This post was co-authored by Susanna Stroik and Rachel Leeson.
Here at Addgene, we like to help share useful new resources with the scientific community – particularly ones that help address ongoing problems, like cells lines. A 2014 study in Science found that approximately one-third of cell lines used in scientific research are misidentified! Rigor and reproducibility of scientific data depends on validated reagents along with clear documentation, so we were excited when Amos Bairoch from the Swiss Institute of Bioinformatics reached out to us to tell us about a resource called Cellosaurus, a growing cell line information database describing nearly 145,000 cell lines.
Cellosaurus
Cellosaurus seeks to identify and describe all cell lines used in research, using four methods to search out cell lines:
- Periodic surveying of cell line collections websites and catalogues
- Literature searches utilizing Google Scholar alerts and the NCBI LitSuggest Tool
- Following the “rabbit hole” of related cell lines when annotating a cell line
- Community submissions via email: cellosaurus@sib.swiss
Entries include accession number, species, and assigned category (cancerous, hybridoma, etc.), as well as sex and age at time of sampling if available. STR profiling for human, mouse, and dog cell lines is included when available, and can be submitted via email for cell lines profiled in the lab. You can browse entries by cell line groups, such as their vaccine production, SARS-CoV-2 research, or adenovirus packaging cell lines dataset collections.
Depending on available information, entries may also include whether a cell line is known to be genetically manipulated, its anatomical origin, if cell line has ever been the target of an ‘-omics’ study or is resistant to specific compounds, links to publication references, and cross-references with cell line catalogs and collections.
Importantly, Cellosaurus also identifies problematic cell lines: ones either suspected to be contaminate or easily contaminated, as well as commonly misidentified lines. Browsing this feature to know if your lines may be especially vulnerable can be very informative. Spoiler alert: HeLa is the most common contaminator line.
For those wanting to access Cellosaurus programatically, their API allows fine grained query and retrieval of all the information in the resource.
This is a labor-intensive project; scripts can be used in some cases, but much of this work has required manually going through the pdfs of the 26,000 publications cited. If the information seems incomplete or unclear, Bairoch follow up via email to the authors of the paper in which the cell line was described.
Using Cellosaurus
Cellosaurus is searchable by cell line name (recommended names and synonyms will all get you to the right entry.) You can look up the cell lines you are currently or planning to use for helpful information on the known features and relevant literature, or to decide which line is best for your experiments.
You can also use the Cellosaurus STR similarity search tool (CLASTR) to find cells lines similar to your STR profiles of interest. You can filter CLASTR results by minimal number of markers and similarity score match. This tool’s algorithm can be adjusted based on query, reference, and shared allele search preferences to help you identify proximal lines you care about.
If you have cell lines without a certified origin (you know, that split you got from the graduate student down the hall who got it from some technician downstairs who got it from who knows where), now is as good a time as ever to find out the true identity of the line? Once you’re run your STR profiling, you can use Cellosaurus, and CLASTR, to finally know whether it was an 'h' or a 'b' written on the flask (and if the name was the correct one!)
|
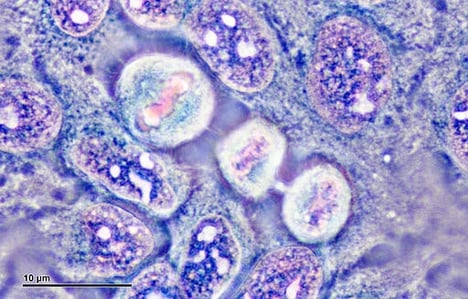
| Fig. 1: HeLa cells. Image courtesy of Josef Reischig. |
Contributing to Cellosaurus
Generated a new cell? Or finding errors in the available information about a cell line you’ve used? Submit your manuscript or data to Cellosaurus via cellosaurus@sib.swiss.This community resource is actively encouraging submissions from scientists all over globe, with some surprising, and important, findings being shared.
For instance, recently Cellosaurus received a report that HT-55, which was described as a colon cancer cell line, is actually a rectal cancer cell line. Using the data provided by the researcher, Cellosaurus was able to both update their database and sent the updated information to the cell line distributor. A success all around!
If you’re excited about this resource and want to learn more, check out the Cellosaurus FAQ!
References
Lorsch JR, Collins FS, Lippincott-Schwartz J. Cell biology. Fixing problems with cell lines. Science 2014; 346:1452–1453.
Bairoch, A. The Cellosaurus, a Cell-Line Knowledge Resource. J Biomol Tech. 2018; 29(2): 25-38.






Leave a Comment