 First described in the 1980s, protein tags are now one of the most useful items in a scientist’s toolbox. As we’ve covered in Plasmids 101, tags can help you determine localization of a protein of interest, purify it, or determine its expression level without the need for a custom antibody. There is one major caveat - a tag may interfere with protein localization and/or function, so each tagged protein must be tested carefully to ensure it retains the attributes of the native protein. Since this process takes a lot of time and energy, Martí Aldea and collaborators have created a set of “innocuous tags” (inntags) less likely to alter a protein’s properties.
First described in the 1980s, protein tags are now one of the most useful items in a scientist’s toolbox. As we’ve covered in Plasmids 101, tags can help you determine localization of a protein of interest, purify it, or determine its expression level without the need for a custom antibody. There is one major caveat - a tag may interfere with protein localization and/or function, so each tagged protein must be tested carefully to ensure it retains the attributes of the native protein. Since this process takes a lot of time and energy, Martí Aldea and collaborators have created a set of “innocuous tags” (inntags) less likely to alter a protein’s properties.
Why do we need new epitope tags?
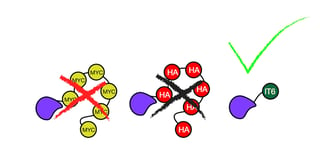
Short epitope tags like FLAG, HA, and Myc can adopt a number of different conformations and thus potentially interact with and alter the proteins they label. In fact, a recent study of 400 proteins found that 20% of tagged proteins did not localize similarly to their untagged counterparts. These findings motivated Georgieva et al. to develop new tags with defined 3D structures less likely to compromise protein folding and function. Using bioinformatic analyses, they also selected for other key properties, notably high solubility, limited affinity for other proteins, and accessibility to an antibody. From their initial pool of candidates, they selected 12 candidate protein domains from non-vertebrate systems. Preliminary testing indicated that inntags IT5, IT6, and IT10 expressed well in cells and did not form undesirable aggregates.
Using budding yeast as a model system, Georgieva et al. examined the effects of GFP fusions to inntags or traditional FLAG, HA, and Myc tags. Impressively, IT6-GFP did not reduce yeast cell size or viability compared to untagged GFP overexpression, while the rest of the fusions had some negative effect on these parameters. IT5 and IT6 displayed good binding efficiency in both immunofluorescence and immunoprecipitation applications, but none of the traditional tags were suitable for both applications.
To prove that inntags are indeed innocuous, Georgieva et al. created tagged versions of the cyclin Cdc28 and the chaperone Ydj1, both of which display sophisticated interactions with multiple other proteins. Again, little to no change in cell morphology was observed with IT5 and IT6 fusions. In contrast, Flag and HA tags had especially detrimental effects on cell size and growth rate, indicating that they compromised protein-protein interactions.
If you’re looking to a tag a protein, inntags IT5 and IT6 are a great place to start. These tags are easy to use in multiple common applications. Due to their limited effects on native protein structure, inntags may become the tags of choice for examining protein-protein interactions, particularly in single-molecule or high-throughput applications. Perhaps IT5 and IT6 will even replace our older epitope tags - only time will tell!
References
1. Georgieva, Maya V., et al. "Inntags: small self-structured epitopes for innocuous protein tagging." Nature methods (2015). PubMed PMID:26322837.
2. Stadler, Charlotte, et al. "Immunofluorescence and fluorescent-protein tagging show high correlation for protein localization in mammalian cells." Nature methods 10.4 (2013): 315-323. Pubmed PMID: 23435261.
Resources at Addgene
Read about other Protein Tags in our Plasmids 101 Series.
Get the Inntag Plasmids
Tag endogenous proteins with CRISPR
Topics: Plasmid Tags, Plasmids






Leave a Comment