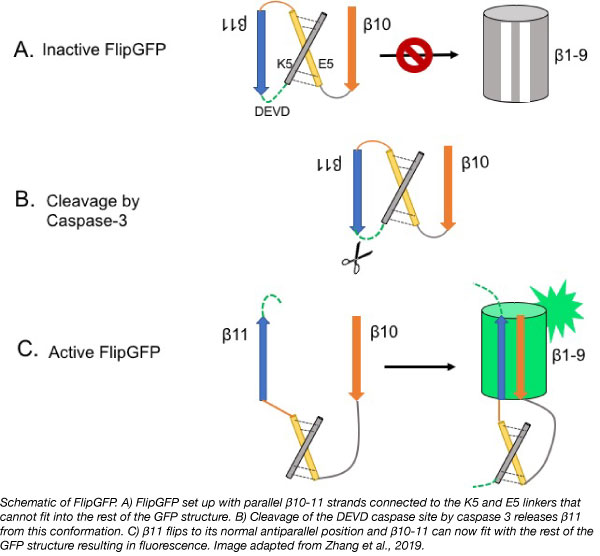
GFP contains 11 β-strands and one central ɑ-helix that can be split into 3 parts: (1) β-strands 1-9 (β1-9) and the one ɑ-helix, (2) β-strand 10 (β10), and (3) β-strand 11 (β11). Each part contains components and residues that are necessary for fluorescence. When these parts are separated from each other, the protein cannot fluoresce. When β10 and β11 are linked together, but separated from β1-9 they will rapidly recombine with β1-9 strands nearby resulting in fluorescence within tens of minutes (Cabantous et al., 2013).
The Shu Lab took advantage of this and came up with a system where fluorescence was dependent on protease activation. Normally, β10-11 are antiparallel. To prevent binding, the β strands in FlipGFP were redesigned so that they are parallel to each other and thus can no longer fit with β1-9. They connected β10-11 with two linkers (E5 and K5) that dimerize together and force the strands into a parallel conformation. Between β11 and one of the linkers is a consensus caspase-3 (an executioner caspase) cleavage site (DEVD). Upon cleavage of the sequence, β10 and β11 can rearrange to their normal antiparallel position and fluoresce.
FlipCherry: Red fluorescence protease reporter
To add to the fluorogenic reporter toolkit, the Shu Lab also designed a red fluorogenic protease reporter based on superfolder Cherry (sfCherry). Like FlipGFP, sfCherry was split into β1-9 and β10-11, where β10-11 was forced into a parallel conformation by the E5 and K5 linkers that are connected to a TEV protease cleavage site. TEV protease is a highly specific protease that is commonly used to cleave proteins. This protease reporter based on sfCherry however did not initially fluoresce in HEK293 cells. To rectify this the Shu lab first improved the self-assembling capability of sfCherry approximately 20 fold by directed evolution. They then optimized the length of both linkers to generate a reporter that would fluoresce: FlipCherry. FlipCherry provides a proof of concept that the designed used in FlipGFP can be applied to other fluorescent proteins to create an array of multicolor reporters that will enable imaging of multiple proteases at once (Zhang et al., 2019).
Studying apoptosis in vivo using the FlipGFP-based caspase reporter
The Shu Lab successfully used FlipGFP to study apoptosis in HeLa cells, zebrafish embryos, and Drosophila. They treated HeLa cells expressing FlipGFP with staurosporine, which induces apoptosis by activating caspase-3, and detected fluorescence within 2 to 5 hours, a time consistent with previous methods. In zebrafish, the Shu Lab used FlipGFP to track the spatiotemporal dynamics of apoptosis during embryo development. GFP fluorescence increased over time and showed a clear pattern in the developing forebrain and retina. This pattern was consistent with TUNEL staining, a popular method to observe apoptotic cells. Lastly, the Shu Lab demonstrated that the FlipGFP-based caspase reporter can be used to visualize apoptosis of enterocytes in the midgut of Drosophila, an observation that has been difficult until now (Zhang et al., 2019).
Conclusion
FlipGFP is a novel fluorogenic reporter that has a large dynamic range and high brightness. This reporter achieves approximately 100-fold increase in fluorescence after cleavage by executioner caspases. Failure for cells to die or cells dying at the wrong time and/or place can lead to an array of diseases and pathologies such as neurodegenerative diseases, autoimmune disorders, and cancer (Elmore, 2007). This reporter will allow scientists to observe both the spatial and temporal dynamics of apoptosis both in vitro and in living organisms to better study and understand these diseases and disorders.
You can find the plasmids for FlipGFP and FlipCherry at Addgene.
References
Cabantous, Stéphanie, et al. "A new protein-protein interaction sensor based on tripartite split-GFP association." Scientific reports 3 (2013): 2854. PubMed PMID: 24092409. PubMed Central PMCID: PMC3790201.
Elmore, Susan. "Apoptosis: a review of programmed cell death." Toxicologic pathology 35.4 (2007): 495-516. PubMed PMID: 17562483. PubMed Central PMCID: PMC2117903.
To, Tsz-Leung, et al. "Rational design of a GFP-based fluorogenic caspase reporter for imaging apoptosis in vivo." Cell chemical biology 23.7 (2016): 875-882. PubMed PMID: 27447051. PubMed Central PMCID: PMC5026494.
Zhang, Qiang, et al. "Designing a green fluorogenic protease reporter by flipping a beta strand of GFP for imaging apoptosis in animals." Journal of the American Chemical Society (2019). PubMed PMID: 30821975.
Additional resources on the Addgene blog
- Read our fluorescent proteins blog posts
- Download the fluorescent proteins 101 eBook
- Read our plasmids 101 blog series
Resources on Addgene.org
- Browse the fluorescent protein collection
- Find empty backbones for fluorescent protein fusions
- Browse the fluorescent proteins plasmid kits
Topics: Fluorescent Proteins, Other






Leave a Comment