In this heyday of molecular biology, many scientists do a lot of DNA work but never get to actually manipulate the organism they study (unless if you count normal human interaction for all of us studying human genes in test tubes and gels). As a freshman in college I studied development and evolution of histone genes in Strongylocentrotus purpuratus, the spiny purple sea urchin. It’s not a standard model organism, but it was easy to harvest eggs and sperm, mix them in a tank, and observe the beauty of embryo development in real time. I was already a biology geek, but this formative experience nailed down my plan to become a research scientist. I recently was invited to attend the first Principal Investigator meeting of the NSF EDGE (Enabling Discovery Through Genomic Tools) Program and I met scientists from 21 different labs who clearly also share this wonder of the organism.
It’s not an overstatement to say that CRISPR/Cas9 editing is one of the key tools now providing access to genotypic study of diverse organisms. Methods for genetic manipulation still need to be developed for each organism, but researchers are optimistic that this can be done for most, if not all, of these new experimental species. Discovering ways to culture, propagate, transform, store, and share all of these new organisms continues to be challenging. Organism specific viral vector systems would also be beneficial for many systems.
How the EDGE Program promotes scientific sharing
The EDGE Program is designed to expand the number of systems that can be studied by providing support "for development of tools, approaches, and infrastructure necessary for direct tests of cause and effect hypotheses between gene function and phenotypes in diverse plants, animals, microbes, viruses, and fungi for which these methods are presently unavailable.” As stated by one of the NSF program directors, this program should “enable the ability to study life’s innovations from anywhere in the tree of life.” Accepted proposals were required to present research plans that went beyond obtaining “omic” data and had to include clear plans to connect Genotype to Phenotype thus making progress on the NSF “G to P grand challenge.”
Creating a productive, new scientific community is no small task. The EDGE program solicitation requires funded labs to rapidly disseminate protocols, methods, data, and tools that result from their work. Researchers are encouraged to not just wait for papers to be published, but to engage in ongoing, real-time dissemination of information and materials. Many groups publicized workshops on their organismal systems for scientists and educational programs for K-12 students. You can see why I was excited to be invited to attend. My job is to ensure that, in general, these labs all include materials sharing in their dissemination plans and in particular, that they all share the plasmids created under this programs via Addgene.
New organisms for genotype to phenotype studies
New organismal systems described at this meeting were extraordinarily diverse. Plans to develop tools to study their genotypes and phenotypes were presented in a series of quick (10 minute) or very quick (2 minute) talks so we got to hear from everyone. Addgene is certainly going to have to find ways to tag and curate reagents from these emerging organisms so scientists can easily find the useful new reagents that come out of this work as the communities using them grow. So, think beyond Drosophila melanogaster or Saccharomyces cerevisiae (both fine research organisms) - perhaps you’ll choose to study Crepidula atrasolea or Prochlorococcus with your next project.
Gymnotiformes and Mormyridae (Jason Gallant)
Weakly electric fish are excellent systems to study for convergent evolution because electric organs arose in different branches of the evolutionary tree at least 6-7 times and on at least 2 separate continents.
Mimulus (Andrea Sweigart)
Monkeyflowers were studied by Darwin and are now studied in 40 labs as a great model for plant adaptation and speciation. Their phenotypes are diverse and they inhabit all habitats - even 50 meters from Old Faithful in 60 °C soils.
Dendrobatidae (Lauren O’Connell)
Poison frogs are great for studying parenting strategies and make important ecological indicators since species live in water and on land. Current tools are all for Xenopus (aquatic frogs), but there are few genetic tools currently for terrestrial frogs.
Cnidarian-Dinoflagellate symbionts (Virginia Weis)
Sea anemone Aiptasia - algae Breviolum minutum symbionts are important models for solutions to the coral reef crisis. I like how they are using protocols.io to create a protocol community!
Asclepias syriaca and Asclepias curassavica (Gerog Jander)
Milkweed plants are known to many as the only Monarch butterfly food. They have chemical pathways generating interesting drug-like compounds especially cardenolides which have been used to treat heart disease. Caterpillars that eat milkweed generate cardenolides that tastes yucky (and is poisonous) for bird predators - birds eat a caterpillar once, vomit, and never do it again.
Cichlids (Scott Juntti)
Cichlids are 2,000 fish species that evolved in the last 10,000 years (which is considered recent). They have diverse traits with good genetic mapping morphology and interesting behaviors. There is diversity in the social bonds that they form including genetic mechanisms that control who a female wants to mate with and if she decides to raise their offspring or leave them to their own devices.
Chytrid fungi (Lillian Fritz-Laylin)
Chytrid fungi are useful for the study of basic evolution of fungi and eukaryotes and includes the fungus that is killing frogs all over the world. These are less well known fungi (they are not yeast or mushrooms) but there are over 1,000 species. No transformation techniques are available yet.
Polistes (Amy Toth)
These wasps are like the ones that nest on your house. Their nests are open so you can see them and even feed them RNA to do silencing in the wild. They also return to their nests so can be studied over time. They recognize individual faces (I guess of other wasps) so can be used to study aspects of cognition.
Spiralians - Crepidula atrasolea (Jonathan Henry)
This black footed slipper snail can be manipulated by injected RNA and can express fluorescent proteins. This lab has created an automated tank for feeding and maintenance that had the audience gasping. Animal husbandry of diverse species is a real bottleneck.
New gif for #MolluscMonday: #Crepidula atrasolea juvenile, light microscope with polarizing filter. Development is direct in this species, and hatchlings are around 1mm long pic.twitter.com/sIIoIsIFd1
— Henry Lab (@EvoEmbryo) September 24, 2018
Xenopus tropicalis (Gary Gorbsky)
This is not a new experimental system, but the Gorbsky Lab is using Xenopus frog cell lines to develop robust methods for frog gene editing. They edit in a cell line, then take the nucleus and put it in a fertilized egg to get edited embryos (replacing the nucleus is still very difficult).
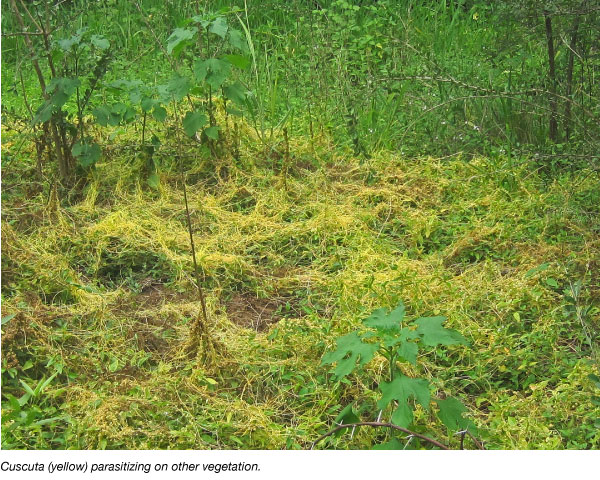
Cuscuta (James Westwood)
Dodders are parasitic plants that steal all their food from other plants and are unusual enough to be interesting biologically. They have economic importance as harmful weeds. The parasitic fusion is such that the two plants share RNA, proteins, and DNA.

Euprymna scolopes (Joshua Rosenthal)
The Hawaiian bobtail squid has an extreme number of nerve cells and an unusually high level of RNA recoding in the brain. RNA editing in the brain allows the squid to create a different protein. Did you know that most cephalopods die after they lay their 1 clutch of eggs? Propagation is thus quite difficult. CRISPR genome editing in squid is possible nonetheless - the first example was of a pigmentation gene.
Genus Microtus (Zoe Donaldson)
Voles have fascinating social behavior. For example, two similar species have very different social behavior. One is monogamous and one is promiscuous and the gene that is responsible is understood. It’s possible to actually silence this gene with RNAi to make a polygamous species monogamous.
Anolis sagrei (Doug Menke)
No methods were available to create targeted gene mutations in anoles lizards or in any reptiles at all. But new success is coming with CRISPR/Cas in lizards. The Menke Lab was able to make an albino lizard with targeted mutations. Now what? What would you study first? They are starting by seeing if they can change long legged lizards into short legged lizards.
Zebra finches (Claudio Mello)
These very cute birds learn their songs like humans learn speech. Finches must learn early, use auditory feedback to learn, babble as baby birds, and have dialects just like people. This is actually rare in the animal kingdom.
Gasterosteus aculeates (Dan Bolnick)
A paper on stickleback fish science was first published in 1867. It said they were hard to raise because they beat on each other in captivity. Sticklebacks demonstrate unusual parallel adaptation to disparate environments so can be used study the genetic basis of evolution in immunology, neurobiology, and more.
Prochlorococcus (Penny Chisholm)
This cyanobacterium is the smallest photosynthetic cell in existence, but most abundant cell on earth (twice the human biomass). The lab is developing a system using novel vesicles to efficiently transform these cells which don’t like foreign DNA at all (maybe because they have only 1,700 genes total).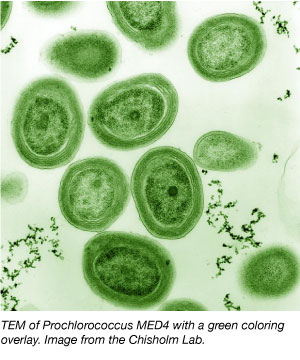
Hydra (Celina Juliano)
This is a known stem cell and regeneration model. Little bits of hydra can grow back into a full hydra and hydra have asexual and sexual reproductive life cycles. One can inject plasmid DNA in the eggs to get transgenic animals. They might be immortal.
Manihot esculenta (Blake Meyers)
Cassava is the fifth most grown crop in the world so it feeds a lot people. This lab is trying to improve homologous recombination efficiency to reduce the number of plants you need to grow to get a recombinant because this process is expensive. They are using a cool fluorescent indicator system to rapidly identify recombinants.
Arthropods (Jason Rasgon)
This lab is developing tools for pan-arthropod genetic manipulation. The ReMOT control system (Receptor-Mediated Ovary Transduction of Cargo) delivers Cas9 RNP to the arthropod germline by injection into adult female mosquitoes. Now they are trying to replace embryo injection (hard to do, expensive, not every lab has the capability) with receptor mediated endocytosis. They used this to edited those brown marmorated stink bugs to make them pink.
We’re excited to see new tools being developed and shared for many different organisms. Do you have a favorite organism you study in the lab? Let us know!
Additional resources on the Addgene blog
- Check out our other model organism blog posts:






Leave a Comment