Transcription, translation, and prone to degradation – those are the words that describe RNA! Double stranded? Well, that’s just for DNA – right?
RNA performs almost all of its biological functions in our cells in the single strand form, but double stranded RNA and RNA:DNA hybrid molecules both exist and have a diverse range of functions within mammalian cells. Here, we will cover how these structures are generated, why it’s important for cells to sense their formation, and how clinicians, researchers, and even farmers are harnessing these molecules for good.
RNA:RNA
dsRNA and viral infection
Viral genomes come in all sorts of shapes and sizes. Many are encoded by single stranded RNA (ssRNA), while some exist as dsRNA. Still others use dsDNA to store their genetic information. All of these viruses have one thing in common – they can generate dsRNA as part of their infection and/or replication process within the host. dsRNA viruses are inherently double stranded, while ssRNA viruses can generate dsRNA intermediates as part of the viral replication process. dsDNA viruses yield dsRNA through a slightly different mechanism thought to be associated with bidirectional transcription. In an uninfected cell, dsRNA rarely circulates, but upon viral infection, this nucleic acid can build up. The foreign dsRNA activates the innate immune response machinery of the host.
This pathway has been clinically exploited for antiviral purposes similar to that of vaccines. If you know the sequence of a virus, you could introduce some of its dsRNA intermediates into a healthy individual to provoke an immune response and then, hopefully, immunity to that virus. This system is certainly a double-edged sword – balancing antiviral benefits with the inflammatory response associated with the introduction of dsRNA.
Synthetic dsRNA
Researchers have been harnessing the power of dsRNA for a long time, even before we knew about all the diverse in vitro production mechanisms. siRNA, also known as RNAi, are short, dsRNA molecules designed to degrade mRNA transcripts. RNAi activates the RNA-induced silencing complexes (RISCs), which detect dsRNA in the cytoplasm and use it as a guide to complementary RNA sequence. Once a compatible sequence is found (typically RNA that the RNAi is designed against), RISC can repress translation and/or degrade target mRNA.
These RNA tools have been used in clinical therapy to target pathogenic proteins, as well as in the lab to deplete expression of proteins of interest. RNAi has even made it into agriculture as species-specific pesticides and insecticides, an advancement over previous chemical versions with toxicity to many organisms.
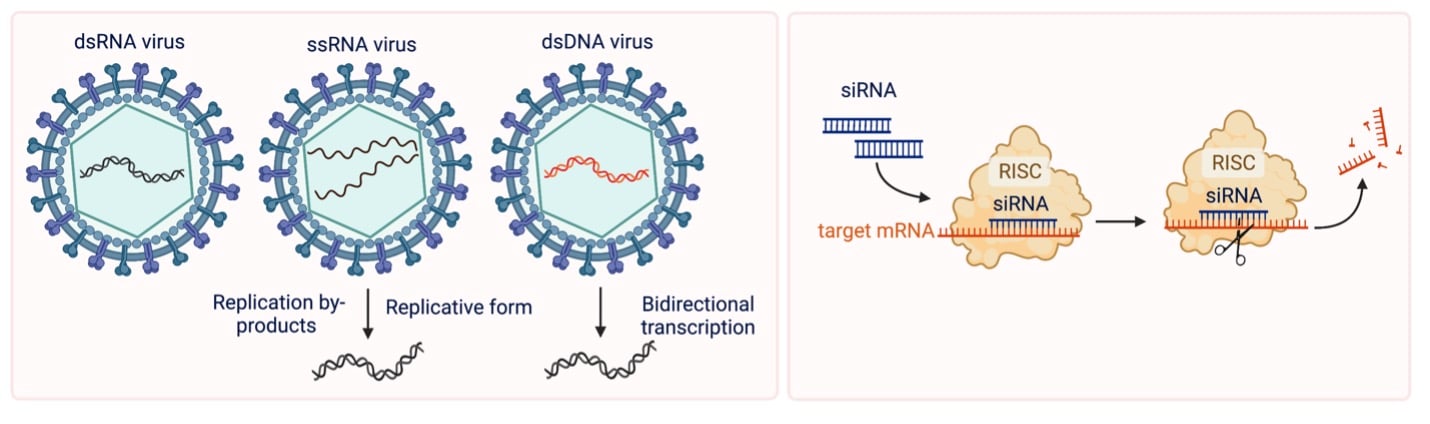 |
| Fig. 1: Exogenous sources of dsRNA |
Cellular sources of dsRNA
Transposable elements such as long/short interspersed nuclear elements can generate dsRNA by bidirectional transcription or as inverted repeats. These elements are almost always epigenetically silenced, but dysregulation of these systems can lead to local dsRNA production. In the same vein, genotoxic stress, such as chemotherapy treatment or ionizing radiation, can also result in dsRNA generation. While these molecules are clearly associated with cellular damage and dysfunction, it is unclear what the precise mechanism behind the production of these dsRNAs is.
A major source of dsRNA (that doesn’t involve cellular dysregulation) is the mitochondria. These organelles have their own circular genomes which can produce dsRNA as a result of bidirectional transcription of their circularized DNA. These dsRNAs are typically contained within the mitochondria, barring a loss of mitochondrial integrity that would release them into the cytosol.
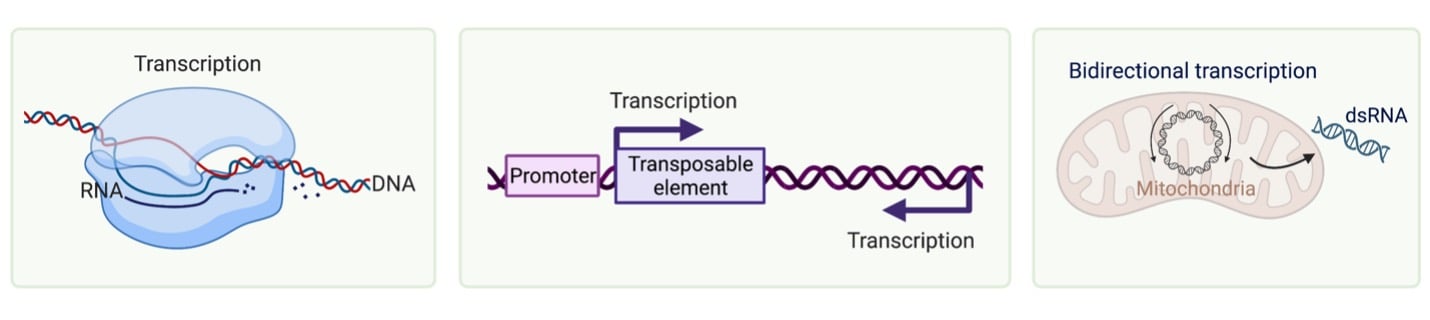 |
| Fig. 2: Endogenous sources of duplexed RNA |
DNA:RNA hybrids
Sometimes ssRNA just needs a buddy, and even DNA will suffice. This can result in DNA:RNA hybrids. Unlike dsRNA, DNA:RNA hybrids are abundant in normal cells and are arise from a ssRNA engaging one strand of a dsDNA molecule; the 3-stranded structure produced is known as an R-loop.
During transcription, RNA is synthesized from a DNA template, inherently generating RNA:DNA intermediates in the process. These transcription-coupled R-loops are transient in nature and will dissociate when gene transcription is complete. However, transcription of thousands of genes is constantly occurring throughout a single cell, so these nucleic acid hybrids are always around. R-loops are particularly enriched in highly transcribed regions and repetitive regions, such as telomeres and centromeres. These genomic regions are prone to persistent R-loop formation, presumably due to their abundance of R-loops and repetitive sequence.
All of the R-loops described above play distinct roles in promoting cellular processes and functions, but they also have a darker side. When ssRNA engages one strand of a dsDNA molecule, it does so at the expense of the unpaired DNA strand. A ssDNA molecule exists within the R-loop that is now vulnerable to various forms of DNA damage, and these structures can in turn lead to genome instability. Thus, it’s important to strike a balance between productive levels of R-loops and putting DNA at risk.
Consequences of duplexed RNA
Mammalian cells have multiple receptors in place to sense dsRNA and respond accordingly. While not perfect, these sensing systems work to distinguish between RNA of cellular vs. viral origin and respond appropriately. When these sensing systems fail, diseases and steep cellular consequences arise.
dsRNA
An immune response is activated if and when dsRNA levels reach a given threshold within the cell, as is often the case upon viral infection. Endogenously produced dsRNA rarely reach this threshold and thus do not typically activate an inflammatory response. However, if dsRNA sensors are mutated or mechanisms which suppress endogenous dsRNA formation are dysregulated, immune responses can mount in the absence of infection. Similarly, if endogenously synthesized dsRNA finds its way outside of the mitochondria, the same inflammatory response can activate. These types of cellular dysregulation have been documented in several autoimmune disorders, including some forms of lupus and rheumatoid arthritis.
These types of immune responses are also a concern when it comes to RNAi therapeutic delivery. Thus, balancing the amount of siRNA to be delivered or anticipating this response are important considerations when using these reagents.
RNA:DNA hybrids
Persistent RNA:DNA hybrids that arise during transcription are typically recognized by RNase H which hydrolyzes the RNA portion, allowing the DNA strands to re-pair. The two dominating RNases in mammalian cells (RNase H1 and H2) handle most of these intermediates, with each one acting on specific substrates and in distinct cell cycles. When RNA:DNA hybrids levels are allowed to reach very high or low levels, possibly due to dysregulation of removal mechanisms, an array of diseases can result (Fanconi Anemia, cancer, Fragile X syndrome, etc.).
Hopefully now you can appreciate double stranded isn’t just for DNA. Untwisting RNA is important for therapeutic delivery, viruses, human disease, and much more!
Resources and References
References
Wang, I. X., Grunseich, C., Fox, J., et al., Human proteins that interact with RNA/DNA hybrids. Genome Res., 28(9): 1405-1414 (2018). 10.1101/gr.237362.118
Chen, G. Y., Hur, S. Cellular origins of dsRNA, their recognition and consequences. Nat Rev Mol Cell Biol, 23: 286-301 (2022). doi.org/10.1038/s41580-021-00430-1
More resources on the Addgene blog
Topics: Miscellaneous





Leave a Comment