Scientists routinely use techniques to alter gene expression or to label specific cells, but there are too few resources to teach students how to perform these experiments in the beginning. In most classrooms, the laboratory experience is focused on classical embryology techniques such as basic observation and dissections. Students don’t usually perform more modern techniques used in genetics or molecular biology because the experiments are either not accessible or too challenging for amateur scientists. Planarians, wormy creatures commonly found in freshwater ponds, provide a good potential solution to this problem. Planarians are easy to buy, cultivate, and have interesting phenotypes to study. In addition, the Sánchez lab has made it easier to perform advanced developmental biology experiments in planarians with their recent plasmid deposit.
These plasmids enable two main types of experiments:
- Whole mount in Situ Hybridization (WISH) for visualizing expression of genes.
- Double-stranded RNA (dsRNA)-mediated RNA interference (RNAi) to knockdown expression of specific gene expression.
The 49 deposited plasmids enable students to study gene expression and functions of key regeneration-related genes in four inexpensive and commercially available planarians: Girardia sp., Dugesia dorotocephala, Phagocata morgani, and Phagocata gracillis. There are 11 different genes cloned into plasmids to investigate for each of these planarian species.
These plasmids all have a PR-T4 vector backbone, which is a modified version of L440. This vector is versatile in that it can be used for both in situ hybridization and dsRNA-induced RNAi experiments. Notably, this plasmid harbors a multi-cloning site flanked by T7 promoters to allow for bidirectional transcription to create dsRNA. Students can produce DNA templates via PCR off of the same plasmid and use these to create riboprobes for in situ hybridization experiments.
A recent publication [1] from the Sánchez lab describes the plasmids and protocols required for both WISH and RNAi experiments, as well as how to perform worm amputations. They also created an online companion resource called “cutting class” which provides more detailed materials and protocol resources.
Visualizing gene expression via Whole Mount In Situ Hybridization (WISH)
WISH is a technique that detects the temporal and spatial localization of mRNA transcripts in tissue. This is useful for labeling specific cell types and for viewing how gene expression changes through time or after genetic manipulation. The technique involves using synthetically made RNA (riboprobes) with modified nucleotides (for example, Digoxigenin-labelled Uracil) that will bind to complementary mRNA within an embryo or tissue. Once the riboprobes are created, the WISH protocol involves preparation of the tissue sample, several washes, and temperature changes to visualize the expression of specific genes. After riboprobe binding (via hybridization) specific antibodies that detect the mRNA-riboprobe complex are added to the tissue. The last step involves a chemical reaction to stain the cells that contain this mRNA-riboprobe-antibody complex and, as a result, these cells turn blue.
Students can create synthetic riboprobes off of linearized DNA templates which they can easily generate off of a circular plasmid via PCR. Typically, to make these constructs, developmental biologists have to extract RNA from their organism of interest, use the RNA to create a complementary DNA (cDNA), and then use PCR and molecular cloning to insert the cDNA into the desired plasmid backbone. The Sánchez lab plasmids allow students and new labs to bypass these steps and focus on the WISH technique. The expression patterns of 11 different planarian genes (all of which are available through the deposit) are described in the Sánchez lab publication [1] so that the students performing these experiments have a good baseline for comparison. Performing WISH using these plasmids can result in the labeling specific cell types or entire organs in planarians. For example, the gene Piwi labels neoblast cells and the gene Prohormone convertase 2 (pc2) labels the central nervous system.
Evaluating Gene function by RNA interference (RNAi)
RNA interference (RNAi) is a powerful tool for performing targeted knockdowns of genes of interest and was first discovered and characterized in an unrelated worm (C. elegans) [2]. Delivering dsRNA matching a specific mRNA sequence triggers a cascade of events that results in the degradation of the target mRNA and silences its expression. More specifically, during the process of RNAi, an enzyme dubbed Dicer cuts or “dices” dsRNA into smaller chunks. These smaller dsRNA fragments are referred to as small interfering RNAs (siRNA) and they bind to a class of Argonaute proteins. The Argonaute proteins remove one strand of the siRNA and are directed by complementarity to the remaining single stranded siRNA to bind natively expressed mRNAs.These native mRNAs are subsequently destroyed by Argonaute-catalyzed nuclease activity or their translation disrupted through physical blocking of the translation machinery.
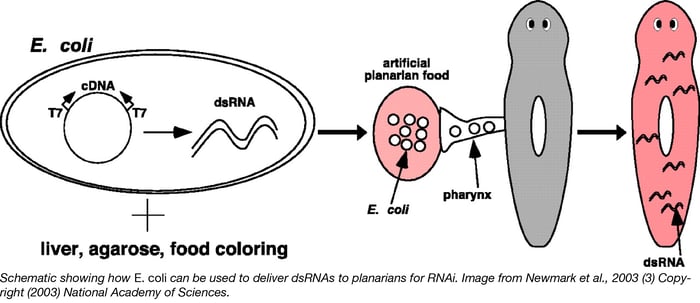
In the Sánchez lab protocol, dsRNAs are delivered by feeding them to planarians. Depending on which planarian species one is working with, the protocol will slightly differ. For example, Girardia sp. and Dugesia dorotocephalacan can be fed dsRNA-expressing E. coli that are mixed with liver paste. Prior to mixing with the liver paste, these E. coli are transformed with the appropriate plasmid and dsRNA expression is activated through the induction of T7 polymerase encoded in the E. coli genome (usually controlled by IPTG). For this IPTG-induced expression to work, students have to use a commercial E. coli strain that is designated as DE3.
For Phagocata morgani and Phagocata gracillis, the process is a little more complicated. These strains are more picky eaters and refuse to eat liver paste mixed with E. coli. Thus, to get the dsRNA into Phagocata morgani and Phagocata gracillis, one has to purify the dsRNA from bacteria and mix the purified dsRNA with liver paste, and feed the mixture to the worms.
The phenotypes of RNAi knockdown treatment should be apparent after three dsRNA feedings. Some of the phenotypes are not evident until after amputation. For example, the dramatic “two-headed” phenotype is observable in post-amputation planarians after knockdown of beta-catenin. Other gene targets do not require amputation to be visible. For example, odf2 gene knockdown results in an observable “sidewinder” phenotype that disrupts wildtype locomotion.
Create your own plasmids to investigate other genes in planarians!
For those who are interested in performing their own molecular cloning and investigating genes not in this collection, they may clone DNA sequences for creating riboprobes and dsRNAs into PR-T4P. The Sánchez lab protocol also provides step-by-step instructions for the “the bravest and more curious” students on how to clone a gene of interest into the empty PR-T4P vector backbone.
The empty vector PLL4, which is similar to PR-T4P, is available from Addgene.
If you use these published protocols and want to share pictures, feedback etc. please write to cuttingclass@stowers.org and if you make new plasmids for investigating different planarian genes, we encourage you to deposit them back with Addgene and expand the number of genes that other classrooms and researchers can easily investigate in planarians.
References
1. Alice Accorsi, Monique M. Williams, Eric J. Ross, Sofia M. C. Robb, Sarah A. Elliott, Kimberly C. Tu, Alejandro Sánchez Alvarado, The American Biology Teacher, Vol. 79 No. 3, March 2017; (pp. 208-223). Link: http://abt.ucpress.edu/content/79/3/208
2. Fire, S. Xu, M.K. Montgomery, S.A. Kostas, S.E. Driver, C.C. Mello. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature, 391 (1998), pp. 806-811. PubMed PMID: 9486653.
3. Newmark, Phillip A., et al. "Ingestion of bacterially expressed double-stranded RNA inhibits gene expression in planarians." Proceedings of the National Academy of Sciences 100.suppl 1 (2003): 11861-11865. PubMed PMID: 12917490. PubMed Central PMCID: PMC304099.
Additional Resources on the Addgene Blog
- Arabidopsis in Education
- Learning Synthetic Biology with BioBuilder
- 3D Models for learning about CRISPR
Resources at Addgene.org
- Find plasmids from the Sánchez lab
- Find plasmids for other developmental models including flies, nematodes, and zebrafish
- Find plasmids for Mammalian RNAi





Leave a Comment