 All organisms share an innate goal to survive. This past year, scientists hijacked survival tactics of prokaryotes to deliver the technological biological blockbuster known as the CRISPR (clustered regularly interspaced palindromic repeats) Cas (CRISPR associated genes) system. This popular genome engineering tool offers flexibility, multiplexibility, and ease of use. In order for this technology to survive the diverse demands of the biotech field, let's look at how prokaryotes originally utilized CRISPR/Cas as a powerful and adaptive defense strategy against life-threatening viruses.
All organisms share an innate goal to survive. This past year, scientists hijacked survival tactics of prokaryotes to deliver the technological biological blockbuster known as the CRISPR (clustered regularly interspaced palindromic repeats) Cas (CRISPR associated genes) system. This popular genome engineering tool offers flexibility, multiplexibility, and ease of use. In order for this technology to survive the diverse demands of the biotech field, let's look at how prokaryotes originally utilized CRISPR/Cas as a powerful and adaptive defense strategy against life-threatening viruses.

Evolving to survive
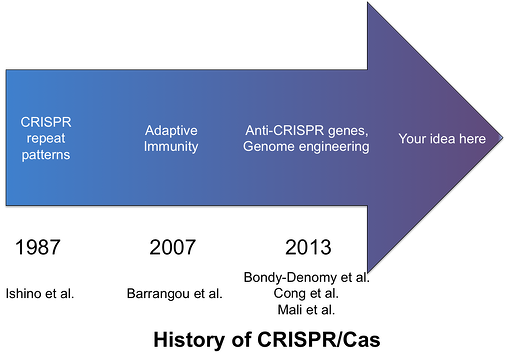
Although the CRISPR sequences were initially discovered in E. coli in 1987 (1), the concept that these clustered repeat sequences operated as a safeguard against bacteriophages did not come to light until 2007. Initial experiments exposed S. thermophilus to predatory phages to test if the exogenous DNA would be incorporated into the bacterial genome. Cas genes, which code for polymerases, nucleases, and helicases, were also disrupted to elucidate their role in the process. The results of these studies led scientists to hypothesize that prokaryotes developed an adaptive immune system that utilized various cas genes not only to store a record of invading phages but to also destroy the phage upon re-exposure (2, 3). More specifically, specialized Cas proteins snip the foreign DNA into small fragments approximately 30bp in length and paste them into the CRISPR sequence. Separate Cas proteins then express and process the CRISPR loci to generate RNA, which guide a Cas nuclease to the specified exogenous genetic material next to the species-specific protospacer adjacent motif (PAM). Once the CRISPR/Cas complex binds to the foreign DNA, a cut is made to destroy the invader.
 |
| Wikipedia, accessed 25 November 2013. Author: James Atmos (2). |
Many CRISPR/Cas moieties carry out these functions using slightly different Cas proteins and PAM sequences. A total of eight evolutionarily conserved CRISPR subtypes comprised of more than 40 cas gene families have been characterized. To date, 83% of archaeal and 45% of bacterial genomes utilize this system (4). Makarova et al. grouped the systems into three distinct types and subtypes using polythetic classification based on evolutionary similarity (5). Type II CRISPR/Cas is the system that scientists have harnessed for genome engineering. The diversity of the CRISPR/Cas systems provides powerful lines of defense against invading phages.
Rising to the challenge - Evolution of anti-CRISPR genes in phages
The CRISPR/Cas adaptive immune system seems like the winning ticket to ensure prokaryotic survival; however, viruses, too, are famous for their ability to devise new strategies to reproduce within a host cell. A few virus particles have learned to circumvent the CRISPR/Cas defense by generating a single point mutation in the PAM sequence, preventing the Cas nucleases from re-identifying it (3). Surprisingly, few genes have been identified that neutralize CRISPR/Cas. Earlier this year, expression of 1 of 5 anti-CRISPR genes were found to inactivate the Type I-F CRISPR/Cas system of Pseudomonas aeruginosa (6). However, these genes only deactivate the Type I-F system and are not translatable to the other CRISPR subtypes. Still, the large and dynamic range of CRISPR systems between and within prokaryotic species suggest the presence of a variety of undiscovered viral anti-CRISPR genes. These data highlight an interesting "arms race" between phages and prokaryotes; the ultimate victor will be the species who has the strongest will to survive.
Genome engineering and future directions
CRISPR/Cas inspired genome editing was initially described by Cong et al. and Mali et al. in January 2013 (7, 8). Nine months later, more than 1500 publications describe work to improve the tool’s specificity, orthogonality, and multiplexibility in various species and describe an assortment of applications. Despite the ubiquity of CRISPR/Cas toolkit, genome engineers continue to work diligently and creatively to deliver a highly specific, programmable platform well-suited for various innovative biological and translative technologies.
Addgene has empowered researchers to harness previous experimental successes and further develop the CRISPR/CAS toolkit by posting generic lab protocols, providing tips from experts in the field, and enabling access to multiple plasmids used for various platform applications. For more information on CRISPR/Cas, check out Addgene’s CRISPR Guide or find CRISPR/Cas plasmids for your research.
Do you have a cool idea or application that involves CRISPR/Cas technology? Let us know.
References
-
Ishino Y, Hideo, S, Makino K, Mitsuko A, Nakata A. J Bacteriol. 1987 Dec; 169 (12): 5429–33.
-
Photo: Wikipedia, accessed 25 November 2013. Author: James Atmos. http://en.wikipedia.org/wiki/File:Crispr.png
-
Barrangou R, Horvath P. Science. 315, 2007 Jan; 1709–1712 (2007).
-
CRISPRdb. Date: 2013-09-18.
-
Makarova K, Haft DH, Barrangou R, Brouns SJJ, Charpentier E, Horvath P, Moineu S, Mojica FJM, Wolf YI, Yakunin JvdO, Koonin EV. Nature Reviews Microbiology. 2011 Jun; 9, 467-477.
-
Bondy-Denomy J, Pawluk A, Maxwell KL, Davidson AR. Nature. 2013 Jan; 493, 429–432.
-
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F. Science. 2013 Jan; 339 (6121): 819–23.
-
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM. Science. 2013 Jan; 339 (6121): 823–6.
Topics: CRISPR, CRISPR 101






Leave a Comment