By mutating one of two Cas9 nuclease domains, researchers created the CRISPR nickase. Nickases create a single-strand rather than a double-strand break, and when used with two adjacent gRNAs, can lower the probability of off-target editing. In this post, we’ll summarize how IDT (Integrated DNA Technologies) first demonstrated how CRISPR nickases improve homology directed repair rates, and share their design rules for your next CRISPR nickase experiment
Overview of Cas9 nickase
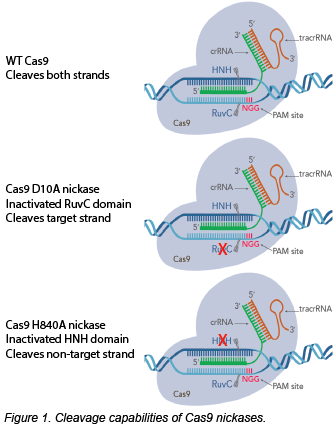
We’ll use SpCas9 nickases as examples for this post. The D10A mutation inactivates the RuvC domain, so this nickase cleaves only the target strand. Conversely, the H840A mutation in the HNH domain creates a non-target strand-cleaving nickase. Instead of cutting both strands bluntly with WT Cas9 and one gRNA, you can create a staggered cut using a Cas9 nickase and two gRNAs.
For nickase applications, a common question is: how should the gRNAs be oriented in comparison to each other? The gRNAs must target different strands to create a DSB, but this can be accomplished with either a PAM-in or PAM-out orientation. Just as the names imply, PAM-out designs have the PAM sequences on the extremes of the targeted region, whereas PAM-in designs place the PAMs closer together in the middle of the targeted region, as shown below.
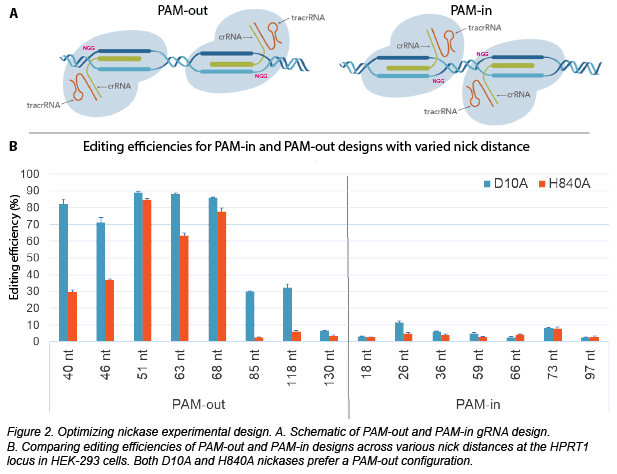
To identify optimal nickase designs, IDT scientists Mollie Schubert and Shuqi Yan designed experiments to test nickase preferences using HEK293 cells. They started by comparing D10A and H840A total editing efficiency in PAM-in and PAM-out configurations with varied nick distances (40 - 130 nt). From these results, it’s clear that editing is much higher in the PAM-out configuration. In addition, D10A editing efficiency is consistently higher than that of H840A, especially with smaller nick distances. Using D10A, they subsequently showed that editing efficiency is very low when the gRNAs are too close (7-23 nt nick distance).
Both D10A and H840A are potent editors, but their indel profiles vary. In the PAM-out configuration, D10A tends to generate small deletions, while H840A is biased towards large insertions. These profiles are likely due to the distinct overhang patterns generated by the nickases; D10A creates 5’ overhangs and H840A creates 3’ overhangs in a PAM-out design.
Exploring nickases for homology directed repair
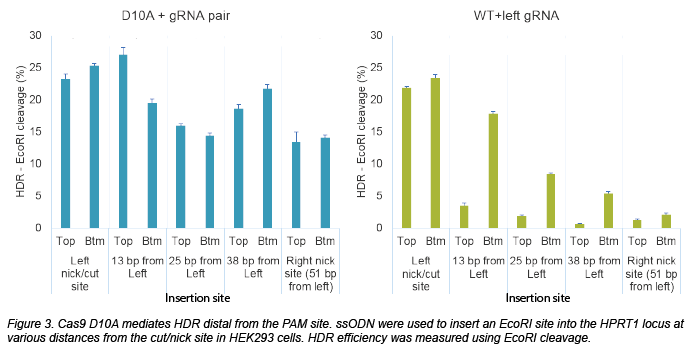
The potential benefit of using nickases for HDR is targeting range: using an individual gRNA with WT Cas9, repair levels decrease rapidly 10 bp from the cut site. So if you can’t find a good gRNA that cuts close to your insertion site, you can’t obtain high HDR efficiency. Nickases create a staggered cut - could this system mediate repair throughout the entire region between the nicks? To find out, Schubert and Yan designed a PAM-out nickase experiment to insert an EcoRI site using a single-stranded oligonucleotide (ssODN) donor. Using a D10A nickase to insert an EcoRI site in various spots between the nick sites, they observed >20% repair (max 27%) across the 51 nt region using either a top or bottom strand ssODN donor with 40 bp homology arms. The HDR efficiency compared favorably to using WT Cas9 with the left gRNA across the window. In a subsequent experiment, they were also able to introduce a longer insertion (mCherry) using IDT Megamer long ssDNA with 100 nt homology arms.
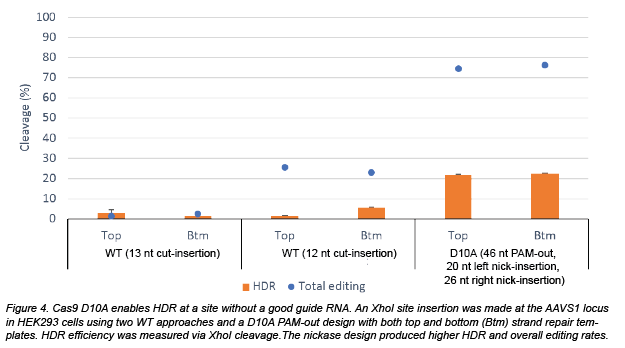
Schubert and Yan next examined a situation thought to be suboptimal for HDR. At the AAVS1 locus, the two nearest gRNAs had cleavage sites 12 nt and 13 nt away from the target site rather than the optimal <10 nt distance. Employing two gRNA sites that are further away from the target site, they designed a D10A nickase strategy (46 nt nick distance, PAM-out) to see if they could solve this problem. D10A nickase outperformed both WT Cas9 sites tested by a wide margin, exhibiting >20% repair efficiency. Clearly, a nickase strategy can expand the targetable region, which is especially useful if there are no available guides close to the desired mutation site.
Quick tips for nickase design
Ready to employ nickases for your next CRISPR experiments? Here’s IDT’s best advice:
- Use a PAM-out configuration
- Optimize your spacing
- D10A: nick sites separated by 37-68 bp
- H840A: nick sites separated by 51-68 bp
- Use D10A for HDR
- Place intended insert in between nick sites
- Use 40 nt homology arms for small insertions or tags (ssODN; IDT Ultramer® Oligonucleotides)
- Use 100 nt homology arms for large insertions (long ssDNA; IDT Megamer® ssDNA Fragments)
- Test both bottom and top strand ssDNA donors (if possible)
The text and images in this post were adapted from an IDT webinar: Optimized methods to use Cas9 nickases in genome editing. For more information and a link to the archived webinar, click here. We would like to thank IDT scientists Mollie Schubert and Shuqi Yan for reviewing this blog post.
Additional Resources on the Addgene Blog
Resources on Addgene.org
- Check out CRISPR Topic Page
- Find CRISPR Nickase Plasmids
- Browse the CRISPR Guide
Topics: CRISPR, CRISPR 101, Cas Proteins






Leave a Comment