RNA-editing Cas13 enzymes have taken the CRISPR world by storm. Like RNA interference, these enzymes can knock down RNA without altering the genome, but Cas13s have higher on-target specificity. New work from Konermann et al. and Yan et al. describes new Cas13d enzymes that average only 2.8 kb in size and are easy to package in low-capacity vectors! These small, but mighty type VI-D enzymes are the latest tools in the transcriptome engineering toolbox.
How were Cas13d enzymes discovered?
Microbial CRISPR diversity is impressive, and researchers are just beginning to tap the wealth of CRISPR possibilities. To identify Cas13d, both groups used very general bioinformatic screens that looked for a CRISPR repeat array near a putative effector nuclease. The Cas13d proteins they identified have little sequence similarity to previously identified Cas13a-c orthologs, but they do include HEPN nuclease domains characteristic of the Cas13 superfamily. Yan et al. proceeded to study orthologs from Eubacterium siraeum (EsCas13d) and Ruminococcus sp. (RspCas13d), while Konermann et al. characterized orthologs from “Anaerobic digester metagenome” (AdmCas13d) and Ruminococcus flavefaciens (nicknamed CasRx), as well as EsCas13d.
How do Cas13d enzymes compare to other Cas13s?
Like other Cas13 enzymes, the Cas13d orthologs described in these papers can independently process their own CRISPR arrays into guide RNAs. crRNA cleavage is retained in dCas13d and is thus HEPN-independent. These enzymes also do not require a protospacer flanking sequence, so you can target virtually any RNA sequence! In bacteria, Cas13d-mediated cleavage promotes collateral cleavage of other RNAs. As with other Cas13s, this collateral cleavage does not occur when Cas13d is expressed in a mammalian system.
Since Cas13d is functionally similar to previously discovered Cas13 enzymes - what makes these orthologs so special? The first property is size - Cas13d enzymes have a median length of ~930aa - making them 17-26% smaller than other Cas13s and a whopping 33% smaller than Cas9! Their small size makes then easy to package in low-capacity vectors like AAV, a popular vector due to its low immunogenicity. But these studies also identified other advantages, including Cas13d-specific regulatory proteins and high targeting efficiency, both of which are described below.
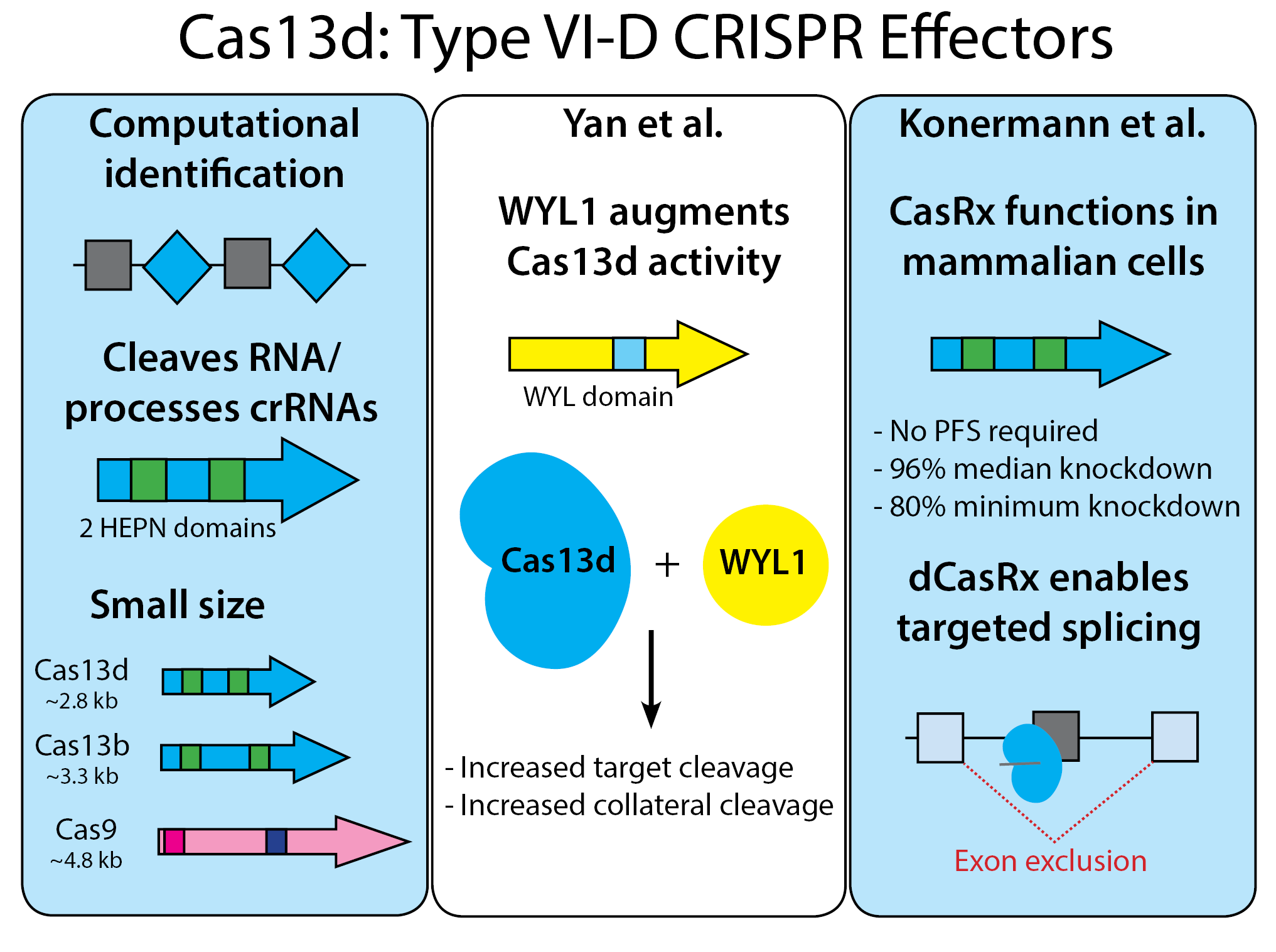
Yan et al.: WYL-domain-containing accessory protein WYL1 increases RspCas13d and EsCas13d cleavage activity
The majority of Type VI-D loci contain accessory proteins with WYL domains (named for the three conserved amino acids in the domain). Yan et al. from Arbor Biotechnologies found that RspCas13d accessory protein RspWYL1 increases both targeted and collateral RNA degradation by RspCas13d. RspWYL1 also increased EsCas13d activity, indicating that WYL domain-containing proteins may be broader regulators of Cas13d activity. This property makes WYL proteins an intriguing counterpart to anti-CRISPR proteins that negatively modulate the activity of Cas enzymes, some of which are also functional in multiple species (read Arbor Biotechnologies' press release about their Cas13d deposit here).
Konermann et al.: CasRx is a highly potent, specific editor in mammalian cells
Not all Cas13d proteins are functional in mammalian cells, but Konermann et al. saw great results with CasRx and AdmCas13d fused to a nuclear localization signal (NLS). In a HEK293 mCherry reporter assay, CasRx and AdmCas13d produced 92% and 87% mCherry protein knockdown measured by flow cytometry, respectively. Cas13d CRISPR array processing is robust, with CasRx and either an unprocessed or processed gRNA array (22 nt spacer with 30 nt direct repeat) mediating potent knockdown. Multiplexing from the CRISPR array yielded >90% knockdown by CasRx for each of four targets, including two mRNAs and two nuclear long non-coding RNAs.
One interesting twist to Cas13d enzymes is their cleavage pattern: EsCas13d produced very similar cleavage products even when guides were tiled across a target RNA, indicating that this enzyme does not cleave at a predictable distance from the targeted region. Konermann et al. show that EsCas13d favors cleavage at uracils, but a more detailed exploration of this cleavage pattern is necessary.
Konermann et al. compared CasRx to multiple RNA regulating methods: small hairpin RNA interference, dCas9-mediated transcriptional inhibition (CRISPRi), and Cas13a/Cas13b RNA knockdown. CasRx was the clear winner with median knockdown of 96% compared to 65% for shRNA, 53% for CRISPRi, and 66-80% for other Cas13a and Cas13b effectors. Like previously characterized Cas13 enzymes, CasRx also displays very high on-target efficiency; where shRNA treatment produced 500-900 significant off-targets, CasRx displayed zero. Unlike Cas9, for which efficiency varies widely across guide RNAs, each guide tested with CasRx yielded >80% knockdown. It seems that CasRx may make it possible to target essentially any RNA in a cell.
Since catalytically dead dCasRx maintains its RNA-binding properties, Konermann et al. tested its ability to manipulate RNA species through exon skipping. Previous CRISPR exon-skipping approaches used two guide RNAs to remove a given exon from the genome, and showed success in models of muscular dystrophy. In this case, Konermann et al. targeted MAPT, the gene encoding dementia-associated tau, delivering dCasRx and a 3-spacer array targeting the MAPT exon 10 splice acceptor and two putative splice enhancers. After AAV-mediated delivery to iPS-derived cortical neurons, dCasRx-mediated exon skipping improved the ratio of pathogenic to non-pathogenic tau by nearly 50%, showing proof-of-concept for pre-clinical and clinical applications of dCasRx.
The identification of Type VI Cas13d enzymes is another win for bioinformatic data mining. As we continue to harness the natural diversity of CRISPR systems, only time will tell how large the genome and transcriptome engineering toolbox will be. It is, however, certain that the impact of CRISPR scientific sharing will continue to grow, and we at Addgene appreciate our depositors for making their tools available to the broader community.
References
Konermann, Silvana, et al. “Transcriptome Engineering with RNA-Targeting Type VI-D CRISPR Effectors.” Cell (2018) pii: S0092-8674(18)30207-1. PubMed PMID: 29551272
- Plasmids from this paper are available at Addgene.
Yan, Winston X., et al. “Cas13d Is a Compact RNA-Targeting Type VI CRISPR Effector Positively Modulated by a WYL-Domain-Containing Accessory Protein.” Mol Cell. (2018) pii: S1097-2765(18)30173-4. PubMed PMID: 29551514
- Plasmids from this paper are available at Addgene.
- Learn more about research at Arbor Biotechnologies.
Additional Resources on the Addgene Blog:
- Learn about CRISPR RNA Editing with REPAIR
- Get tips for using Cas9 Transcriptional Activators
- Learn about Cas13a (C2c2) Characterization
Resources on Addgene.org:
- Find RNA Knockdown Plasmids
- Check out our CRISPR Guide
- Read our CRISPR History
Topics: CRISPR, Cas Proteins





Leave a Comment